Abstract
Understanding the targeting and spreading patterns of long non-coding RNAs (lncRNAs) on chromatin requires a technique that can detect both high-intensity binding sites and reveal genome-wide changes in spreading patterns with high precision and confidence. Here we determine lncRNA localization using biotinylated locked nucleic acid (LNA)-containing oligonucleotides with toehold architecture capable of hybridizing to target RNA through strand-exchange reaction. During hybridization, a protecting strand competitively displaces contaminating species, leading to highly specific RNA capture of individual RNAs. Analysis of Drosophila roX2 lncRNA using this approach revealed that heat shock, unlike the unfolded protein response, leads to reduced spreading of roX2 on the X chromosome, but surprisingly also to relocalization to sites on autosomes. Our results demonstrate that this improved hybridization capture approach can reveal previously uncharacterized changes in the targeting and spreading of lncRNAs on chromatin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
Analysis pipeline is available at https://bitbucket.org/machyna/thchart-pipeline. All computational scripts are available upon request.
Change history
18 March 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41594-020-0413-9
References
Engreitz, J. M., Ollikainen, N. & Guttman, M. Long non-coding RNAs: spatial amplifiers that control nuclear structure and gene expression. Nat. Rev. Mol. Cell Biol. 17, 756–770 (2016).
Rinn, J. L. lncRNAs: linking RNA to chromatin. Cold Spring Harb. Perspect. Biol. 6, a018614 (2014).
Samata, M. & Akhtar, A. Dosage compensation of the X chromosome: a complex epigenetic assignment involving chromatin regulators and long noncoding RNAs. Annu. Rev. Biochem. 87, 323–350 (2018).
Straub, T. et al. The chromosomal high-affinity binding sites for the Drosophila dosage compensation complex. PLoS Genet. 4, e1000302 (2008).
Alekseyenko, A. A. et al. A sequence motif within chromatin entry sites directs MSL establishment on the Drosophila X chromosome. Cell 134, 599–609 (2008).
Larschan, E. et al. MSL complex is attracted to genes marked by H3K36 trimethylation using a sequence-independent mechanism. Mol. Cell 28, 121–133 (2007).
Bell, O. et al. Transcription-coupled methylation of histone H3 at lysine 36 regulates dosage compensation by enhancing recruitment of the MSL complex in Drosophila melanogaster. Mol. Cell Biol. 28, 3401–3409 (2008).
Chu, C. et al. Genomic maps of long noncoding RNA occupancy reveal principles of RNA–chromatin interactions. Mol. Cell 44, 667–678 (2011).
Simon, M. D. et al. The genomic binding sites of a noncoding RNA. Proc. Natl Acad. Sci. USA 108, 20497–20502 (2011).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Machyna, M. & Simon, M. D. Catching RNAs on chromatin using hybridization capture methods. Brief. Funct. Genomics 17, 96–103 (2017).
Koshkin, A. A. et al. LNA (locked nucleic acids): synthesis of the adenine, cytosine, guanine, 5-methylcytosine, thymine and uracil bicyclonucleoside monomers, oligomerisation, and unprecedented nucleic acid recognition. Tetrahedron 54, 3607–3630 (1998).
Braasch, D. A. & Corey, D. R. Locked nucleic acid (LNA): fine-tuning the recognition of DNA and RNA. Chem. Biol. 8, 1–7 (2001).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Vieregg, J. R. et al. Selective nucleic acid capture with shielded covalent probes. J. Am. Chem. Soc. 135, 9691–9699 (2013).
Karkare, S. & Bhatnagar, D. Promising nucleic acid analogs and mimics: characteristic features and applications of PNA, LNA, and morpholino. Appl. Microbiol. Biotechnol. 71, 575–586 (2006).
Murugan, R. On the origin of affinity–specificity negative correlation in DNA–probe interactions. Biophys. Chem. 116, 105–109 (2005).
Demidov, V. V. & Frank-Kamenetskii, M. D. Two sides of the coin: affinity and specificity of nucleic acid interactions. Trends Biochem. Sci. 29, 62–71 (2004).
Wu, L. R. et al. Continuously tunable nucleic acid hybridization probes. Nat. Methods 12, 1191–1196 (2015).
Lingner, J. & Cech, T. R. Purification of telomerase from Euplotes aediculatus: requirement of a primer 3’ overhang. Proc. Natl Acad. Sci. USA 93, 10712–10717 (1996).
