Abstract
DNA cytosine methylation plays a vital role in repressing retrotransposons, and such derepression is linked with developmental failure, tumorigenesis and aging. DNA methylation patterns are formed by precisely regulated actions of DNA methylation writers (DNA methyltransferases) and erasers (TET, ten-eleven translocation dioxygenases). However, the mechanisms underlying target-specific oxidation of 5mC by TET dioxygenases remain largely unexplored. Here we show that a large low-complexity domain (LCD), located in the catalytic part of Tet enzymes, negatively regulates the dioxygenase activity. Recombinant Tet3 lacking LCD is shown to be hyperactive in converting 5mC into oxidized species in vitro. Endogenous expression of the hyperactive Tet3 mutant in mouse oocytes results in genome-wide 5mC oxidation. Notably, the occurrence of aberrant 5mC oxidation correlates with a consequent loss of the repressive histone mark H3K9me3 at ERVK retrotransposons. The erosion of both 5mC and H3K9me3 causes ERVK derepression along with upregulation of their neighboring genes, potentially leading to the impairment of oocyte development. These findings suggest that Tet dioxygenases use an intrinsic auto-regulatory mechanism to tightly regulate their enzymatic activity, thus achieving spatiotemporal specificity of methylome reprogramming, and highlight the importance of methylome integrity for development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-seq, WGBS and ACE-seq data were deposited in the Gene Expression Omnibus under the accession code GSE222501. CUT&Tag data were deposited in GSE222618. Source data are provided with this paper.
References
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Li, E. & Zhang, Y. DNA methylation in mammals. Cold Spring Harb. Perspect. Biol. 6, a019133 (2014).
Yoder, J. A., Walsh, C. P. & Bestor, T. H. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 13, 335–340 (1997).
Reik, W., Dean, W. & Walter, J. Epigenetic reprogramming in mammalian development. Science 293, 1089–1093 (2001).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Lio, C. J. et al. TET methylcytosine oxidases: new insights from a decade of research. J. Biosci. 45, 21 (2020).
Iyer, L. M., Tahiliani, M., Rao, A. & Aravind, L. Prediction of novel families of enzymes involved in oxidative and other complex modifications of bases in nucleic acids. Cell Cycle 8, 1698–1710 (2009).
Hu, L. et al. Crystal structure of TET2-DNA complex: insight into TET-mediated 5mC oxidation. Cell 155, 1545–1555 (2013).
Upadhyay, A. K., Horton, J. R., Zhang, X. & Cheng, X. Coordinated methyl-lysine erasure: structural and functional linkage of a Jumonji demethylase domain and a reader domain. Curr. Opin. Struct. Biol. 21, 750–760 (2011).
Bauer, C. et al. Phosphorylation of TET proteins is regulated via O-GlcNAcylation by the O-linked N-acetylglucosamine transferase (OGT). J. Biol. Chem. 290, 4801–4812 (2015).
Gu, T. P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Chen, B. et al. Maternal inheritance of glucose intolerance via oocyte TET3 insufficiency. Nature 605, 761–766 (2022).
Zeng, T. B., Han, L., Pierce, N., Pfeifer, G. P. & Szabo, P. E. EHMT2 and SETDB1 protect the maternal pronucleus from 5mC oxidation. Proc. Natl Acad. Sci. USA 116, 10834–10841 (2019).
Nakamura, T. et al. PGC7 binds histone H3K9me2 to protect against conversion of 5mC to 5hmC in early embryos. Nature 486, 415–419 (2012).
Zhang, Q. et al. Differential regulation of the ten-eleven translocation (TET) family of dioxygenases by O-linked beta-N-acetylglucosamine transferase (OGT). J. Biol. Chem. 289, 5986–5996 (2014).
Lewandoski, M., Wassarman, K. M. & Martin, G. R. Zp3-cre, a transgenic mouse line for the activation or inactivation of loxP-flanked target genes specifically in the female germ line. Curr. Biol. 7, 148–151 (1997).
