Abstract
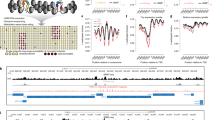
Determining the spatial organization of chromatin in cells mainly relies on crosslinking-based chromosome conformation capture techniques, but resolution and signal-to-noise ratio of these approaches is limited by interference from DNA-bound proteins. Here we introduce chemical-crosslinking assisted proximity capture (CAP-C), a method that uses multifunctional chemical crosslinkers with defined sizes to capture chromatin contacts. CAP-C generates chromatin contact maps at subkilobase (sub-kb) resolution with low background noise. We applied CAP-C to formaldehyde prefixed mouse embryonic stem cells (mESCs) and investigated loop domains (median size of 200 kb) and nonloop domains (median size of 9 kb). Transcription inhibition caused a greater loss of contacts in nonloop domains than loop domains. We uncovered conserved, transcription-state-dependent chromatin compartmentalization at high resolution that is shared from Drosophila to human, and a transcription-initiation-dependent nuclear subcompartment that brings multiple nonloop domains in close proximity. We also showed that CAP-C could be used to detect native chromatin conformation without formaldehyde prefixing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
CAP-C, in situ Hi-C, ChIP–seq and PLAC-seq raw sequencing data are available on Gene Expression Omnibus accession: GSE110061. ChIP–seq data can also be found in the following links on UCSD genome browser: CAP-C–mESC: https://genome.ucsc.edu/s/anton386/CAPC%2DmESC; CAP-C–CTCF-AID: https://genome.ucsc.edu/s/anton386/CAPC%2DCTCF%2DAID; CAP-C–induction: https://genome.ucsc.edu/s/anton386/CAPC%2DInduction. Source data are provided with this paper.
Code availability
Code for CAP-C and ChIP–seq analysis is available on GitHub: http://github.com/ouyang-lab/CAPC.
References
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell. 160, 1049–1059 (2015).
Pombo, A. & Dillon, N. Three-dimensional genome architecture: players and mechanisms. Nat. Rev. Mol. Cell Biol. 16, 245–257 (2015).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178. e1120 (2018).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Dostie, J. et al. Chromosome conformation capture carbon copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Dowen, J. M. et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 159, 374–387 (2014).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Ramani, V. et al. Mapping 3D genome architecture through in situ DNase Hi-C. Nat. Protoc. 11, 2104–2121 (2016).
Maiti, P. K., Çaǧın, T., Wang, G. & Goddard, W. A. Structure of PAMAM dendrimers::generations 1 through 11. Macromol. 37, 6236–6254 (2004).
Astruc, D., Boisselier, E. & Ornelas, C. Dendrimers designed for functions: from physical, photophysical, and supramolecular properties to applications in sensing, catalysis, molecular electronics, photonics, and nanomedicine. Chem. Rev. 110, 1857–1959 (2010).
Eichman, B. F. et al. The crystal structures of psoralen cross-linked DNAs: drug-dependent formation of Holliday junctions. J. Mol. Biol. 308, 15–26 (2001).
Liang, Z. et al. BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions. Nat. Commun. 8, 1622 (2017).
Kolb, H. C., Finn, M. & Sharpless, K. B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. 40, 2004–2021 (2001).
Hnisz, D., Day, D. S. & Young, R. A. Insulated neighborhoods: structural and functional units of mammalian gene control. Cell 167, 1188–1200 (2016).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 e922 (2017).
Davuluri, R. V. et al. The functional consequences of alternative promoter use in mammalian genomes. Trends Genet. 24, 167–177 (2008).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell. 67, 837–852 e837 (2017).
Rowley, M. J. et al. Condensin II counteracts cohesin and RNA polymerase II in the establishment of 3D chromatin organization. Cell Rep. 26, 2890–2903. e2893 (2019).
Mitchell, J. A. & Fraser, P. Transcription factories are nuclear subcompartments that remain in the absence of transcription. Genes Dev. 22, 20–25 (2008).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598 (2015).