Kennedy-Darling, J. et al. Multiplexed programmable release of captured DNA. Calbiochem 15, 2353–2356 (2014).
Quinn, J. J. et al. Revealing long noncoding RNA architecture and functions using domain-specific chromatin isolation by RNA purification. Nat. Biotechnol. 32, 933–940 (2014).
Alekseyenko, A. A. et al. Heterochromatin-associated interactions of Drosophila HP1a with dADD1, HIPP1, and repetitive RNAs. Genes Dev. 28, 1445–1460 (2014).
Villa, R. et al. PionX sites mark the X chromosome for dosage compensation. Nature 537, 244–248 (2016).
Fauth, T. et al. The DNA binding CXC domain of MSL2 is required for faithful targeting the dosage compensation complex to the X chromosome. Nucleic Acids Res. 38, 3209–3221 (2010).
Sural, T. H. et al. The MSL3 chromodomain directs a key targeting step for dosage compensation of the Drosophila melanogaster X chromosome. Nat. Struct. Mol. Biol. 15, 1318–1325 (2008).
Kharchenko, P. V. et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 471, 480–485 (2011).
Ramírez, F. et al. High-affinity sites form an interaction network to facilitate spreading of the MSL complex across the X chromosome in Drosophila. Mol. Cell 60, 146–162 (2015).
Schauer, T. et al. Chromosome topology guides the Drosophila dosage compensation complex for target gene activation. EMBO Rep. 18, 1854–1868 (2017).
West, J. A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Quinn, J. J. et al. Rapid evolutionary turnover underlies conserved lncRNA-genome interactions. Genes Dev. 30, 191–207 (2016).
Cheetham, S. W. & Brand, A. H. RNA-DamID reveals cell-type-specific binding of roX RNAs at chromatin-entry sites. Nat. Struct. Mol. Biol. 25, 109–114 (2017).
O’Brien, T. & Lis, J. T. Rapid changes in Drosophila transcription after an instantaneous heat shock. Mol. Cell Biol. 13, 3456–3463 (1993).
Schofield, J. A. et al. TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding. Nat. Methods 15, 221–225 (2018).
Demakova, O. V. et al. The MSL complex levels are critical for its correct targeting to the chromosomes in Drosophila melanogaster. Chromosoma 112, 103–115 (2003).
Palmer, M. J. et al. The male-specific lethal-one (msl-1) gene of Drosophila melanogaster encodes a novel protein that associates with the X chromosome in males. Genetics 134, 545–557 (1993).
Soruco, M. M. L. et al. The CLAMP protein links the MSL complex to the X chromosome during Drosophila dosage compensation. Genes Dev. 27, 1551–1556 (2013).
Kuzu, G. et al. Expansion of GA dinucleotide repeats increases the density of CLAMP binding sites on the X-chromosome to promote Drosophila dosage compensation. PLoS Genet. 12, e1006120 (2016).
Straub, T. et al. Different chromatin interfaces of the Drosophila dosage compensation complex revealed by high-shear ChIP-seq. Genome Res. 23, 473–485 (2013).
Valsecchi, C. I. K. et al. Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells. Nat. Commun. 9, 3626 (2018).
Duarte, F. M. et al. Transcription factors GAF and HSF act at distinct regulatory steps to modulate stress-induced gene activation. Genes Dev. 30, 1731–1746 (2016).
Bowman, S. K. et al. Multiplexed Illumina sequencing libraries from picogram quantities of DNA. BMC Genom. 14, 466 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Song, Q. & Smith, A. D. Identifying dispersed epigenomic domains from ChIP-Seq data. Bioinformatics 27, 870–871 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Shen, L. et al. diffReps: detecting differential chromatin modification sites from ChIP-seq data with biological replicates. PLoS One 8, e65598 (2013).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Wang, Q. et al. Sub-kb Hi-C in D. melanogaster reveals conserved characteristics of TADs between insect and mammalian cells. Nat. Commun. 9, 188 (2018).
