Abstract
In meiosis, a supramolecular protein structure, the synaptonemal complex (SC), assembles between homologous chromosomes to facilitate their recombination. Mammalian SC formation is thought to involve hierarchical zipper-like assembly of an SYCP1 protein lattice that recruits stabilizing central element (CE) proteins as it extends. Here we combine biochemical approaches with separation-of-function mutagenesis in mice to show that, rather than stabilizing the SYCP1 lattice, the CE protein SYCE3 actively remodels this structure during synapsis. We find that SYCP1 tetramers undergo conformational change into 2:1 heterotrimers on SYCE3 binding, removing their assembly interfaces and disrupting the SYCP1 lattice. SYCE3 then establishes a new lattice by its self-assembly mimicking the role of the disrupted interface in tethering together SYCP1 dimers. SYCE3 also interacts with CE complexes SYCE1–SIX6OS1 and SYCE2–TEX12, providing a mechanism for their recruitment. Thus, SYCE3 remodels the SYCP1 lattice into a CE-binding integrated SYCP1–SYCE3 lattice to achieve long-range synapsis by a mature SC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
This study used the publicly available dataset PDB 6H86. Source data are provided with this paper.
References
Hunter, N. Meiotic recombination: the essence of heredity. Cold Spring Harb. Perspect. Biol. 7, a016618 (2015).
Zickler, D. & Kleckner, N. Recombination, pairing, and synapsis of homologs during meiosis. Cold Spring Harb. Perspect. Biol. 7, a016626 (2015).
Romanienko, P. J. & Camerini-Otero, R. D. The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol. Cell 6, 975–987 (2000).
Geisinger, A. & Benavente, R. Mutations in genes coding for synaptonemal complex proteins and their impact on human fertility. Cytogenet, Genome Res. 150, 77–85 (2016).
Fan, S. et al. Homozygous mutations in C14orf39/SIX6OS1 cause non-obstructive azoospermia and premature ovarian insufficiency in humans. Am. J. Hum. Genet 108, 324–336 (2021).
Schilit, S. L. P. et al. SYCP2 translocation-mediated dysregulation and frameshift variants cause human male infertility. Am. J. Hum. Genet 106, 41–57 (2020).
Solari, A. J. Synaptosomal complexes and associated structures in microspread human spermatocytes. Chromosoma 81, 315–337 (1980).
Cahoon, C. K. & Hawley, R. S. Regulating the construction and demolition of the synaptonemal complex. Nat. Struct. Mol. Biol. 23, 369–377 (2016).
Fraune, J., Schramm, S., Alsheimer, M. & Benavente, R. The mammalian synaptonemal complex: protein components, assembly and role in meiotic recombination. Exp. Cell Res. 318, 1340–346 (2012).
de Vries, F. A. et al. Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev. 19, 1376–1389 (2005).
Schucker, K., Holm, T., Franke, C., Sauer, M. & Benavente, R. Elucidation of synaptonemal complex organization by super-resolution imaging with isotropic resolution. Proc. Natl Acad. Sci. USA 112, 2029–2033 (2015).
Hamer, G. et al. Characterization of a novel meiosis-specific protein within the central element of the synaptonemal complex. J. Cell Sci. 119, 4025–4032 (2006).
Hamer, G. et al. Progression of meiotic recombination requires structural maturation of the central element of the synaptonemal complex. J. Cell Sci. 121, 2445–2451 (2008).
Schramm, S. et al. A novel mouse synaptonemal complex protein is essential for loading of central element proteins, recombination, and fertility. PLoS Genet. 7, e1002088 (2011).
Gomez, H. L. et al. C14ORF39/SIX6OS1 is a constituent of the synaptonemal complex and is essential for mouse fertility. Nat. Commun. 7, 13298 (2016).
Bolcun-Filas, E. et al. SYCE2 is required for synaptonemal complex assembly, double strand break repair, and homologous recombination. J. Cell Biol. 176, 741–747 (2007).
Bolcun-Filas, E. et al. Mutation of the mouse Syce1 gene disrupts synapsis and suggests a link between synaptonemal complex structural components and DNA repair. PLoS Genet. 5, e1000393 (2009).
Dunce, J. M. et al. Structural basis of meiotic chromosome synapsis through SYCP1 self-assembly. Nat. Struct. Mol. Biol. 25, 557–569 (2018).
Dunne, O. M. & Davies, O. R. Molecular structure of human synaptonemal complex protein SYCE1. Chromosoma 128, 223–236 (2019).
Dunne, O. M. & Davies, O. R. A molecular model for self-assembly of the synaptonemal complex protein SYCE3. J. Biol. Chem. 294, 9260–9275 (2019).
