Abstract
The transient receptor potential cation channel subfamily V member 3 (TRPV3) channel plays a critical role in skin physiology, and mutations in TRPV3 result in the development of a congenital skin disorder, Olmsted syndrome. Here we describe multiple cryo-electron microscopy structures of human TRPV3 reconstituted into lipid nanodiscs, representing distinct functional states during the gating cycle. The ligand-free, closed conformation reveals well-ordered lipids interacting with the channel and two physical constrictions along the ion-conduction pore involving both the extracellular selectivity filter and intracellular helix bundle crossing. Both the selectivity filter and bundle crossing expand upon activation, accompanied by substantial structural rearrangements at the cytoplasmic intersubunit interface. Transition to the inactivated state involves a secondary structure change of the pore-lining helix, which contains a π-helical segment in the closed and open conformations, but becomes entirely α-helical upon inactivation. Together with electrophysiological characterization, structures of TRPV3 in a lipid membrane environment provide unique insights into channel activation and inactivation mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM maps of the wild-type human TRPV3 and K169A have been deposited in the Electron Microscopy Data Bank with accession codes EMD-20917 (apo TRPV3), EMD-20918 (apo K169A), EMD-20919 (K169A with 3 min exposure to 2-APB) and EMD-20920 (K169A in the presence of 2-APB). Atomic coordinates have been deposited in the Protein Data Bank with accession codes 6UW4 (apo TRPV3), 6UW6 (apo K169A), 6UW8 (K169A with 3 min exposure to 2-APB) and 6UW9 (K169A in the presence of 2-APB).
References
Clapham, D. E. TRP channels as cellular sensors. Nature 426, 517–524 (2003).
Ramsey, I. S., Delling, M. & Clapham, D. E. An introduction to TRP channels. Annu. Rev. Physiol. 68, 619–647 (2006).
Julius, D. TRP channels and pain. Annu. Rev. Cell Dev. Biol. 29, 355–384 (2013).
Xu, H. et al. TRPV3 is a calcium-permeable temperature-sensitive cation channel. Nature 418, 181–186 (2002).
Peier, A. M. et al. A heat-sensitive TRP channel expressed in keratinocytes. Science 296, 2046–2049 (2002).
Smith, G. D. et al. TRPV3 is a temperature-sensitive vanilloid receptor-like protein. Nature 418, 186–190 (2002).
Cheng, X. et al. TRP channel regulates EGFR signaling in hair morphogenesis and skin barrier formation. Cell 141, 331–343 (2010).
Bang, S., Yoo, S., Yang, T. J., Cho, H. & Hwang, S. W. Farnesyl pyrophosphate is a novel pain-producing molecule via specific activation of TRPV3. J. Biol. Chem. 285, 19362–19371 (2010).
Miyamoto, T., Petrus, M. J., Dubin, A. E. & Patapoutian, A. TRPV3 regulates nitric oxide synthase-independent nitric oxide synthesis in the skin. Nat. Commun. 2, 369 (2011).
Lin, Z. et al. Exome sequencing reveals mutations in TRPV3 as a cause of Olmsted syndrome. Am. J. Hum. Genet. 90, 558–564 (2012).
Ni, C. et al. A novel mutation in TRPV3 gene causes atypical familial Olmsted syndrome. Sci. Rep. 6, 21815 (2016).
Hu, H. Z. et al. 2-Aminoethoxydiphenyl borate is a common activator of TRPV1, TRPV2, and TRPV3. J. Biol. Chem. 279, 35741–35748 (2004).
Xu, H., Delling, M., Jun, J. C. & Clapham, D. E. Oregano, thyme and clove-derived flavors and skin sensitizers activate specific TRP channels. Nat. Neurosci. 9, 628–635 (2006).
Moqrich, A. et al. Impaired thermosensation in mice lacking TRPV3, a heat and camphor sensor in the skin. Science 307, 1468–1472 (2005).
Chung, M. K., Lee, H., Mizuno, A., Suzuki, M. & Caterina, M. J. 2-Aminoethoxydiphenyl borate activates and sensitizes the heat-gated ion channel TRPV3. J. Neurosci. 24, 5177–5182 (2004).
Xiao, R. et al. Calcium plays a central role in the sensitization of TRPV3 channel to repetitive stimulations. J. Biol. Chem. 283, 6162–6174 (2008).
Phelps, C. B., Wang, R. R., Choo, S. S. & Gaudet, R. Differential regulation of TRPV1, TRPV3, and TRPV4 sensitivity through a conserved binding site on the ankyrin repeat domain. J. Biol. Chem. 285, 731–740 (2010).