Geis, F. K. & Goff, S. P. Silencing and transcriptional regulation of endogenous retroviruses: an overview. Viruses 12, 884 (2020).
Sakashita, A. et al. Endogenous retroviruses drive species-specific germline transcriptomes in mammals. Nat. Struct. Mol. Biol. 27, 967–977 (2020).
Yan, R. et al. Dynamics of DNA hydroxymethylation and methylation during mouse embryonic and germline development. Nat. Genet. 55, 130–143 (2023).
Guo, F. et al. Active and passive demethylation of male and female pronuclear DNA in the mammalian zygote. Cell Stem Cell 15, 447–459 (2014).
Shirane, K. et al. Mouse oocyte methylomes at base resolution reveal genome-wide accumulation of non-CpG methylation and role of DNA methyltransferases. PLoS Genet. 9, e1003439 (2013).
Kaneda, M. et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429, 900–903 (2004).
Lane, N. et al. Resistance of IAPs to methylation reprogramming may provide a mechanism for epigenetic inheritance in the mouse. Genesis 35, 88–93 (2003).
Lees-Murdock, D. J., De Felici, M. & Walsh, C. P. Methylation dynamics of repetitive DNA elements in the mouse germ cell lineage. Genomics 82, 230–237 (2003).
Hedges, D. J. & Deininger, P. L. Inviting instability: transposable elements, double-strand breaks, and the maintenance of genome integrity. Mutat. Res 616, 46–59 (2007).
Marangos, P. & Carroll, J. Oocytes progress beyond prophase in the presence of DNA damage. Curr. Biol. 22, 989–994 (2012).
Eymery, A., Liu, Z., Ozonov, E. A., Stadler, M. B. & Peters, A. H. The methyltransferase Setdb1 is essential for meiosis and mitosis in mouse oocytes and early embryos. Development 143, 2767–2779 (2016).
Kim, J. et al. Maternal Setdb1 is required for meiotic progression and preimplantation development in mouse. PLoS Genet. 12, e1005970 (2016).
Karimi, M. M. et al. DNA methylation and SETDB1/H3K9me3 regulate predominantly distinct sets of genes, retroelements, and chimeric transcripts in mESCs. Cell Stem Cell 8, 676–687 (2011).
Walsh, C. P., Chaillet, J. R. & Bestor, T. H. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat. Genet. 20, 116–117 (1998).
Jonsson, M. E. et al. Activation of endogenous retroviruses during brain development causes an inflammatory response. EMBO J. 40, e106423 (2021).
Matsui, T. et al. Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 464, 927–931 (2010).
Riso, V. et al. ZFP57 maintains the parent-of-origin-specific expression of the imprinted genes and differentially affects non-imprinted targets in mouse embryonic stem cells. Nucleic Acids Res. 44, 8165–8178 (2016).
Inoue, A., Shen, L., Matoba, S. & Zhang, Y. Haploinsufficiency, but not defective paternal 5mC oxidation, accounts for the developmental defects of maternal Tet3 knockouts. Cell Rep. 10, 463–470 (2015).
Tsukada, Y., Akiyama, T. & Nakayama, K. I. Maternal TET3 is dispensable for embryonic development but is required for neonatal growth. Sci. Rep. 5, 15876 (2015).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Deplus, R. et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J. 32, 645–655 (2013).
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal 14, 1 (2016).
Deniz, O., Frost, J. M. & Branco, M. R. Regulation of transposable elements by DNA modifications. Nat. Rev. Genet. 20, 417–431 (2019).
Kellinger, M. W. et al. 5-Formylcytosine and 5-carboxylcytosine reduce the rate and substrate specificity of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 19, 831–833 (2012).
Garcia-Perez, J. L., Widmann, T. J. & Adams, I. R. The impact of transposable elements on mammalian development. Development 143, 4101–4114 (2016).
Peaston, A. E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004).
Liu, S. et al. Setdb1 is required for germline development and silencing of H3K9me3-marked endogenous retroviruses in primordial germ cells. Genes Dev. 28, 2041–2055 (2014).
