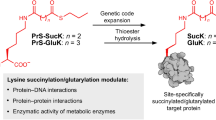
Abstract
Protein post-translational modification (PTM) regulates nearly every aspect of cellular processes in eukaryotes. However, the identification of new protein PTMs is very challenging. Here, using genetically encoded unnatural amino acids as chemical probes, we report the identification and validation of a previously unreported form of protein PTM, aminoacylated lysine ubiquitination, in which the modification occurs on the α-amine group of aminoacylated lysine. We identify more than 2,000 ubiquitination sites on all 20 aminoacylated lysines in two human cell lines. The modifications can mediate rapid protein degradation, complementing the canonical lysine ubiquitination-mediated proteome degradation. Furthermore, we demonstrate that the ubiquitin-conjugating enzyme UBE2W acts as a writer of aminoacylated lysine ubiquitination and facilitates the ubiquitination event on proteins. More broadly, the discovery and validation of aminoacylated lysine ubiquitination paves the way for the identification and verification of new protein PTMs with the genetic code expansion strategy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Any Supplementary Information (methods, figures, DNA sequences and protein sequences) and chemical compound information is available in the online version of the paper. The MS data have been deposited to the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD035166. UbiSite enrichment MS/MS raw files were obtained from PXD006201 (ref. 21) deposited to the ProteomeXchange Consortium36. Correspondence and requests for materials should be addressed to S.L. Source data are provided with this paper.
Code availability
The source codes for R and Python used in this paper have been deposited in GitHub (https://github.com/yuriychen/2022_KXUb_UbiSite_Codes).
References
Ciechanover, A. & Ben-Saadon, R. N-terminal ubiquitination: more protein substrates join in. Trends Cell. Biol. 14, 103–106 (2004).
Jiang, H. et al. Protein lipidation: occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 118, 43–112 (2018).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018).
Luo, M. Chemical and biochemical perspectives of protein lysine methylation. Chem. Rev. 118, 6656–6705 (2018).
He, X.-D. et al. Sensing and transmitting intracellular amino acid signals through reversible lysine aminoacylations. Cell Metab. 27, 151–166 (2018).
Giglione, C., Fieulaine, S. & Meinnel, T. N-terminal protein modifications: bringing back into play the ribosome. Biochimie 114, 134–146 (2015).
Varland, S., Osberg, C. & Arnesen, T. N-terminal modifications of cellular proteins: the enzymes involved, their substrate specificities and biological effects. Proteomics 15, 2385–2401 (2015).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Park, H.-S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Yang, M., Li, J. & Chen, P. R. Transition metal-mediated bioorthogonal protein chemistry in living cells. Chem. Soc. Rev. 43, 6511–6526 (2014).
Lang, K. & Chin, J. W. Cellular incorporation of unnatural amino acids and bioorthogonal labeling of proteins. Chem. Rev. 114, 4764–4806 (2014).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Krauskopf, K. & Lang, K. Increasing the chemical space of proteins in living cells via genetic code expansion. Curr. Opin. Chem. Biol. 58, 112–120 (2020).
Wang, J. et al. Palladium-triggered chemical rescue of intracellular proteins via genetically encoded allene-caged tyrosine. J. Am. Chem. Soc. 138, 15118–15121 (2016).
Elliott, T. S. et al. Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat. Biotechnol. 32, 465–472 (2014).
Elliott, T. S., Bianco, A., Townsley, F. M., Fried, S. D. & Chin, J. W. Tagging and enriching proteins enables cell-specific proteomics. Cell. Chem. Biol. 23, 805–815 (2016).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21, 921–926 (2003).
Xu, G., Paige, J. S. & Jaffrey, S. R. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat. Biotechnol. 28, 868–873 (2010).
Akimov, V. et al. UbiSite approach for comprehensive mapping of lysine and N-terminal ubiquitination sites. Nat. Struct. Mol. Biol. 25, 631–640 (2018).
Lide, D. R. Handbook of Chemistry and Physics, 83rd ed (CRC, 2002).
Bae, S. et al. Akt is negatively regulated by the MULAN E3 ligase. Cell Res. 22, 873–885 (2012).
Nguyen, D. P. et al. Genetic encoding and labeling of aliphatic azides and alkynes in recombinant proteins via a pyrrolysyl-tRNA synthetase/tRNACUA pair and click chemistry. J. Am. Chem. Soc. 131, 8720–8721 (2009).
Kolb, H. C., Finn, M. G. & Sharpless, K. B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. 40, 2004–2021 (2001).
Stewart, M. D., Ritterhoff, T., Klevit, R. E. & Brzovic, P. S. E2 enzymes: more than just middle men. Cell Res. 26, 423–440 (2016).