Singh, A. K., McGoldrick, L. L. & Sobolevsky, A. I. Structure and gating mechanism of the transient receptor potential channel TRPV3. Nat. Struct. Mol. Biol. 25, 805–813 (2018).
Zubcevic, L. et al. Conformational ensemble of the human TRPV3 ion channel. Nat. Commun. 9, 4773 (2018).
Zubcevic, L., Borschel, W. F., Hsu, A. L., Borgnia, M. J. & Lee, S.-Y. Regulatory switch at the cytoplasmic interface controls TRPV channel gating. Elife 8, e47746 (2019).
Singh, A. K. et al. Structural basis of temperature sensation by the TRP channel TRPV3. Nat. Struct. Mol. Biol. 26, 994–998 (2019).
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113–118 (2013).
Zubcevic, L. et al. Cryo-electron microscopy structure of the TRPV2 ion channel. Nat. Struct. Mol. Biol. 23, 180–186 (2016).
Deng, Z. et al. Cryo-EM and X-ray structures of TRPV4 reveal insight into ion permeation and gating mechanisms. Nat. Struct. Mol. Biol. 25, 252–260 (2018).
Grandl, J. et al. Temperature-induced opening of TRPV1 ion channel is stabilized by the pore domain. Nat. Neurosci. 13, 708–714 (2010).
Jordt, S. E., Tominaga, M. & Julius, D. Acid potentiation of the capsaicin receptor determined by a key extracellular site. Proc. Natl Acad. Sci. USA 97, 8134–8139 (2000).
Cui, Y. et al. Selective disruption of high sensitivity heat activation but not capsaicin activation of TRPV1 channels by pore turret mutations. J. Gen. Physiol. 139, 273–283 (2012).
Bohlen, C. J. et al. A bivalent tarantula toxin activates the capsaicin receptor, TRPV1, by targeting the outer pore domain. Cell 141, 834–845 (2010).
Myers, B. R., Bohlen, C. J. & Julius, D. A yeast genetic screen reveals a critical role for the pore helix domain in TRP channel gating. Neuron 58, 362–373 (2008).
Yang, F., Cui, Y., Wang, K. & Zheng, J. Thermosensitive TRP channel pore turret is part of the temperature activation pathway. Proc. Natl Acad. Sci. USA 107, 7083–7088 (2010).
Jara-Oseguera, A., Huffer, K. E. & Swartz, K. J. The ion selectivity filter is not an activation gate in TRPV1-3 channels. Elife 8, e51212 (2019).
Grandl, J. et al. Pore region of TRPV3 ion channel is specifically required for heat activation. Nat. Neurosci. 11, 1007–1013 (2008).
Kim, S. E., Patapoutian, A. & Grandl, J. Single residues in the outer pore of TRPV1 and TRPV3 have temperature-dependent conformations. PLoS One 8, e59593 (2013).
Hille, B., Dickson, E. J., Kruse, M., Vivas, O. & Suh, B. C. Phosphoinositides regulate ion channels. Biochim. Biophys. Acta 1851, 844–856 (2015).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Denisov, I. G. & Sligar, S. G. Nanodiscs for structural and functional studies of membrane proteins. Nat. Struct. Mol. Biol. 23, 481–486 (2016).
Dang, S. et al. Cryo-EM structures of the TMEM16A calcium-activated chloride channel. Nature 552, 426–429 (2017).
Kern, D. M., Oh, S., Hite, R. K. & Brohawn, S. G. Cryo-EM structures of the DCPIB-inhibited volume-regulated anion channel LRRC8A in lipid nanodiscs. Elife 8, e42636 (2019).
Doerner, J. F., Hatt, H. & Ramsey, I. S. Voltage- and temperature-dependent activation of TRPV3 channels is potentiated by receptor-mediated PI(4,5)P2 hydrolysis. J. Gen. Physiol. 137, 271–288 (2011).
Bang, S., Yoo, S., Yang, T. J., Cho, H. & Hwang, S. 17(R)-resolvin D1 specifically inhibits transient receptor potential ion channel vanilloid 3 leading to peripheral antinociception. Br. J. Pharmacol. 165, 683–692 (2012).
Hu, H. Z. et al. Potentiation of TRPV3 channel function by unsaturated fatty acids. J. Cell. Physiol. 208, 201–212 (2006).
Hughes, T. E. T. et al. Structural insights on TRPV5 gating by endogenous modulators. Nat. Commun. 9, 4198 (2018).
Saotome, K., Singh, A. K., Yelshanskaya, M. V. & Sobolevsky, A. I. Crystal structure of the epithelial calcium channel TRPV6. Nature 534, 506–511 (2016).
McGoldrick, L. L. et al. Opening of the human epithelial calcium channel TRPV6. Nature 553, 233–237 (2018).