Comiskey, M. & Warner, C. M. Spatio-temporal localization of membrane lipid rafts in mouse oocytes and cleaving preimplantation embryos. Dev. Biol. 303, 727–739 (2007).
Cangelosi, A. L. et al. Zonated leucine sensing by Sestrin-mTORC1 in the liver controls the response to dietary leucine. Science 377, 47–56 (2022).
Jin, S. G. et al. Tet3 reads 5-carboxylcytosine through Its CXXC domain and is a potential guardian against neurodegeneration. Cell Rep. 14, 493–505 (2016).
Rodriguez, C. I. et al. High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat. Genet. 25, 139–140 (2000).
Jackson-Grusby, L. et al. Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation. Nat. Genet. 27, 31–39 (2001).
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. A FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Zhong, C. et al. CRISPR-Cas9-mediated genetic screening in mice with haploid embryonic stem cells carrying a guide RNA library. Cell Stem Cell 17, 221–232 (2015).
Xue, J. H. et al. A vitamin-C-derived DNA modification catalysed by an algal TET homologue. Nature 569, 581–585 (2019).
Li, Z. et al. Gadd45a promotes DNA demethylation through TDG. Nucleic Acids Res. 43, 3986–3997 (2015).
Yin, R. et al. Ascorbic acid enhances Tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J. Am. Chem. Soc. 135, 10396–10403 (2013).
Zhang, R. et al. Manganese salts function as potent adjuvants. Cell Mol. Immunol. 18, 1222–1234 (2021).
Ge, Y. Z. et al. Chromatin targeting of de novo DNA methyltransferases by the PWWP domain. J. Biol. Chem. 279, 25447–25454 (2004).
Castrillon, D. H., Miao, L., Kollipara, R., Horner, J. W. & DePinho, R. A. Suppression of ovarian follicle activation in mice by the transcription factor Foxo3a. Science 301, 215–218 (2003).
Zhang, C. et al. The chromatin remodeler Snf2h is essential for oocyte meiotic cell cycle progression. Genes Dev. 34, 166–178 (2020).
Yang, H., Wang, H. & Jaenisch, R. Generating genetically modified mice using CRISPR/Cas-mediated genome engineering. Nat. Protoc. 9, 1956–1968 (2014).
Gu, C., Liu, S., Wu, Q., Zhang, L. & Guo, F. Integrative single-cell analysis of transcriptome, DNA methylome and chromatin accessibility in mouse oocytes. Cell Res 29, 110–123 (2019).
Qian, J., Zhu, R., Yan, R., Long, X. & Guo, F. Isolation of mouse ovarian follicles for single-cell RNA-seq and in vitro culture. STAR Protoc. 3, 101537 (2022).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Yan, R. et al. Decoding dynamic epigenetic landscapes in human oocytes using single-cell multi-omics sequencing. Cell Stem Cell 28, 1641–1656.e7 (2021).
Yan, R., Cheng, X. & Guo, F. Protocol for scChaRM-seq: simultaneous profiling of gene expression, DNA methylation, and chromatin accessibility in single cells. STAR Protoc. 2, 100972 (2021).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform 14, 178–192 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Schutsky, E. K. et al. Nondestructive, base-resolution sequencing of 5-hydroxymethylcytosine using a DNA deaminase. Nat. Biotechnol. 36, 1083–1090 (2018).