Sanchez-Saez, F. et al. Meiotic chromosome synapsis depends on multivalent SYCE1-SIX6OS1 interactions that are disrupted in cases of human infertility. Sci. Adv. 6, eabb1660 (2020).
Syrjanen, J. L., Pellegrini, L. & Davies, O. R. A molecular model for the role of SYCP3 in meiotic chromosome organisation. eLife 3, e02963 (2014).
Dunce, J. M., Salmon, L. J. & Davies, O. R. Structural basis of meiotic chromosome synaptic elongation through hierarchical fibrous assembly of SYCE2-TEX12. Nat. Struct. Mol. Biol. 28, 681–693 (2021).
Ollinger, R., Alsheimer, M. & Benavente, R. Mammalian protein SCP1 forms synaptonemal complex-like structures in the absence of meiotic chromosomes. Mol. Biol. Cell 16, 212–217 (2005).
Hernandez-Hernandez, A. et al. The central element of the synaptonemal complex in mice is organized as a bilayered junction structure. J. Cell Sci. 129, 2239–2249 (2016).
Lu, J. et al. Structural insight into the central element assembly of the synaptonemal complex. Sci. Rep. 4, 7059 (2014).
Teboul, L., Murray, S. A. & Nolan, P. M. Phenotyping first-generation genome editing mutants: a new standard? Mamm. Genome 28, 377–382 (2017).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Crichton, J. H. et al. Tex19.1 promotes Spo11-dependent meiotic recombination in mouse spermatocytes. PLoS Genet. 13, e1006904 (2017).
Lizatovic, R. et al. A de novo designed coiled-coil peptide with a reversible pH-induced oligomerization switch. Structure 24, 946–955 (2016).
Roder, K. & Wales, D. J. Transforming the energy landscape of a coiled-coil peptide via point mutations. J. Chem. Theory Comput. 13, 1468–1477 (2017).
Croasdale, R. et al. An undecided coiled coil: the leucine zipper of Nek2 kinase exhibits atypical conformational exchange dynamics. J. Biol. Chem. 286, 27537–27547 (2011).
Jordan, P. W., Karppinen, J. & Handel, M. A. Polo-like kinase is required for synaptonemal complex disassembly and phosphorylation in mouse spermatocytes. J. Cell Sci. 125, 5061–5072 (2012).
Spindler, M. C., Filbeck, S., Stigloher, C. & Benavente, R. Quantitative basis of meiotic chromosome synapsis analyzed by electron tomography. Sci. Rep. 9, 16102 (2019).
Libuda, D. E., Uzawa, S., Meyer, B. J. & Villeneuve, A. M. Meiotic chromosome structures constrain and respond to designation of crossover sites. Nature 502, 703–706 (2013).
Woglar, A. & Villeneuve, A. M. Dynamic architecture of DNA repair complexes and the synaptonemal complex at sites of meiotic recombination. Cell 173, 1678–1691 e16 (2018).
Pancsa, R., Schad, E., Tantos, A. & Tompa, P. Emergent functions of proteins in non-stoichiometric supramolecular assemblies. Biochim. Biophys. Acta Proteins Proteom. 1867, 970–979 (2019).
Davies, O. R., Maman, J. D. & Pellegrini, L. Structural analysis of the human SYCE2–TEX12 complex provides molecular insights into synaptonemal complex assembly. Open Biol. 2, 120099 (2012).
Keller, S. et al. High-precision isothermal titration calorimetry with automated peak-shape analysis. Anal. Chem. 84, 5066–5073 (2012).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT–a platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Brautigam, C. A., Zhao, H., Vargas, C., Keller, S. & Schuck, P. Integration and global analysis of isothermal titration calorimetry data for studying macromolecular interactions. Nat. Protoc. 11, 882–894 (2016).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Kochb, M. H. J. & Sverguna, D. I. PRIMUS—a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Crystallogr. 42, 342–346 (2009).
Quadros, R. M. et al. Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol. 18, 92 (2017).
Shen, B. et al. Efficient genome modification by CRISPR–Cas9 nickase with minimal off-target effects. Nat. Methods 11, 399–402 (2014).
Ollinger, R. et al. Deletion of the pluripotency-associated Tex19.1 gene causes activation of endogenous retroviruses and defective spermatogenesis in mice. PLoS Genet. 4, e1000199 (2008).
Soriano, P. & Jaenisch, R. Retroviruses as probes for mammalian development: allocation of cells to the somatic and germ cell lineages. Cell 46, 19–29 (1986).