Wingett, S. et al. HiCUP: pipeline for mapping and processing Hi-C data. F1000Res. 4, 1310 (2015).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 11 (2015).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Acknowledgements
The authors thank J. Steitz, D. Schatz, C. Lin, N. Dimitrova and K. Neugebauer for insightful comments; members of the Simon Lab for critical feedback and prior work on alternative oligonucleotide designs; and The Charles H. Revson Foundation (M.M.), The American Heart Association (L.K.), the Searle foundation (M.D.S.), and NIH New Innovator Award DP2 HD083992 (M.D.S.) for funding.
Author information
Authors and Affiliations
Contributions
M.M. and M.D.S. designed thCHART. M.M. performed most experiments and data analysis. L.K. performed TT-TimeLapse-seq and M.D.S. analyzed the resulting data. M.M. and M.D.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare that thCHART technology has been protected under patent application no. 16/412,527.
Additional information
Peer review information Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Development of thCHART approach.
a, Sequence of the capture oligonucleotide (green) with DNA backbone contains LNA nucleotides (uppercase letters and green knobs) in its central part and is pre-hybridized with complementary RNA protecting oligonucleotide (grey). b, Electrophoretic mobility shift assay (EMSA) of fluorescent RNA target hybridized to toehold capture oligonucleotide (CO). CO:PO pairs hybridized via strand exchange reactions at increasing ratios of CO:PO (1:1, 1:3, 1:10) for 1h at 20 °C show tunability for RNA PO, while DNA PO yield remains unchanged. Uncropped gel images are shown in the Source Data. c, CHART enriched RNA was analyzed by RT-qPCR for roX2 RNA and two off-target RNAs (Act5C, CG14438). Shown are averages of n = 2 technical replicates normalized to input. Enrichment was performed with DNA CO (DNA), unprotected LNA CO (LNA) or protected LNA CO (thLNA) followed by mild wash conditions (Simon 2011). d, RNA retention assay shows relative amount of RNA target remaining hybridized to CO after subjected to stringent washes with wash buffer (10 mM NaCl, 10 mM HEPES pH 7.4, 2 mM EDTA, 1 mM EGTA, 0.2% SDS). e, Enrichment performed with DNA CO (DNA), or protected LNA CO (thLNA) followed by mild or stringent washes analyzed as in c. Shown is average of n = 2 technical replicates. f, Schematic representation of elution strategy. g, EMSA analysis of efficiency and specificity of target RNA displacement from CO by elution oligonucleotide (EO). Fluorescent target RNA was prehybridized to LNA–DNA CO and then incubated with excess of EO to trigger RNA release. h, Enrichment was performed with DNA or thLNA CO design, subjected to stringent washes and eluted either with proteinase K or excess of EO. Note that in thCHART Ct values for off-target RNAs reached 40 cycles. Data were analyzed as in c. i, Analysis of enriched DNA from roX2 CHART assayed with qPCR. Two X-linked CES loci (roX2, CES-5C2) and an autosomal locus (Act87E) were investigated. Shown are averages of technical duplicates normalized to input. j, Predicted roX2 secondary structure with highlighted regions used for CO design. k, Proportion (%) of reads that were discarded during analysis. Data for graphs in c-e, h, i are available as source data.
Extended Data Fig. 2 thCHART shows high reproducibility and specificity.
a, Genomic tracks show read coverage from RNA-seq after roX2 enrichment. Experiments were performed with DNA or unprotected LNA CO and compared to thCHART enrichment with roX2-antisense or non-targeting CO (control). Note that the bottom tracks are in log scale to allow visualization of low signal in control and input. b, Comparing proportion of mapped reads in thCHART and published ChIP40 and ChIRP32 datasets. c, Comparison of chromosome X signal enrichment. d, DNA enrichment tracks show reproducibility of thCHART with two distinct COs. This experiment was performed using a DNA PO. e, Proportion of reads uniquely mapping to D. melanogaster chromosomes from indicated experiments. f, Enrichment of chromosome X signal over autosomes. The number of reads were normalized to the chromosome length before calculating the enrichment. g, h, Correlation of chromosome X roX2 signal enrichment (1 kb bins) between (g) odd and even ChIRP datasets and (h) CO1 and CO2 thCHART experiments. i, Proportion of reads uniquely mapping to mouse chromosomes from indicated experiments. j, Genome track of mouse Xist thCHART performed with single toehold CO. k, Enrichment of chromosome X signal over autosomes from indicated experiments similar to e. l, Cumulative mean distribution of read coverage in 1 kb bins for thCHART, CHART9 and ChIRP8 datasets. m, Comparing relation between FDR score and peak calling accuracy. High-quality thCHART data allow more accurate peak calling, which is consistent with low FDR score (FDR = 0.1). n, Contrasting FDR scores of thCHART with published roX1 or roX2 ChIRP and MSL2 or MSL3 ChIP datasets. Data for graphs in b, c, e, f, i, k are available as source data.