Acknowledgements
We thank M. Dawlaty, J. Walter, Q. Wang, T. Chen, A. Susor, Q. Xu and T.-P. Gu for critical reading and discussion of the manuscript; J. Li (Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences) for providing AG-haESCs and Blimp1-Cre knockin mice; Z. Jiang (Peking University) for providing MnJ adjuvant; L. Wu for bioinformatic analysis; and G.-F. Xu and S.-S. Cheng for constructing the Tet3 mutant mouse line. This study was supported by grants from National Science Foundation of China (grant nos. 82088101 and 31991163 to G.-L.X., 32370632 to Y.-R.D.), the Ministry of Science and Technology China (grant nos. 2018YFA0800302 to G.-L.X., 2019YFA0109900 and 2017YFC1001301 to Y.-R.D.), the Sino-German Mobility Program (grant no. M-0313 to G.-L.X.), Shanghai Municipal Science and Technology Major Project (to G.-L.X.), the Open Project Program of Shanghai Key Laboratory of Embryo Original Diseases (grant no. Shelab2020ZD02 to Y.-R.D.) and the CAS Project for Young Scientists in Basic Research (grant no. YSBR-012 to F.G.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
G.-L.X. and Y.-R.D. designed the experiments. Y.-R.D., B.-B.H. X.-J.Z. and Z.-Y.J. carried out CRISPR–Cas9 editing, mouse modeling, phenotyping, bisulfite-seq and ACE-seq analysis. X.-J.Z. performed biochemistry, qPCR, MAB-seq and CUT&Tag library building. F.G. supervised and R.Y. performed RNA-seq, WGBS and ACE-seq library building experiments. Z.-Y.S., C.G., F.Z. and Y.W. performed RNA-seq, WGBS, ACE-seq and CUT&Tag analysis. J.G. performed micromanipulation and oocyte IVM. Y.-R.D. and T.L. conducted immunostaining. Z.-M.X. and Y.W. assisted for animal housing and genotyping. W.L., C.-H.W. and H.W. performed MS analysis. Y.-R.D., G.-L.X. and X.-J.Z. wrote the paper; F.G. and C.P.W. revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Albert Jeltsch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Preparation of anti-Tet3 antibodies and validation of their specificity.
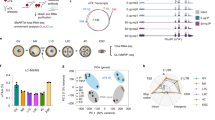
a, Locations of the Tet3 N1 (aa 1–285) and Tet3 LCD-C (aa 1294–1553, within the LCD) antigen regions in relation to the domains of three isoforms (Tet3o, Tet3FL, Tet3s). b, Western detection of overexpressed Tet3 in the whole lysate of HEK293T cells transfected with WT or Tet3 ∆LCD expression plasmid. c, Western detection of the endogenous Tet3 protein in mouse oocytes with two anti-Tet3 antibodies. The detection of β-actin served as a loading control. 100 oocytes were pooled as one sample for the analysis.
Extended Data Fig. 2 Generation of Tet3-LCD conditional knockout mouse model and their phenotypic analysis.
a, Extended DataRNA-seq tracks of the Tet3 locus in GV oocytes of WT (blue) and LCD KO (red) mice. The deleted LCD-coding region is indicated by a dashed line box. b, Confirmation of the GFP expression in MII oocytes upon Cre-mediated inversion of the loxP/loxN-flanked region. MII oocytes were obtained 20 hours after IVM. BF, bright field. Scale bar, 100 μm. c, Representative images of 5caC and 5mC immunostaining in GV oocytes. Dashed lines represent cell boundaries (from differential interference contrast view). Scale bar, 20 μm. d, Representative images of ovaries collected from adult CKO female mice and controls. Scale bar, 500 μm. e, Representative images of H&E-stained cross-sections of ovaries from adult CKO female mice and controls. Scale bar, 200 μm. f, Classification of follicles per ovarian section at different ages. Six sections per ovary from three mice per genotype were analyzed. Statistical significance (indicated P value) was analyzed using an unpaired two-tailed Student’s t-test. Data in b and c represent three independent experiments.