Scaglione, K. M. et al. The ubiquitin-conjugating enzyme (E2) Ube2w ubiquitinates the N terminus of substrates. J. Biol. Chem. 288, 18784–18788 (2013).
Tatham, M. H., Plechanovova, A., Jaffray, E. G., Salmen, H. & Hay, R. T. Ube2W conjugates ubiquitin to alpha-amino groups of protein N-termini. Biochem. J. 453, 137–145 (2013).
Vittal, V. et al. Intrinsic disorder drives N-terminal ubiquitination by Ube2w. Nat. Chem. Biol. 11, 83–89 (2015).
Kravtsova-Ivantsiv, Y. & Ciechanover, A. Non-canonical ubiquitin-based signals for proteasomal degradation. J. Cell Sci. 125, 539–548 (2012).
McClellan, A. J., Laugesen, S. H. & Ellgaard, L. Cellular functions and molecular mechanisms of non-lysine ubiquitination. Open Biol. 9, 190147 (2019).
Otten, E. G. et al. Ubiquitylation of lipopolysaccharide by RNF213 during bacterial infection. Nature 594, 111–116 (2021).
Ding, W. et al. Chimeric design of pyrrolysyl-tRNA synthetase/tRNA pairs and canonical synthetase/tRNA pairs for genetic code expansion. Nat. Commun. 11, 3154 (2020).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253–1260 (2017).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Vizcaíno, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Giai Gianetto, Q., Coute, Y., Bruley, C. & Burger, T. Uses and misuses of the fudge factor in quantitative discovery proteomics. Proteomics 16, 1955–1960 (2016).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Luo, W., Pant, G., Bhavnasi, Y. K., Blanchard, S. G. & Brouwer, C. Pathview Web: user friendly pathway visualization and data integration. Nucleic Acids Res. 45, W501–W508 (2017).
Luo, W. & Brouwer, C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinf. 10, 161 (2009).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Schneider, T. D. & Stephens, R. M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Wang, M., Herrmann, C. J., Simonovic, M., Szklarczyk, D. & von Mering, C. Version 4.0 of PaxDb: protein abundance data, integrated across model organisms, tissues, and cell-lines. Proteomics 15, 3163–3168 (2015).
Wang, L. H. et al. pFind 2.0: a software package for peptide and protein identification via tandem mass spectrometry. Rapid Commun. Mass Spectrom. 21, 2985–2991 (2007).
Carvalho, A. F. et al. High-yield expression in Escherichia coli and purification of mouse ubiquitin-activating enzyme E1. Mol. Biotechnol. 51, 254–261 (2012).
Wenzel, D. M., Lissounov, A., Brzovic, P. S. & Klevit, R. E. UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 474, 105–108 (2011).
Li, X., Fekner, T. & Chan, M. K. N6-(2-(R)-propargylglycyl)lysine as a clickable pyrrolysine mimic. Chem. Asian J. 5, 1765–1769 (2010).
Acknowledgements
We thank the National Key R&D Program of China (grant no. 2019YFA09006600), the National Natural Science Foundation of China (grant nos. 91953113, 21877096, 22222705) and the Fundamental Research Funds for the Zhejiang Provincial Universities (grant no. 2019XZZX003-19) for financial support. We thank L. Liu and X. Zhang from the core facility of Life Sciences Institute and H. Song for sharing stable knockdown plasmids. We thank X. He, B. Yang, X. Guo and V. Y. Yu for helpful discussions.
Author information
Authors and Affiliations
Contributions
S.L. conceived the idea and supervised the study. J.Z. and Y.C. designed and conducted most experiments, and analyzed the data together. C.L. assisted with compound preparation. L.H. helped with sample preparation. H.Z., W.D. and Y.Y. provided technical advice. S.L., J.Z. and Y.C. wrote the manuscript. All authors commented on the final draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Jeff Ranish and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Florian Ullrich, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The PylRS mutation table.
The amino acids at the indicated position of WT-PylRS are shown. And the mutated amino acids in the PylRS variants are highlighted by red.