Ueno, H., Turnbull, B. B. & Weissman, I. L. Two-step oligoclonal development of male germ cells. Proc. Natl Acad. Sci. USA 106, 175–180 (2009).
Costa, Y. et al. Two novel proteins recruited by synaptonemal complex protein 1 (SYCP1) are at the centre of meiosis. J. Cell Sci. 118, 2755–2762 (2005).
Acknowledgements
We thank Diamond Light Source and the staff of beamline B21 (proposals sm14435, sm15580, sm15897 and sm15836). We thank H. Waller for assistance with CD data collection, and V. A. Jatikusumo and M. Ratcliff for work in the early stages of this project. We thank A. Wheeler, M. Pearson and L. Murphy in the MRC HGU advanced imaging resource for help and guidance with imaging and image analysis, the Edinburgh Super-Resolution Imaging Consortium for super-resolution imaging and the University of Edinburgh Bioresearch and Veterinary Services for mouse husbandry. We thank W. Bickmore, J. Caceres, C. Vara and A. Marston for critically reviewing the manuscript. This study was supported by MRC University Unit grant MC_UU_00007/6 (I.R.A. and J.H.C.) and a Wellcome Senior Research Fellowship (grant number 219413/Z/19/Z) (O.R.D.).
Author information
Authors and Affiliations
Contributions
J.M.D., O.M.D. and L.J.S. performed biochemical and biophysical experiments. J.H.C. performed mouse phenotyping, developed imaging analysis pipelines and analyzed imaging. P.S.D. performed CRISPR–Cas9 injections. J.L. performed mouse genotyping. O.R.D. and I.R.A. analyzed data, designed experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Abrahan Hernandez-Hernandez and John Weir for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling editor: Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
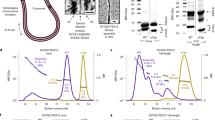
Extended Data Fig. 1 SYCP1 forms a high-affinity complex with SYCE3.
(a) Recombinant co-expression and co-purification of SYCP1αNcore-SYCE3 through amylose and anion exchange chromatography, followed by TEV cleavage to remove N-terminal expression tags, with subsequent anion exchange and size exclusion chromatography. (b) SDS-PAGE of elution fractions corresponding to the size-exclusion chromatography analysis shown in Fig. 1f. (c) Isothermal calorimetry (ITC) of SYCE3 titrated into SYCP1αNcore, demonstrating an apparent affinity of 170 ± 30 nM (mean ± SEM, n = 3 biologically independent replicates), corresponding to Fig. 1h. The injections (top), fit (middle) and residuals (bottom) are shown for the three biological replicates, with individually determined apparent affinities of 230 nM, 120 nM and 160 nM (the binding curve of the 160 nM replicate is shown in Fig. 1h). Error bars correspond to the estimated error of each integrated isotherm based on baseline uncertainty (calculated in NITPIC).
Extended Data Fig. 2 Structure of the SYCP1αNcore-SYCE3 complex.
(a) SEC-MALS analysis of MBP-SYCP1αNcore-SYCE3 (blue), MBP-SYCP1αNcore-MBP-SYCE3 (red), SYCP1αNcore-MBP-SYCE3 (yellow), revealing 2:1 complexes of 134 kDa, 163 kDa and 86 kDa, respectively (theoretical – 134 kDa, 175 kDa and 94 kDa). (b) SEC-MALS analysis of SYCP1 αCore-ΔNtip in isolation and in complex with SYCE3, demonstrating a 302 kDa tetramer and 195 kDa 2:1 complex, respectively (theoretical – 320 kDa and 171 kDa). (c) Far UV CD spectra and (d) CD thermal denaturation of SYCP1αNcore-SYCE3 (purple) and SYCP1αNcore (blue). (c) Secondary structure composition was estimated through deconvolution of spectra with data fitted at normalised rms deviation values of 0.006 and 0.001, respectively. (d) Thermal denaturation recorded for SYCP1αNcore-SYCE3 and SYCP1αNcore as % unfolded based on the helical signal at 222 nm; melting temperatures were estimated at 38 °C and 37 °C, respectively. (e–g) SEC-SAXS analysis. (e) Scattering intensity plots, (f) Guinier analysis to determine the radius of gyration (Rg) with linear fits shown in black (Q.Rg values were < 1.3) and (g) Guinier analysis to determine the radius of gyration of the cross-section (Rc) (Q.Rc values were < 1.3) for SYCP1αNcore-SYCE3 and SYCP1αNcore. Corresponding P(r) distributions and ab initio models are shown in Fig. 2b. (h) SYCE3-binding analysis through co-expression with MBP-SYCP1 or free MBP and co-purification by amylose, ion exchange and size-exclusion chromatography using SYCP1αNcore and SYCE3 truncations.