Extended Data Fig. 3 Transcriptomic profiling of GV oocytes with indicated genotypes.
a, Scatter plot of the differentially expressed genes and Transposon Elements (TEs) in Loop KO oocytes. The TEcount function of TE transcripts (v2.2.1)67 was applied to count TEs and genes. b, Gene ontology analysis of the DEGs in the LCD KO oocytes. A two-tailed test for log2FoldChange was performed for analysis, and the p-values were corrected for multiple testing. c, Clustering of GV oocytes according to the distinct genotypes as shown by principal component analysis on TE transcription. Each dot represents a biological replicate. Ten oocytes were pooled for the preparation of each RNA-seq library. WT, LCD KO, LCD, Dnmt3a DKO, LCD KO HD-Mut, Tet3 KO and Dnmt TKO (Dnmt1f/f, Dnmt3af/f, Dnmt3bf/f; Zp3-cre): n = 6, 7, 2, 5, 2 and 2 replicates respectively. d, Transcriptional profiles of IAPs in RNA-seq analysis (raw counts are normalized by library size). WT, LCD KO: n = 6, 7 replicates respectively. Statistical significance (indicated P value) was analyzed using an unpaired two-tailed Student’s t-test.
Extended Data Fig. 4 RT-qPCR analysis of GV oocytes with indicated genotypes.
a, RT-qPCR analysis of LTR-ERV subfamilies and representative non-LTR TEs in oocytes with the indicated genotypes. b, Pie chart illustrating the proportion of upregulated genes with an adjacent ERVK in 500 kb. c, RT-qPCR analysis of 5 representative genes upregulated in correlation with ERVK derepression in oocytes within the indicated groups. a, c, 200 oocytes were pooled together for each sample. The expression levels were normalized to 18s rRNA, and then compared to WT, which were set to 1.0. Mean and s.e.m. were determined from five biological replicates. Statistical significance (indicated P value) was analyzed using an unpaired two-tailed Student’s t-test.
Extended Data Fig. 5 Whole-genome bisulfite sequencing analysis of Tet3-LCD deleted GV oocytes.
a, Global CpG methylation levels of WT and Tet3 LCD KO GV oocytes. Methylation level was calculated using CpG sites with at least 5 × coverage in WGBS. Error bars represent ± s.e.m from 3 biological replicates. Thirty oocytes were pooled for the preparation of each WGBS library. b, Average CpG methylation levels along the gene bodies and 3-kb regions upstream and downstream of the gene body. TSS, transcription start site. TES, transcription end site. c, DMRs of WT and Tet3 LCD KO GV oocytes. At least 20% methylation level difference between LCD KO and WT samples was required to be defined as differentially hypermethylated or hypomethylated DMRs. d, Accumulative bar diagrams illustrating the proportions of the indicated genomic elements with various methylation levels.
Extended Data Fig. 6 5hmC profiling of Tet3 LCD KO GV oocytes by ACE-seq.
a, CpG methylation and hydroxymethylation in the gene bodies and 3-kb flanking regions. The profiles of combined 5mC and 5hmC determined by WGBS were included for comparison. b, Correlation between the signals of WGBS and ACE-seq data across the whole genome. Each dot is a 10-kb bin. c, A snapshot of chr 6: 84777942-85943262 locus showing modification levels determined by WGBS and ACE-seq in WT and LCD KO GV oocytes. Vertical lines denote individual CpG. d, 5mC levels determined by WGBS and 5hmC levels determined by ACE-seq of distinct IAPs in LCD KO GV oocytes. e, Heat map showing the comparison in methylation (5mC) and hydroxymethylation (5hmC) states of the indicated gICRs between WT and Tet3 LCD KO GV oocytes. 5mC levels were calculated by extracting the difference between the WGBS (5mC+5hmC) and ACE-seq (5hmC) values. The crosses (e) denote DMRs not detected by sequencing. f-g, Bisulfite (f) and ACE (g) Sanger-sequencing analyses of gICRs of the indicated maternally imprinted genes in WT and LCD KO GV oocytes. 500 oocytes were pooled for each genotype.