Extended Data Fig. 2 Screening and verification of PylRS variants for UAA recognition.
a, The schematic illustration of the screening process. The GFP-190TAG gene was used as the reporter gene in the assay. And the amber suppression efficiency of the PylRS variants showed in Extended Data Fig. 1 was assayed in the presence or absence of UAA. b, Screening of PylRS variants for MetK recognition. The MetKRS (variant 5) used in this study is highlighted in red. c, Screening of PylRS variants PraK recognition. The potential candidates (variant 4 and 41 for PraK) are shown in bold. d, Assessment of amber suppression efficiency of variants 4 and 41 for PraK recognition. The PraKRS used in this study is highlighted in red. e, The amino acid mutations of MetKRS and PraKRS used in this study. f, Analysis of amber suppression efficiency of MetKRS and PraKRS in the presence or absence of the corresponding UAAs in E. coli. Mass spectrometry characterization of the fidelity of MetK (g) and PraK (h) incorporation on GFP. i, the FACS gating method in Fig. 2d sorts the EGFP+ mCherry+ population of HEK293T cells. FSC and SSC are used to exclude cell debris. Cells expressing EGFP or mCherry fluorescent proteins are used to set the gain and determine the threshold for positive and negative cells in each channel. For d and f, data are presented as mean values ± SEM (n = 3 biologically independent experiments). Source data are provided as a Source Data file.
Extended Data Fig. 3 The representative spectra of K-XUb modification.
a, A representative spectrum of K-MUb on K98 of PSB3 protein. Inset: enlarged signal for the characteristic fragment (GGM). b, A representative spectrum of K-oxMUb on K1214 of TOP2B protein. Inset: enlarged signal for the characteristic fragment (GGoxM, oxM refers to the oxidized form of Met). c, The spectrum of synthesized peptide with a branching GGM modification on lysine. Inset: enlarged signal for the characteristic fragment (GGM). d, The spectrum of synthesized peptide without a branching GGM modification on lysine. Inset: enlarged signal for the non-existence of characteristic fragment (GGM). e, A representative spectrum of K-XleUb on K338 of TBA1A protein in UbiSite enrichment. Inset: enlarged signal for the characteristic fragment (GGXle, Xle refers to the Leu or Ile). f, A spectrum of K-NUb on K49 of 14-3-3Z protein in UbiSite enrichment. Inset: enlarged signal for the characteristic fragment (GGN).
Extended Data Fig. 4 Analysis of K-XUb modification enriched by the UbiSite antibody.
a, Consensus amino acid frequencies surrounding the aminoacylated lysine (K-XUb) and canonical lysine (K-Ub) ubiquitination sites containing characteristic fragment. b, Multidimensional scaling (MDS) plot of samples. Different replicates of the same cell line and treatment were in the same color. c, Enrichment analysis of enriched proteins in Jurkat cells treated with vehicle (left columns colored in green) or inhibitor (Bort or AP15) (right columns colored in red). d, Enrichment analysis of inhibitors enriched proteins in Hep2 (left columns colored in red) and Jurkat cells (right columns colored in blue). e, Overlap of inhibitors enrichment sites and proteins in Hep2 and Jurkat cell lines, detailed figures are shown in Supplementary Data 3. f, Circos plot of overlapping ubiquitinated proteins that carried both aminoacylated lysine and lysine ubiquitination (Same as Ub) or carried only aminoacylated lysine ubiquitination (K-XUb only). Blue lines link proteins in the same gene set analyzed by GO enrichment. g, Boxplot of protein abundance distribution of ubiquitinated proteins that carried both aminoacylated lysine and lysine ubiquitination (Same as Ub) or carried only aminoacylated lysine ubiquitination (K-XUb only). Protein abundance data were acquired from PAXdb. P-value was calculated by two-sided t-test (n = 990 for Same as Ub proteins and n = 173 K-XUb only proteins). Boxplot bounds depict quartile 1, median and quartile 3, with whiskers at 1.5× interquartile range and outlier points. h, The type and number of K-XUb modification distributed in each protein. For c and d, p-values are calculated based on the cumulative hypergeometric distribution by Metascape.
Extended Data Fig. 5 Pathway analysis of proteins with enriched K-XUb modification.
K-XUb modified proteins that enrich in the inhibitor-treated group (red), the vehicle-treated group (green), and both groups (blue) are involved in cell cycle (a), and protein processing in endoplasmic reticulum pathway (b).
Extended Data Fig. 6 Western blot analysis of turnover rates and ubiquitination of identified aminoacylated lysine ubiquitinated proteins.
a, The turnover rates of proteins with lysine aminoacylation incorporated at the indicated position were analyzed by western blot with antibody against Myc-tag. Actin was used as loading control. Results are representative of three independent experiments with similar results. b, The level of polyubiquitination of proteins with lysine aminoacylation-incorporated at the indicated position were analyzed, using their corresponding WT proteins as controls. Protein samples were enriched by Ni-NTA resin and analyzed by western blot with antibodies against Flag-tag (Flag-Ub) and Myc-tag (substrate protein). Western blot analysis of anti-Myc and anti-Actin in whole cell lysates was performed as controls. The experiment in the figure was repeated twice with similar results. Source data are provided as a Source Data file.