Cubeñas-Potts, C. et al. Different enhancer classes in Drosophila bind distinct architectural proteins and mediate unique chromatin interactions and 3D architecture. Nucleic Acids Res. 45, 1714–1730 (2016).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell. 174, 744–757.e724 (2018).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res. 26, 1345 (2016).
Hsieh, T.-H. S. et al. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell. 78, 539–553.e538 (2020).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature. 547, 232–235 (2017).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell. 169, 216–228 e219 (2017).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell. 171, 557–572 e524 (2017).
Lu, H. et al. Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 58, 318–323 (2018). 1.
Schoenfelder, S. et al. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42, 53 (2010).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature. 544, 503 (2017).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature. 547, 236–240 (2017).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science. 361, eaar3958 (2018).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature. 572, 543–548 (2019).
Kubo, N. et al. Preservation of chromatin organization after acute loss of CTCF in mouse embryonic stem cells. Preprint at bioRxiv https://doi.org/10.1101/118737 (2017).
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Law, C. et al. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Res. 5, https://doi.org/10.12688/f1000research.9005.3 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997v2 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 26, 841–842 (2010).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Yang, T. et al. HiCRep: assessing the reproducibility of Hi-C data using a stratum-adjusted correlation coefficient. Genome Res. 27, 1939–1949 (2017).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell. 171, 305–320 e324 (2017).
DeMare, L. E. et al. The genomic landscape of cohesin-associated chromatin interactions. Genome Res. 23, 1224–1234 (2013).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature. 551, 51–56 (2017).
Acknowledgements
We thank A. Andersen (Life Science Editors) for editing the manuscript, P.W. Faber for helping with high-throughput sequencing, and J. Fei and J. Zhang for helping with DNA–FISH experiments. This work was supported by grant nos. NIH F32CA221007 (B.T.H.), NIH RM1HG008935 (C.H.), NIH U54CA193419 (C.H.), the Ludwig Institute for Cancer Research (B.R. and C.H.) and NIH/NIGMS grant no. R35 GM124998 (Z.O.). C.H. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.H. and Q.Y. conceived the original idea. Q.Y., A.Y.C., X.G., Z.O. and C.H. designed the experiments. Q.Y., X.G., T.W. and M.Y., performed the experiments, A.Y.C. and B.L. performed analysis of the sequencing data. B.T.H. aided in the analysis of the FISH data. A.Y.C., Q.Y., X.G., B.R., Z.O. and C.H. analyzed the data and interpreted the finding. Q.Y., X.G., B.T.H. and A.Y.C. wrote the manuscript with input from B.R., Z.O. and C.H.
Corresponding authors
Ethics declarations
Competing interests
C.H. is a scientific founder and a member of the scientific advisory board of Accent Therapeutics, Inc. and a shareholder of Epican Genetech. B.R. is a co-founder and a member of the scienceitific advisory board of Arima Genomics Inc.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–30 and Notes 1–5.
Supplementary Table 1
DNA–FISH probes
Supplementary Table 2
The sequencing information of CAP-C and in situ Hi-C
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
You, Q., Cheng, A.Y., Gu, X. et al. Direct DNA crosslinking with CAP-C uncovers transcription-dependent chromatin organization at high resolution. Nat Biotechnol 39, 225–235 (2021). https://doi.org/10.1038/s41587-020-0643-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0643-8
This article is cited by
-
KARR-seq reveals cellular higher-order RNA structures and RNA–RNA interactions
Nature Biotechnology (2024)
-
KARR-seq maps higher-order RNA structures and RNA–RNA interactions across the transcriptome
Nature Biotechnology (2024)
-
A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
Nature Communications (2024)
-
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Nature Communications (2024)
-
3D genomics and its applications in precision medicine
Cellular & Molecular Biology Letters (2023)