Abstract
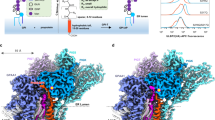
Glycosylphosphatidylinositol (GPI) molecules are complex glycophospholipids and serve as membrane anchors for tethering many proteins to the cell surface. Attaching GPI to the protein in the endoplasmic reticulum (ER) is catalyzed by the transmembrane GPI transamidase (GPIT) complex, which is essential for maturation of the GPI-anchored proteins. The GPIT complex is known to be composed of five subunits: PIGK, PIGU, PIGT, PIGS and GPAA1. Here, we determined the structure of the human GPIT complex at a resolution of 3.1 Å using single-particle cryo-EM, elucidating its overall assembly. The PIGK subunit functions as the catalytic component, in which we identified a C206-H164-N58 triad that is critical for the transamination reaction. Transmembrane helices constitute a widely opened cleft, which is located underneath PIGK, serving as a GPI substrate-binding site. The ubiquitin E3 ligase RNF121 is visualized at the back of the complex and probably serves as a quality control factor for the GPIT complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The three-dimensional cryo-EM density maps of the GPIT and GPIT-RNF121 complexes have been deposited in the EM Database under accession codes EMD-32336 and EMD-32452, respectively, and the coordinates of the GPIT complex have been deposited in the Protein Data Bank under accession code 7W72. Source data are provided with this paper.
References
Ikezawa, H. Glycosylphosphatidylinositol (GPI)-anchored proteins. Biol. Pharm. Bull. 25, 409–417 (2002).
Orlean, P. & Menon, A. K. Thematic review series: lipid posttranslational modifications. GPI anchoring of protein in yeast and mammalian cells, or: how we learned to stop worrying and love glycophospholipids. J. Lipid Res. 48, 993–1011 (2007).
Pittet, M. & Conzelmann, A. Biosynthesis and function of GPI proteins in the yeast Saccharomyces cerevisiae. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1771, 405–420 (2007).
Borner, G. H., Lilley, K. S., Stevens, T. J. & Dupree, P. Identification of glycosylphosphatidylinositol-anchored proteins in Arabidopsis. A proteomic and genomic analysis. Plant Physiol. 132, 568–577 (2003).
Kinoshita, T., Inoue, N. & Takeda, J. Defective glycosyl phosphatidylinositol anchor synthesis and paroxysmal nocturnal hemoglobinuria. Adv. Immunol. 60, 57–103 (1995).
Kinoshita, T., Fujita, M. & Maeda, Y. Biosynthesis, remodelling and functions of mammalian GPI-anchored proteins: recent progress. J. Biochem. 144, 287–294 (2008).
Kinoshita, T. Biosynthesis and biology of mammalian GPI-anchored proteins. Open Biol. 10, 190290 (2020).
Yu, J. et al. The affected gene underlying the class K glycosylphosphatidylinositol (GPI) surface protein defect codes for the GPI transamidase. Proc. Natl Acad. Sci. USA 94, 12580–12585 (1997).
Hiroi, Y. et al. Molecular cloning of human homolog of yeast GAA1 which is required for attachment of glycosylphosphatidylinositols to proteins 1. FEBS Lett. 421, 252–258 (1998).
Ohishi, K., Inoue, N. & Kinoshita, T. PIG-S and PIG-T, essential for GPI anchor attachment to proteins, form a complex with GAA1 and GPI8. EMBO J. 20, 4088–4098 (2001).
Hong, Y. et al. Human PIG-U and yeast Cdc91p are the fifth subunit of GPI transamidase that attaches GPI-anchors to proteins. Mol. Biol. Cell 14, 1780–1789 (2003).
Nguyen, T. T. M. et al. Mutations in GPAA1, encoding a GPI transamidase complex protein, cause developmental delay, epilepsy, cerebellar atrophy, and osteopenia. Am. J. Hum. Genet. 101, 856–865 (2017).
Nakashima, M. et al. Novel compound heterozygous PIGT mutations caused multiple congenital anomalies-hypotonia-seizures syndrome 3. Neurogenetics 15, 193–200 (2014).
Nguyen, T. T. M. et al. Mutations in PIGS, encoding a GPI transamidase, cause a neurological syndrome ranging from fetal akinesia to epileptic encephalopathy. Am. J. Hum. Genet. 103, 602–611 (2018).
Knaus, A. et al. Mutations in PIGU impair the function of the GPI transamidase complex, causing severe intellectual disability, epilepsy, and brain anomalies. Am. J. Hum. Genet. 105, 395–402 (2019).