Extended Data Fig. 3 High significance roX2 thCHART regions are enriched in active chromatin marks and insulator proteins.
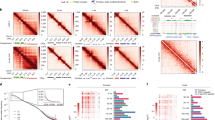
a, Heatmap of ChIP-chip signal (modENCODE) around the significant roX2 regions identified with thCHART. b, A representative HiC contact heat map (200 bp resolution) and TADs (Wang at al.)54 showed overlap with roX2 thCHART regions and insulator proteins ChIP-seq (modENCODE). c, Distribution of TAD boundaries (identified by Wang et al.) in vicinity of significant roX2 thCHART regions. d-i, Heatmaps of ChIP-chip or -seq signal (modENCODE) around the significant roX2 regions identified with thCHART. j-l, Extent of chromosome X associated with hallmarks of active chromatin (H3K36me3 and RNAPII ChIP-seq) and actively transcribed genes demined by RNA-seq (modEncode). thCHART signal (FDR < 0.1) showed higher degree of overlap with these regions than CHART and ChIRP. m-o, Heatmaps for insulators presented similar to b-g. p, Each roX2 region was split in the middle into two equal parts and number of HiC contacts between left half (5’ boundary) and the remaining portion of the roX2 region (in) or adjacent outside region of the same length (out) was calculated. The same procedure was used for the right half (3’ boundary) of roX2 regions. In box plots of n=313 roX2 regions, center lines indicate the median, box limits indicate the first and third quartiles, whiskers indicate 1.5× interquartile range (IQR), notches indicate 95% confidence interval. (* P-value = 1.121e-05, 1.277e-08; paired wilcox-test) Data for graphs in j-l are available as source data.
Extended Data Fig. 4 Heat shock restricts MSL complex chromosome X spreading and binding to some CES sites.
a, 20-Hydroxyecdysone treatment for 24h (dark grey) leads to significant activation of several X-linked genes (* P values: br = 0.046, Pvf1 = 0.029, Sgg = 0.039; Mct = 0.135, unpaired t-test). Expression was tested by RT-qPCR and the fold enrichment over mock treated sample values were normalized to 18S levels. Average of n=2 biologically independent samples ± s.e.m. shown. b, Treatment with 20-Hydroxyecdysone for 24 h does not lead to extension of MSL complex spreading as shown for two representative ecdysone-regulated genes. c, Heat map of Pearson correlation coefficient between individual thCHART replicates. d, MA-plot of average heat shock thCHART signal. Bins (1kb) with more than 2-fold significant change (FDR < 0.05) are indicated in red. e, Changes in spliced Xbp1 and nascent RNA levels in response to unfolded protein response (DTT) measured by RT-qPCR (* P values: Xbp1 = 0.0084, Hsc70-3 = 0.0194, Act5C = 0.00323, unpaired t-test). Bars are mean of n=3 biologically independent samples ± s.e.m. f, Correlation of thCHART signal at CES. Average read coverage was calculated in 500 bp window around CES summit. The CES detected by Diffreps to be significantly (FDR < 0.05) downregulated or upregulated by more than 2-fold are indicated in red and green, respectively. g, roX2 localization to CES is differentially affected after heat shock. We analyzed DNA fraction of roX2 thCHART with qPCR at six X-linked CES loci and control autosomal locus (Act87E) and detected significant changes caused by heat shock conditions (* P values: CG8636 = 0.0052, CG32758 = 0.0057, AP-1-2-beta = 0.0064; Gbeta5, P = 0.053, unpaired t-test). Bars are mean of n=3 biologically independent samples ± s.e.m. h, Example of roX2 thCHART signal at CES unaffected (left), downregulated (center) and upregulated (right) by heat shock. (i) Fold change of thCHART chromatin qPCR signal after heat shock performed with one of two distinct COs targeting roX2 regions shown in Extended Data Fig. 1j. Left side of figure is reused from Fig. 3c. (* P values: roX2 = 0.19; 0.009, CES-5C2 = 0.18; 0.078, IntS6 = 0.00026; 0.002, CG8636 = 6.7x10-5; 3.9x10-5, unpaired t-test) Bars are mean of n=3 biologically independent samples ± s.e.m. j, k, Heat shock (37 °C, 1 h, dark grey) caused significant upregulation of heat shock genes and downregulation of non-heat shock related genes (* P values: HSP70Aa = 0.0034, Act5C < 0.0001, RpL17 = 0.0096, roX2-pre = 0.0058, roX2-total = 0.3330, unpaired t-test) as detected by RT-qPCR. Shown are averages of n=3 independent biological replicates ± s.e.m. l, m, TT-TimeLapse-seq read coverage is color-coded according to number of detected T-to-C mutations. Heat shock (l) induces expression of heat shock genes, while (m) globally suppresses transcription at Pol II genes on chromosome X (and autosomes). Data for graphs in a, e, g, i-k are available as source data.