Long, S. B., Tao, X., Campbell, E. B. & MacKinnon, R. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007).
Dang, S. et al. Structural insight into TRPV5 channel function and modulation. Proc. Natl Acad. Sci. USA 116, 8869–8878 (2019).
Chung, M. K., Güler, A. D. & Caterina, M. J. Biphasic currents evoked by chemical or thermal activation of the heat-gated ion channel, TRPV3. J. Biol. Chem. 280, 15928–15941 (2005).
Luo, J., Stewart, R., Berdeaux, R. & Hu, H. Tonic inhibition of TRPV3 by Mg2+ in mouse epidermal keratinocytes. J. Invest. Dermatol. 132, 2158–2165 (2012).
Cheng, W. et al. Heteromeric heat-sensitive transient receptor potential channels exhibit distinct temperature and chemical response. J. Biol. Chem. 287, 7279–7288 (2012).
Palovcak, E., Delemotte, L., Klein, M. L. & Carnevale, V. Comparative sequence analysis suggests a conserved gating mechanism for TRP channels. J. Gen. Physiol. 146, 37–50 (2015).
Hinman, A. A., Chuang, H. H. A. B., Bautista, D. M. A. & Julius, D. A. TRP channel activation by reversible covalent modification. Proc. Natl Acad. Sci. USA 103, 19564–19568 (2006).
Li, M., Jiang, J. & Yue, L. Functional characterization of homo- and heteromeric channel kinases TRPM6 and TRPM7. J. Gen. Physiol. 127, 525–537 (2006).
Lievremont, J. P., Bird, G. S. & Putney, J. W. Mechanism of inhibition of TRPC cation channels by 2-aminoethoxydiphenylborane. Mol. Pharmacol. 68, 758–762 (2005).
Togashi, K., Inada, H. & Tominaga, M. Inhibition of the transient receptor potential cation channel TRPM2 by 2-aminoethoxydiphenyl borate (2-APB). Br. J. Pharmacol. 153, 1324–1330 (2008).
Singh, A. K., Saotome, K., McGoldrick, L. L. & Sobolevsky, A. I. Structural bases of TRP channel TRPV6 allosteric modulation by 2-APB. Nat. Commun. 9, 2465 (2018).
Liu, B., Yao, J., Zhu, M. X. & Qin, F. Hysteresis of gating underlines sensitization of TRPV3 channels. J. Gen. Physiol. 138, 509–520 (2011).
Yang, F., Vu, S., Yarov-Yarovoy, V. & Zheng, J. Rational design and validation of a vanilloid-sensitive TRPV2 ion channel. Proc. Natl Acad. Sci. USA 113, E3657–E3666 (2016).
Zhang, F. et al. Engineering vanilloid-sensitivity into the rat TRPV2 channel. Elife 5, e16409 (2016).
Zhang, F., Swartz, K. J. & Jara-Oseguera, A. Conserved allosteric pathways for activation of TRPV3 revealed through engineering vanilloid-sensitivity. Elife 8, e42756 (2019).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Schnorf, M., Potrykus, I. & Neuhaus, G. Microinjection technique: routine system for characterization of microcapillaries by bubble pressure measurement. Exp. Cell. Res. 210, 260–267 (1994).
Acknowledgements
This work was supported by National Institutes of Health Grant R01NS099341 and the Mallinckrodt Foundation grant (to P.Y.), and by the McDonnell Center for Cellular and Molecular Neurobiology Postdoctoral Fellowship (to Z.D.). M.R. and J.A.J.F. are supported by the Washington University Center for Cellular Imaging, which is funded in part by Washington University School of Medicine through the Precision Medicine Initiative, the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CDI-CORE-2015-505 and CDI-CORE-2019-813) and the Foundation for Barnes-Jewish Hospital (3770).
Author information
Authors and Affiliations
Contributions
Z.D. performed biochemical preparations, cryo-EM experiments, structural determination and analysis. G.M. conducted electrophysiology experiments. M.R. and J.A.J.F. performed cryo-EM data acquisition in conjunction with Z.D. Z.X. and H.H. helped with functional studies. P.Y. designed and supervised the project. Z.D., G.M. and P.Y. analyzed the results and prepared the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer review information: Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Reconstitution of human TRPV3 into lipid nanodiscs.
a, Size-exclusion chromatography of TRPV3 reconstituted into lipid nanodiscs made of soybean polar lipids and the scaffold protein MSP2N2 (left panel). Peaks indicating the void, the TRPV3-embedded nanodiscs, the empty nanodiscs, and GFP were labeled. The peak fraction corresponding to TRPV3 channels in nanodiscs was shown on SDS-PAGE (right panel). b, The collected TRPV3-nanodisc fraction ran as a monodisperse peak on size-exclusion chromatography. c, Representative micrograph for negative stain and reference-free 2D class averages indicating a tetrameric channel inserted into nanodiscs (right).