Nguyen, T. T. M. et al. Bi-allelic variants in the GPI transamidase subunit PIGK cause a neurodevelopmental syndrome with hypotonia, cerebellar atrophy, and epilepsy. Am. J. Hum. Genet. 106, 484–495 (2020).
Chen, X. et al. Loss of PIGK function causes severe infantile encephalopathy and extensive neuronal apoptosis. Hum. Genet. 140, 791–803 (2021).
Meyer, U., Benghezal, M., Imhof, I. & Conzelmann, A. Active site determination of Gpi8p, a caspase-related enzyme required for glycosylphosphatidylinositol anchor addition to proteins. Biochemistry 39, 3461–3471 (2000).
Holm, L. & Rosenström, P. I. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Bloch, J. S. et al. Structure and mechanism of the ER-based glucosyltransferase ALG6. Nature 579, 443–447 (2020).
Eisenhaber, B., Maurer-Stroh, S., Novatchkova, M., Schneider, G. & Eisenhaber, F. Enzymes and auxiliary factors for GPI lipid anchor biosynthesis and post‐translational transfer to proteins. Bioessays 25, 367–385 (2003).
Ohishi, K., Nagamune, K., Maeda, Y. & Kinoshita, T. Two subunits of glycosylphosphatidylinositol transamidase, GPI8 and PIG-T, form a functionally important intermolecular disulfide bridge. J. Biol. Chem. 278, 13959–13967 (2003).
Yi, L. et al. Disulfide bond formation and N-glycosylation modulate protein–protein interactions in GPI-transamidase (GPIT). Sci. Rep. 7, 45912 (2017).
Kvarnung, M. et al. A novel intellectual disability syndrome caused by GPI anchor deficiency due to homozygous mutations in PIGT. J. Med. Genet. 50, 521–528 (2013).
Yang, L. et al. Homozygous PIGT mutation lead to multiple congenital anomalies-hypotonia seizures syndrome 3. Front. Genet. https://doi.org/10.3389/fgene.2018.00153 (2018).
Ohishi, K. et al. Gaa1p and gpi8p are components of a glycosylphosphatidylinositol (GPI) transamidase that mediates attachment of GPI to proteins. Mol. Biol. Cell 11, 1523–1533 (2000).
Benghezal, M., Benachour, A., Rusconi, S., Aebi, M. & Conzelmann, A. Yeast Gpi8p is essential for GPI anchor attachment onto proteins. EMBO J. 15, 6575–6583 (1996).
Udenfriend, S. & Kodukula, K. How glycosylphosphatidylinositol-anchored membrane proteins are made. Annu. Rev. Biochem. 64, 563–591 (1995).
Dall, E. & Brandstetter, H. Mechanistic and structural studies on legumain explain its zymogenicity, distinct activation pathways, and regulation. Proc. Natl Acad. Sci. USA 110, 10940–10945 (2013).
Ellis, M., Sharma, D. K., Hilley, J. D., Coombs, G. H. & Mottram, J. C. Processing and trafficking of Leishmania mexicana GP63 analysis using gpi8 mutants deficient in glycosylphosphatidylinositol protein anchoring. J. Biol. Chem. 277, 27968–27974 (2002).
Legler, D. F. et al. Differential insertion of GPI-anchored GFPs into lipid rafts of live cells. FASEB J. 19, 73–75 (2005).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Darom, A., Bening-Abu-Shach, U. & Broday, L. RNF-121 is an endoplasmic reticulum-membrane E3 ubiquitin ligase involved in the regulation of β-integrin. Mol. Biol. Cell 21, 1788–1798 (2010).
Zavodszky, E., Peak-Chew, S.-Y., Juszkiewicz, S., Narvaez, A. J. & Hegde, R. S. Identification of a quality-control factor that monitors failures during proteasome assembly. Science 373, 998–1004 (2021).