Extended Data Fig. 5 Heat shock causes genome-wide redistribution of MSL complex to autosomes.
a, Treatment with DRB (3 h, dark grey) caused significant reduction in RNAPII transcription (* P values: RpL17 = 0.0003, Act5C < 0.0001, unpaired t-test). Shown are averages of n = 3 biologically independent samples ± s.e.m. b, Genomic tracks of roX2 thCHART signal at a representative X-linked locus. c, Correlation of roX2 thCHART signal between DRB and mock-treated cells. d-i, Read coverage per 1 kb genomic regions was compared between heat shock and control replicates averages (dark grey) and two replicates of control samples (light grey). j, Genome-wide view of roX2 thCHART replicates show the appearance of autosomal signals after heat shock. k, Enriched motif (E-value = 1.7e-429) in autosomal roX2 peaks agrees well with sequence identified in CES and previously described MRE motif4. l, ChIP-seq signal heatmaps centered around summits of roX2 autosomal peaks (n = 468). Shown are MSL complex components and CLAMP38,40,41. m, Fold change of thCHART chromatin qPCR signal after heat shock performed with one of two distinct COs targeting roX2 regions shown in Extended Data Fig. 1j. Assayed are autosomal regions with roX2 heat shock-induced peaks. (* P values: CG1399 = 0.051; 0.037, CR44856 = 0.049; 0.037, CG1943 = 0.023; 0.065, unpaired t-test) Bars are mean of n = 3 biologically independent samples ± s.e.m. n, Top 5 most enriched GO-terms for genes containing heat shock-induced roX2 peaks. Data for graphs in a, m are available as source data.
Extended Data Fig. 6 thCHART further refines ncRNA chromatin binding pattern.
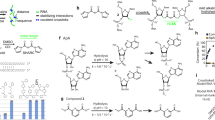
a, For all genes on chrX (within n = 1483, outside n = 581), PRO-seq signal42 between 20 min heat shock-treated and control cells was compared (*P value < 2.2e-16, unpaired t-test) . Genes found inside roX2 spreading regions are on average downregulated, while transcription of genes not covered by roX2 signal is on average unchanged. b, Secondary structure of human TERC RNA with highlighted (green) region that was used for designing of CO for thCHART. c, Sequence design of CO (green) and PO (grey) with marked LNA modifications (green knobs). d, Ratio between number of reads originating from telomeric regions in pulldown / input after normalization to sequencing depth compared to data from ref. 8. e, Genomic view of representative Myc locus, comparing ChIRP published data and thCHART TERC pulldown. f-h, Comparison of DNA fold enrichment between ChIPR even/odd and combined dataset and thCHART experiment. After aligning, the reads having multiple perfect alignments (q < 2) or aligning to repetitive parts of genome (simple repeats, low complexity, satellite repeats) were removed. Each dot represents average signal fold enrichment in genomic 10 kb region overlapping with ChIRP peak from ref. 8. Data for graph in d is available as source data.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Unprocessed western blots
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed gels
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Rights and permissions
About this article
Cite this article
Machyna, M., Kiefer, L. & Simon, M.D. Enhanced nucleotide chemistry and toehold nanotechnology reveals lncRNA spreading on chromatin. Nat Struct Mol Biol 27, 297–304 (2020). https://doi.org/10.1038/s41594-020-0390-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0390-z