Extended Data Fig. 2 Cryo-EM reconstruction of the wild-type full-length human TRPV3 in lipid nanodiscs.
a, Flowchart of cryo-EM data processing. b, Fourier shell correlation before and after post-processing in RELION2. c, Fourier shell correlation between the refined model and the full map. d, Angular distribution plot of particles used for final reconstruction. e, Cryo-EM density map colored by local resolution. f, Representative cryo-EM density shown as blue mesh contoured at 5.0 σ.
Extended Data Fig. 3 Channel-lipid interactions in human TRPV3.
a, Lipid densities at sites 1, 2, and 3 located in the proximity of the outer pore region. Lipids are putatively modeled as phosphatidylethanolamine to illustrate interactions with the channel. Channel subunits are uniquely colored. Polar and charged residues potentially interacting with lipid head groups are highlighted in stick representation. b, Lipid density at site 4 in the intracellular cavity of the S1-S4 domain. c, Lipid density between the S4-S5 linkers of two neighboring subunits. The putative lipid densities are shown as orange mesh contoured at 3.5 σ.
Extended Data Fig. 4 Cryo-EM reconstruction of the human TRPV3 K169A variant in lipid nanodiscs.
a, Flowchart of cryo-EM data processing. Two major conformations, presumably representing the open (~69%) and inactivated states (31%), were refined to resolutions of 3.66 Å and 4.4 Å, respectively. b, Fourier shell correlation before and after post-processing in RELION3 for the open state. c, Fourier shell correlation between the refined model and the full map for the open state. d, Angular distribution plot of particles used for final reconstruction for the open state. e, Cryo-EM density map colored by local resolution for the open state. f, Representative cryo-EM density shown as blue mesh contoured at 5.0 σ.
Extended Data Fig. 6 Lipid in the analogous vanilloid binding pocket in TRPV3.
a,b, Putative lipid density shown as orange mesh contoured at 4.5 σ in the analogous vanilloid binding pocket in the closed (a) and open (b) conformations.
Extended Data Fig. 7 Cryo-EM reconstruction of K169A accompanied by 2-APB in an inactivated state.
a, Flowchart of cryo-EM data processing. b, Fourier shell correlation before and after post-processing in RELION3. c, Fourier shell correlation between the refined model and the full map. d, Angular distribution plot of particles used for final reconstruction. e, Cryo-EM density map colored by local resolution. f, Representative cryo-EM density shown as blue mesh contoured at 5.0 σ.
Extended Data Fig. 8 Cryo-EM reconstruction of K169A briefly exposed to 2-APB for 3 minutes in an open conformation.
a, Flowchart of cryo-EM data processing. b, Fourier shell correlation before and after post-processing in RELION3. c, Fourier shell correlation between the refined model and the full map. d, Angular distribution plot of particles used for final reconstruction. e, Cryo-EM density map colored by local resolution. f, Representative cryo-EM density shown as blue mesh contoured at 5.0 σ.
Extended Data Fig. 9 TRPV3 pore structures in distinct functional states.
a–d, The pore structures in the apo closed wild-type TRPV3 channel (a), the apo open K169 mutant (b), the 2-APB-bound open K169A (c), and 2-APB-bound inactivated states (d). Also shown are cryo-EM densities of the normalized cryo-EM maps contoured at 6.0 σ.
Extended Data Fig. 10 Mapping the pore mutations enabling TRPV3-6M activation by RTX.
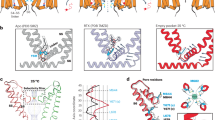
Two views of the closed TRPV3 pore, highlighting the pore mutations that render TRPV3-6M sensitive to RTX activation. These mutations, including V587L, A606V, F625L, F656I, and F666Y, are shown as stick representation and colored in green.
Supplementary information
Rights and permissions
About this article
Cite this article
Deng, Z., Maksaev, G., Rau, M. et al. Gating of human TRPV3 in a lipid bilayer. Nat Struct Mol Biol 27, 635–644 (2020). https://doi.org/10.1038/s41594-020-0428-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0428-2
This article is cited by
-
Structural basis of TRPV3 inhibition by an antagonist
Nature Chemical Biology (2023)
-
A pentameric TRPV3 channel with a dilated pore
Nature (2023)
-
FRET analysis of the temperature-induced structural changes in human TRPV3
Scientific Reports (2023)
-
Structural insights into TRPV2 activation by small molecules
Nature Communications (2022)
-
Structural mechanism of TRPV3 channel inhibition by the anesthetic dyclonine
Nature Communications (2022)