Extended Data Fig. 3 SYCP1 binds to the SYCE3 WY mutant.
(a) Schematic of the SYCE3 chain, dimeric structure and self-assembly into tetramers and higher-order structures. SYCE3 consists of two α-helices, α1 (blue) and α2 (red), which are linked together by the P53 loop. In the SYCE3 dimer, the P53 loop adopts a closed conformation. This structure self-assembles through one of its P53 loops opening, creating a tetramer consisting of two linear chains and two helix-loop-helix chains. SYCE3 tetramer formation is blocked by the PPP-loop mutation which supports the dimeric closed loop conformation, but is incompatible with the assembled open loop conformation. Similarly, SYCE3 constitutively assembles upon P53Q mutation, which is incompatible with the closed loop conformation. SYCE3 tetramers undergo higher-order assembly through lateral interaction of their W41 Y44 sites. The resultant higher-order structures are held together by the combined actions of the end-on interface of the tetramer and the lateral interfaces mediated by W41 and Y44. Higher-order assembly through lateral interactions is blocked by the W41E Y44E (WY) mutation. (b) ITC analysis of SYCE3 WY titrated into SYCP1αNcore, demonstrating an apparent affinity of 16 ± 3 nM (mean ± SEM, n = 3 biologically independent replicates), corresponding to Fig. 2g. The injections (top), fit (middle) and residuals (bottom) are shown for the three biological replicates, with individually determined apparent affinities of 16 nM, 10 nM and 22 nM (the binding curve of the 16 nM replicate is shown in Fig. 2g). (c) ITC of SYCE3 PPP-loop titrated into SYCP1αNcore, in which no interaction was not observed and the binding affinity was not determined (n.d.). (b, c) Error bars correspond to the estimated error of each integrated isotherm based on baseline uncertainty (calculated in NITPIC).
Extended Data Fig. 4 Structure of the SYCP1αN-SYCE3 complex.
(a) SDS-PAGE of size-exclusion chromatography elution fractions of 127 μM SYCP1αN upon incubation with SYCE3 at stoichiometric ratios (per molecule) of 1:0.5, 1:5 and 1:10; free SYCP1αN and SYCE3 are shown for comparison. (b) SEC-MALS analysis (using a Superose 6 increase 10/300 GL column) of MBP-SYCP1αN-His-SYCE3 revealing 2:1 and 4:2 species of 164 kDa and 335 kDa, respectively (theoretical – 160 kDa and 319 kDa). (c,d) SEC-SAXS analysis. (c) Scattering intensity plots and (d) Guinier analysis to determine the radius of gyration (Rg) with linear fits shown in black (Q.Rg values were < 1.3) for SYCP1αN-SYCE3 2:1 and 4:2 complexes. Corresponding P(r) distributions and ab initio models are shown in Fig. 3c, d.
Extended Data Fig. 5 SYCP1-SYCE3 integrated lattice formation through SYCE3 self-assembly.
(a) Size-exclusion chromatography of (a, b) 95 μM SYCP1αN-SYCE3 and (c) 95 μM SYCP1αNcore-SYCE3 upon incubation with a 10-fold stoichiometric excess (per molecule) of SYCE3 wild-type or WY, corresponding to Fig. 4f–h. (a, c) UV absorbance (280 nm) chromatograms normalised to the same maximum peak height shown in Fig. 3f, g with additional chromatograms for free SYCE3 wild-type and WY.
Extended Data Fig. 6 Syce3 mutant allele sequences and meiotic phenotypes.
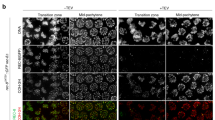
(a) Syce3 nucleotide and predicted protein sequences of mice used in this study. Mouse IDs are shown on the left and the genotype group on the right. Animal 5959 in the Syce3WY/WY group did not incorporate the silent C:A mutation in PAM1 and was mosaic/heterozygous for the silent G:C mutation in PAM2. For the Syce3Δ/Δ group the number of nucleotides deleted is indicated in the allele name. The PAM sequences are highlighted with blue boxes, the W41 and Y44 sequences with orange boxes. (b) Chromatograms showing examples of sequencing the Syce3 locus from mice with the indicated genotypes and IDs. A black arrow indicates nucleotides deleted at this position. Mice that lacked potential mosaicism/heterogeneity at the W41/Y44 codons were used in this study. (c) Percentage of SYCP3-positive spermatocytes at the indicated stage of meiosis I in Syce3 chromosome spreads based on SYCP3 and SYCP1 immunostaining (Fig. 4f). Mean percentages for each genotype are indicated by the bars, percentages from individual animals are indicated by filled circles. Asterisks indicate a significant difference (p < 0.05, Student’s t-test, n = 3) relative to Syce3PAM/PAM controls.