Extended Data Fig. 7 The degradation of individual protein and whole proteome in the presence or absence of E3 ligase or proteasomal inhibitors.
a, A representative western blot of the indicated AKT1 proteins in the presence or absence of E3 ligase. The quantification analysis was shown in Fig. 5d in the main text. The experiment in the figure was repeated twice with similar results. b, Residue-specific incorporation of UAAs via SORT in mammalian cells. The HEK293T proteomes were labelled with AlkK or PraK in a residue-specific manner via SORT. The successful incorporation of UAAs was verified by the azide-PEG3-biotin probe through click reaction and visualized with streptavidin (SA)-HRP blot. The Coomassie brilliant blue staining gel is provided as the loading control. The experiment in the figure was repeated twice with similar results. c, Dot blot analysis of the proteasomal inhibitors rescue of the proteome labelled with PraK using SORT. MG132, MG132 treatment; Bort, bortezomib treatment. Three biological replicates (Re) were performed. And the normalized data was shown in (d). Data are presented as mean values ± SEM. The experiment in the figure was repeated twice with similar results. Source data are provided as a Source Data file.
Extended Data Fig. 8 In vitro nucleophile reactivity screening.
All Ub-transferred E2s were purified and the reactivity of E2~Ubs towards buffer (-), K, PraK and AsnK was determined. Reactions were quenched by a non-reducing loading buffer. Reactivity of E2~Ub with lysine or aminoacylated lysines is indicated by the decrease of E2~Ub conjugates and the increase of free E2 and free Ub. E2~Ub and free E2 are indicated by the red and black triangles, respectively. Free Ub is highlighted by the dashed boxes. The experiment in the figure was repeated twice with similar results. Source data are provided as a Source Data file.
Extended Data Fig. 9 Liquid chromatography–mass spectrometry (LC–MS) analysis of the products in nucleophile reactivity screening.
a, LC–MS analysis of the results of incubation of E2~Ub with AsnK in Supplementary Fig. 11. Freshly purified Ub protein has two MS peaks, Ub (Ubiquitin lacking N terminal Met) and Ub* (acetylated Ub carrying N terminal Met). The desired products are indicated by arrows. And the unwanted products (or the side products) were detected and indicated by triangles in UBE2J2 and UBE2R1. b, LC–MS analysis of the results of incubation of UBE2R1~Ub with AsnK in the presence or absence of ATP. The side products were generated by reaction with ATP instead of AsnK. c, LC–MS analysis of the results of incubation of UBE2W~Ub with the indicated aminoacylated lysine individually or simultaneously. In the figure, the pKa value of AsnK is lower than that of GlnK. And the pKa value of MetK is lower than that of ThrK.
Extended Data Fig. 10 UBE2W promotes the ubiquitination of PraK-incorporated proteome.
a, The endogenous UBE2W expression level in several commonly used cell lines. b, shRNAs sequence information used in this study and western blot analysis of knockdown efficiency of UBE2W in HEK293T cells. Tubulin was used as a loading control. HEK293T-shUBE2W-2 cell line was used for the following study. c, The ubiquitination of PraK-labelled proteomes of WT cells and UBE2W knockdown cells generated in the presence of proteasomal inhibitor bortezomib. The proteome was labelled with PraK via SORT, conjugated with biotin, enriched by streptavidin (SA) resin and subsequently analyzed by western blot with antibodies against biotin and ubiquitin. Western blot analysis of anti-Actin and anti-UBE2W in whole cell lysate was performed as controls. All experiments were repeated twice with similar results. Source data are provided as a Source Data file.
Supplementary information
Supplementary Information
Supplementary sequences 1–3.
Supplementary Data 1
Summary of K-XUb modified sites and peptides in the SORT-labeled proteome.
Supplementary Data 2
Summary of K-XUb profiling results with the UbiSite antibody.
Supplementary Data 3
Summary of K-XUb quantification results with the UbiSite antibody.
Supplementary Data 4
Summary of K-Ub profiling results with the UbiSite antibody.
Source data
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots and dot blot.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Unprocessed western blots, dot blot and SDS–PAGE.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed SDS–PAGE.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zang, J., Chen, Y., Liu, C. et al. Genetic code expansion reveals aminoacylated lysine ubiquitination mediated by UBE2W. Nat Struct Mol Biol 30, 62–71 (2023). https://doi.org/10.1038/s41594-022-00866-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00866-9
This article is cited by
-
Trim-Away ubiquitinates and degrades lysine-less and N-terminally acetylated substrates
Nature Communications (2023)