Ferguson, M. A., Kinoshita, T. & Hart, G. W. in Essentials of Glycobiology 2nd edn (eds Varki, A. et al.) Ch.11 (Cold Spring Harbor Laboratory Press, 2009).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2858 (2014).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
DeLano, W. L. PyMOL: an open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 40, 82–92 (2002).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Acknowledgements
We thank B. Zhu, J. Xiong, Z. Liu and S. Zhou from the Institute of Biophysics, Chinese Academy of Science, for helpful discussion about the PIGK knockout experiment. We thank M. Dong from the National Institute of Biological Science, Beijing, and M. Zhou from Nanjing University of Science and Technology, China, for helpful discussion about the mass spectrometry experiment. We thank X. Huang, B. Zhu, X. Li and other staff members at the Center for Biological Imaging (CBI), Core Facilities for Protein Science at the Institute of Biophysics, Chinese Academy of Science (IBP, CAS) for support in cryo-EM data collection. We thank J. Jia and S. Meng (Core Facility of Institute of Biophysics, Chinese Academy of Sciences) for flow cytometric analysis. We thank X. Ding and M. Zhang from the Laboratory of Proteomics, Core Facility of Protein Science, Institute of Biophysics, Chinese Academy of Sciences, for mass spectrometry analysis. We thank Y. Wu for his research assistant service. This work is funded by National Natural Science Foundation of China (grants 92157102, to Y.Z., and 31971134, to X.C.Z.), National Key Research and Development Program of China (grant 2021YFA1301501, to Y.Z.) and Chinese Academy of Sciences Strategic Priority Research Program (grants XDB37030304, to Y.Z., and XDB08020301 and XDB37030301, to X.C.Z.).
Author information
Authors and Affiliations
Contributions
Y.Z. conceived and supervised the project. H.Z. and M.L. carried out molecular cloning. H.Z. expressed purified protein complex samples and prepared samples for cryo-EM study. H.Z. and Y.G. carried out cryo-EM data collection. B.L., Y.G., H.X. and Y.Z. processed the cryo-EM data, built models and prepared figures. J.S. and L.H. made the PIGK-deficient HEK293 cell line. H.Z., J.S., L.H. and Y.D. carried out confocal microscopy and flow cytometric analyses. X.C.Z. and Y.Z. analyzed the structure and wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks Calvin Yip and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Florian Ullrich was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification of the human GPIT complex.
a, Size−exclusive chromatography (Superose 6 increase) of purified sample of human GPIT complex. Peak fractions within the dashed lines were collected for cryo-EM sample preparation. b, Coomassie−blue-stained SDS-PAGE gel of SEC peak fractions. GPIT components are labeled. The bands of PIGS and PIGK are smeared because of varied extent of glycosylation. The experiments were repeated independently more than 3 times with similar results.
Extended Data Fig. 2 Cryo-EM analysis of the GPITX complex.
a, Flow chart of cryo-EM data processing of the human GPITX complex. A total of 2,795 movie stacks were collected. A representative motion-corrected micrograph of this dataset is shown here. Particles were picked using Topaz and Gautomatch, and were then submitted to 2D and 3D classifications, followed by Bayesian polish, and CTF refinement and 3D-auto refinement. The final map of GPITX and GPIT was reported at 3.8 Å and 3.4 Å according to GSFSC criterion, respectively. b, Sharpened map of human GPITX complex, colored according to the estimation of local resolution. c, Angular distribution of particles contributed to the final reconstruction. The length of cylinders indicates the number of particles in the designated orientation. d, Half-map FSC curves of human GPIT complex, calculated before (red) and after (blue) PostProcesss.
Extended Data Fig. 3 Cryo-EM data processing of the human GPIT complex.
a, Flow chart of cryo-EM data processing of the human GPIT complex. A total of 2,186 movie stacks were collected (Bar = 40 nm). A representative motion-corrected micrograph of this dataset is shown here. Particles were picked using Gautomatch, Topaz, and Template Picker in cryoSPARC, and were then submitted to several rounds of 3D classification, Bayesian polish, and CTF and 3D-auto refinement. The final map was reported at 3.1 Å according to GSFSC criterion. b, Sharpened map of human GPIT complex, colored according to the estimation of local resolution. c, Angular distribution of particles contributed to the final reconstruction. The length of cylinders indicates the number of particles in the designated orientation. d, Half-map and model-map FSC curves of human GPIT complex. Half-map FSC curves were calculated before (red) and after (blue) PostProcesss, overlaid with a model-map FSC which is calculated between the final map and structural model. e, Representative EM maps for the human GPIT complex and the GPI molecule.
Extended Data Fig. 4 Structural comparison of PIGU and GPAA1 with homologous structures.
a, Side view and top-down view of the subunit PIGU, colored in a rainbow scheme. b, Structural comparison between PIGU and ALG6. The substrate of ALG6 (Dol25-P-Glc) is shown as sticks. c, Structural comparison between GPAA1 and AMPX. Critical residues and hydroxamate are shown as sticks. The zinc ions are shown as spheres.
Extended Data Fig. 5 Distribution of the pathogenic mutations in the GPIT complex.
a-b, Side-view of the structures of the GPIT complex. The Cα atoms of the pathogenic mutations are shown in spheres. c, Reported pathogenic mutations and related diseases are listed.