Extended Data Fig. 7 H3K9 methylation levels of TEs and epigenetic status of the representative upregulated genes and adjacent ERVKs.
a, Comparison of the upregulated ERVKs between LCD KO vs. WT and Setdb1 KO vs. WT GV oocytes. Out of the 17 shared subfamilies, 15 are ERVKs (highlighted in black). TEs marked in bold are those upregulated in LCD KO GV oocytes shown in Fig. 3b. The RNA-seq data of Setdb1 KO GV oocytes were from GSE82002. b, Heat maps of H3K9me3 signals (CPM) within 2-kb peak summits at LINE, SINE and ERV1 in WT, LCD KO and LCD, Dnmt3a DKO GV oocytes. Note that H3K9 methylation appears unaltered in transposon elements except for ERVK in oocytes with LCD-deleted Tet3. c-d, Tracks displaying the epigenetic status (5mC, 5hmC, and H3K9me3) of Cyp7a1 and Asb4, two representative upregulated genes, and their adjacent ERVKs in WT and LCD KO GV oocytes.
Extended Data Fig. 8 Modest upregulation of IAPs and reduction in H3K9me3 enrichment in D1+/−, D3a+/− GV oocytes.
a, Bisulfite sequencing analyses of an IAP element (accession M17551) in WT and D1+/−, D3a+/− GV oocytes. 200 oocytes were used for each genotype. b, RT-qPCR analysis of IAPs in GV oocytes with the indicated genotypes. 200 oocytes were pooled for each sample. Expression levels were normalized to 18s rRNA, and then compared to WT, which was set to 1.0. Mean and s.e.m. were determined from six biological replicates. c, No significant change in H3K9me3 level at IAPs in oocytes with Dnmt1 and Dnmt3a haploinsufficiency. Shown are H3K9me3 enrichment profiles (CPM) in a region including IAPs and their 2-kb flanking regions (upper panel), and heat maps of H3K9me3 signals (CPM) within 2-kb peak summits at IAPs in WT and D1+/−, D3a+/− GV oocytes (bottom panel). 500 GV oocytes were pooled for analysis.
Extended Data Fig. 9 Overexpression of WT TET3 has no noticeable effect on the maturation process of GV oocytes.
a, Representative images of 5hmC and 5mC immunostaining in SN GV oocytes from BDF1 female mice. The oocytes had been injected with H2O (Mock) or Tet3 mRNA. Note that the 5hmC intensity was increased significantly in GV oocytes injected with Tet3 mRNA. Scale bar, 10 μm. b, The GVBD rate of oocytes at different time points during in vitro maturation. Data are represented as mean ± s.e.m. from three independent experiments. A total of 52 (Mock) and 43 (+Tet3) GV oocytes were analyzed, respectively. c, RT-qPCR analysis of IAP expression in oocytes with the indicated treatment. 50 GV oocytes were pooled for each sample. Expression levels were normalized dto 18s rRNA, and then compared to mock, which was set to 1.0. Mean and s.e.m. were determined from six biological replicates. d, Representative images of GV oocytes stained with anti-γH2AX (green) and DAPI (blue). Scale bar, 10 μm.
Extended Data Fig. 10 Tet-mediated 5mC oxidation reactivates expression of a methylated IAP reporter in HEK293T cells.
a, Schematic of the IAPEz reporter (upper panel) and confirmation of 5mC formation on an M.SssI-treated reporter DNA fragments and 5fC/5caC formation on an M.SssI plus Tet2CD-treated reporter DNA fragments by bisulfite sequencing analysis (bottom panel). Tet2CD denotes truncated Tet2 protein with the core catalytic domain. b, Luciferase activity assay to test the effect of Tet-mediated 5mC oxidation on the reactivation of the methylated IAP reporter in transfected HEK293T cells. Mean and s.e.m. were determined from three biological replicates. Statistical significance (indicated P value) was analyzed using an unpaired two-tailed Student’s t-test.
Supplementary information
Supplementary Information
Supplementary Tables 1–5.
Source data
Source Data Figs. 1, 3, 4 and 6 and Extended Data Figs. 2–5 and 8
Single file containing all statistical source data.
Source Data Figs. 1 and 2 and Extended Data Fig. 1
Single file containing all unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, XJ., Han, BB., Shao, ZY. et al. Auto-suppression of Tet dioxygenases protects the mouse oocyte genome from oxidative demethylation. Nat Struct Mol Biol 31, 42–53 (2024). https://doi.org/10.1038/s41594-023-01125-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01125-1