Extended Data Fig. 7 Quantitative analysis of SYCP1 foci in Syce3 spermatocytes.
(a) SIM images of pachytene Syce3PAM/PAM meiotic chromosome spreads immunostained for SYCP3 (magenta) and SYCP1 (green). Scale bars, 10 µm for low magnification images, 1 µm for enlarged regions. Patches where SYCP1 extends linearly along one axis are indicated with arrows. The spread shown is the same as in Fig. 5a (b) SYCP1 foci-SYCE3 axis mask distances. Distances from the centroid of each SYCP1 focus to the nearest point on the SYCP3 axis mask are shown alongside distances from shuffled datasets obtained by assigning all SYCP1 foci in each nucleus to a random nuclear location for twenty iterations. The red dotted horizontal line represents the 35 nm threshold distinguishing axial and non-axial foci. Crossbars represent quartiles; *, p < 0.01 (Mann-Whitney U test, paired test used to compare observed with shuffled datasets, nuclei medians are 16, 603, 91 and 494 nm, n = 25, 26 nuclei); 3 animals analysed for each Syce3 genotype. (c) Total SYCP1 signal in each axial SYCP1 focus as shown in Fig. 5d, with data segmented for individual animals. Crossbars represent quartiles; medians are 152975, 116615, 101019, 8067, 24188 and 1855 arbitrary units); mouse IDs are shown below each dataset. (d) Violin plots showing the anti-SYCP1 immunostaining signal per focus, intensity within foci and focus area for axial and non-axial SYCP1 foci in Syce3Δ/Δ and Syce3WY/WY spermatocytes. Crossbars represent quartiles. Median SYCP1 signals per focus: 60101, 123441, 9061 and 8485 arbitrary units. Median SYCP1 intensities within foci; 2205, 2525, 255, and 203 arbitrary units per px2. Median areas; 29, 48, 48 and 53 nm2. 3 animals analysed for each Syce3 genotype. (e) Non-axial SYCP1 foci frequencies in asynapsed pachytene Syce3Δ/Δ and Syce3WY/WY spermatocytes. *, p < 0.05 (Mann-Whitney U test, medians are 86 and 66 foci, n = 25, 26 nuclei); 3 animals analysed for each Syce3 genotype.
Extended Data Fig. 8 SYCE3 interacts with SYCE1-SIX6OS1 and SYCE2-TEX12 complexes.
(a) Size-exclusion chromatography of co-expressed and co-purified SYCE1core-SYCE3, with UV chromatograms of SYCE1core-SYCE3 (blue), SYCE1core (yellow) and SYCE3 (red), corresponding to Fig. 6e. (b) Size-exclusion chromatography of SYCE2-TEX12 core (green), SYCE3 (red) and an equimolar mixture of SYCE2-TEX12 core and SYCE3 (blue), shown as UV absorbance (280 nm) and SDS-PAGE of elution fractions. (c-i) MST analysis of SYCE3 titrated into 150 nM SYCE2-TEX12 core, corresponding to Fig. 6 f. (c) Full dataset in which the final two datapoints were excluded from analysis. The apparent binding affinity was determined to be 21.8 ± 2.1 μM (mean ± SEM,n = 3 biologically independent replicates). (d) Overlaid capillary scans. (e,f) Initial fluorescence for three data series (blue, yellow, green) displayed as (e) individual data series and (f) data represented as mean ± SEM (n = 3 biologically independent replicates). (g–i) Relative fluorescence for each of the three data series.
Extended Data Fig. 9 Electron microscopy analysis of SYCE2-TEX12 following incubation with a two-fold excess of SYCE3.
Full panels corresponding to Fig. 6 g. Scale bar, 200 nm.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed gels.
Source Data Fig. 3
Unprocessed gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed gels.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 4
Unprocessed gels.
Source Data Extended Data Fig. 5
Unprocessed gels.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Crichton, J.H., Dunce, J.M., Dunne, O.M. et al. Structural maturation of SYCP1-mediated meiotic chromosome synapsis by SYCE3. Nat Struct Mol Biol 30, 188–199 (2023). https://doi.org/10.1038/s41594-022-00909-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00909-1
This article is cited by
-
SCEP1 and SCEP2 are two new components of the synaptonemal complex central element
Nature Plants (2023)
-
The plant synaptonemal complex: identification of the first central element components
Nature Plants (2023)