Extended Data Fig. 6 More analyses on the catalytic site in PIGK.
a, Sequence alignment of human legumain and PIGK. Secondary structures are marked above the alignment. Red triangles indicate residues that are critical for the endopeptidase activity. b, Florescent confocal microscopy analyses of effects of N58KA and D120KN on GPIT activity using HEK293ΔPIGK cells as the host. The first, second, and third columns represent the confocal fluorescence microscopy of GFP, mCherry, and merged images, respectively. The scale bar represents 10 μm. The confocal microscopy experiment was repeated at least five times.
Extended Data Fig. 7 GPIT activity detected in flow cytometry experiment.
a, Cartoon scheme of GPIT activity analysis using GSTCy5-nanobodyGFP. b, The gating strategy for FACS. HEK293T cells was first gated on FSC and SSC signals (R1). GFP and Cy5 positive cells were defined as cells exceeding fluorescence of wild type cells (R2). c, The HEK293 cells (WT or PIGK deficient) expressing GFP-GPI and PIGK (WT or mutants) are analyzed according to GFP and Cy5 fluorescence using flow cytometry. Cells was first gated on FSC and SSC signals. GFP and Cy5 positive cells were defined as cells exceeding fluorescence of wild type cells. The flow cytometry experiment was repeated at least five times.
Extended Data Fig. 8 Sequence alignment of the GPI binding site.
Analysis of amino acid sequences of PIGU and PIGT homologous. HsPIGU and HsPIGT is from Homo sapiens (Uniprot ID: Q9H490 and Q969N2); MmPIGU-MmPIGT, Mus musculus (Q8K358 and Q8BXQ2); DrPIGU-DrPIGT, Danio rerio (F1RES7 and E9QH65); BdPIGU-BdPIGT, Bactrocera dorsalis (A0A6I9VB87 and A0A034WF00); CePIGU-CePIGT, Caenorhabditis elegans (Q22672 and Q19518); ScPIGU-ScPIGT, Saccharomyces cerevisiae (P41733 and P38875); AtPIGU-AtPIGT, Arabidopsis thaliana (F4I1Z0 and Q949U5); AfPIGU-AfPIGT, Aspergillus fumigatus (A0A0J5PDP8 and Q4WMR2); DdPIGU-DdPIGT, Dictyostelium discoideum (Q54T45 and B0G182). Secondary structures are marked above the alignment. Red triangles indicate residues that are involved in binding of the GPI molecule.
Extended Data Fig. 9 Analysis of the extra subunit of GPIT complex.
a, Cryo-EM map of the GPIT complex with six additional transmembrane helices, viewed in parallel to the membrane plane. Subunits PIGK, GPAA1, PIGU, PIGS, and PIGT are colored in cyan, blue, purple, pink, and red, respectively. The extra helices are colored in grey. b, Mass-spectrometry analysis of the purified GPIT protein sample. Subunits of GPIT complexes are highlighted in green. The putative extra subunit RNF121 is highlighted in red. c, Manually-built poly-alanine model of the extra subunit, overlaid with corresponding cryo-EM map (left). 3D structure of the RNF121 predicted by AlphaFold2 (middle); superimposition of structures of extra subunit and RNF121 (right). d, Top-down view of the transmembrane domain of the GPIT-RNF121 complex. e, Western blot analysis of strep tag purified samples (co-expressed GPIT-RNF121 or RNF121 alone) using antibodies against His tag and strep tag to monitor RNF121 and PIGU, respectively. The letters S and E represent supernatant of detergent solubilized cell membrane and elution sample from streptavidin beads, respectively. The experiments were repeated independently 3 times with similar results.
Supplementary information
Source data
Source Data Fig. 3
Unprocessed microscopy images.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 6
Unprocessed microscopy images.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Zhang, H., Su, J., Li, B. et al. Structure of human glycosylphosphatidylinositol transamidase. Nat Struct Mol Biol 29, 203–209 (2022). https://doi.org/10.1038/s41594-022-00726-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00726-6
This article is cited by
-
Molecular basis of the inositol deacylase PGAP1 involved in quality control of GPI-AP biogenesis
Nature Communications (2024)
-
Protein lipidation in health and disease: molecular basis, physiological function and pathological implication
Signal Transduction and Targeted Therapy (2024)
-
Structures of liganded glycosylphosphatidylinositol transamidase illuminate GPI-AP biogenesis
Nature Communications (2023)
-
Structural insights into the regulation of Cas7-11 by TPR-CHAT
Nature Structural & Molecular Biology (2023)
-
Molecular insights into biogenesis of glycosylphosphatidylinositol anchor proteins
Nature Communications (2022)