Abstract
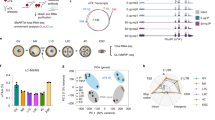
The ten-eleven translocation 2 (TET2) protein, which oxidizes 5-methylcytosine in DNA, can also bind RNA; however, the targets and function of TET2–RNA interactions in vivo are not fully understood. Using stringent affinity tags introduced at the Tet2 locus, we purified and sequenced TET2-crosslinked RNAs from mouse embryonic stem cells (mESCs) and found a high enrichment for tRNAs. RNA immunoprecipitation with an antibody against 5-hydroxymethylcytosine (hm5C) recovered tRNAs that overlapped with those bound to TET2 in cells. Mass spectrometry (MS) analyses revealed that TET2 is necessary and sufficient for the deposition of the hm5C modification on tRNA. Tet2 knockout in mESCs affected the levels of several small noncoding RNAs originating from TET2-bound tRNAs that were enriched by hm5C immunoprecipitation. Thus, our results suggest a new function of TET2 in promoting the conversion of 5-methylcytosine to hm5C on tRNA and regulating the processing or stability of different classes of tRNA fragments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA sequencing data generated for this study have been deposited in the NCBI GEO with accession number GSE133472. Raw mass spectrometry data are available on figshare (https://doi.org/10.6084/m9.figshare.c.5133581).
Code availability
Software utilized for each analysis is detailed in the relevant Methods section. Scripts and R markdown documents to generate figures are available from the corresponding author upon request.
References
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Gilbert, W. V., Bell, T. A. & Schaening, C. Messenger RNA modifications: Form, distribution, and function. Science 352, 1408–1412 (2016).
Alarcon, C. R., Lee, H., Goodarzi, H., Halberg, N. & Tavazoie, S. F. N6-methyladenosine marks primary microRNAs for processing. Nature 519, 482–485 (2015).
Patil, D. P. et al. m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Ke, S. et al. m6A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes Dev. 31, 990–1006 (2017).
Squires, J. E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Delatte, B. et al. RNA biochemistry. transcriptome-wide distribution and function of RNA hydroxymethylcytosine. Science 351, 282–285 (2016).
Huang, T., Chen, W., Liu, J., Gu, N. & Zhang, R. Genome-wide identification of mRNA 5-methylcytosine in mammals. Nat. Struct. Mol. Biol. 26, 380–388 (2019).
Hussain, S., Aleksic, J., Blanco, S., Dietmann, S. & Frye, M. Characterizing 5-methylcytosine in the mammalian epitranscriptome. Genome Biol. 14, 215 (2013).
Legrand, C. et al. Statistically robust methylation calling for whole-transcriptome bisulfite sequencing reveals distinct methylation patterns for mouse RNAs. Genome Res. 27, 1589–1596 (2017).
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010).
Tuorto, F. et al. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat. Struct. Mol. Biol. 19, 900–905 (2012).
Gkatza, N. A. et al. Cytosine-5 RNA methylation links protein synthesis to cell metabolism. PLoS Biol. 17, e3000297 (2019).
Blanco, S. et al. Aberrant methylation of tRNAs links cellular stress to neuro-developmental disorders. EMBO J. 33, 2020–2039 (2014).
Wu, X. & Zhang, Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat. Rev. Genet. 18, 517–534 (2017).
Dawlaty, M. M. et al. Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev. Cell 29, 102–111 (2014).
Li, X. et al. Ten-eleven translocation 2 interacts with forkhead box O3 and regulates adult neurogenesis. Nat. Commun. 8, 15903 (2017).
Shen, Q. et al. Tet2 promotes pathogen infection-induced myelopoiesis through mRNA oxidation. Nature 554, 123–127 (2018).
Fu, L. et al. Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J. Am. Chem. Soc. 136, 11582–11585 (2014).
DeNizio, J. E., Liu, M. Y., Leddin, E. M., Cisneros, G. A. & Kohli, R. M. Selectivity and promiscuity in TET-mediated oxidation of 5-methylcytosine in DNA and RNA. Biochemistry 58, 411–421 (2019).
Guallar, D. et al. RNA-dependent chromatin targeting of TET2 for endogenous retrovirus control in pluripotent stem cells. Nat. Genet. 50, 443–451 (2018).
Kumar, P., Kuscu, C. & Dutta, A. Biogenesis and function of transfer RNA-related fragments (tRFs). Trends Biochem. Sci. 41, 679–689 (2016).
Ivanov, P., Emara, M. M., Villen, J., Gygi, S. P. & Anderson, P. Angiogenin-induced tRNA fragments inhibit translation initiation. Mol. Cell 43, 613–623 (2011).
Sobala, A. & Hutvagner, G. Small RNAs derived from the 5′ end of tRNA can inhibit protein translation in human cells. RNA Biol. 10, 553–563 (2013).
Blanco, S. et al. Stem cell function and stress response are controlled by protein synthesis. Nature 534, 335–340 (2016).
Schorn, A. J., Gutbrod, M. J., LeBlanc, C. & Martienssen, R. LTR-retrotransposon control by tRNA-derived small RNAs. Cell 170, 61–71.e11 (2017).
Chen, Q. et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 351, 397–400 (2016).
Sharma, U. et al. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 351, 391–396 (2016).
Conine, C. C., Sun, F., Song, L., Rivera-Perez, J. A. & Rando, O. J. Small RNAs gained during epididymal transit of sperm are essential for embryonic development in mice. Dev. Cell 46, 470–480.e3 (2018).
Sharma, U. et al. Small RNAs are trafficked from the epididymis to developing mammalian sperm. Dev. Cell 46, 481–494.e6 (2018).
He, C. et al. High-resolution mapping of RNA-binding regions in the nuclear proteome of embryonic stem cells. Mol. Cell 64, 416–430 (2016).
Koh, K. P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Pastor, W. A., Aravind, L. & Rao, A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat. Rev. Mol. Cell Biol. 14, 341–356 (2013).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Friedersdorf, M. B. & Keene, J. D. Advancing the functional utility of PAR-CLIP by quantifying background binding to mRNAs and lncRNAs. Genome Biol. 15, R2 (2014).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Gajjar, M. et al. The p53 mRNA-Mdm2 interaction controls Mdm2 nuclear trafficking and is required for p53 activation following DNA damage. Cancer Cell 21, 25–35 (2012).
Zarnegar, B. J. et al. irCLIP platform for efficient characterization of protein-RNA interactions. Nat. Methods 13, 489–492 (2016).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Hu, X. et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14, 512–522 (2014).
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Huber, S. M., van Delft, P., Tanpure, A., Miska, E. A. & Balasubramanian, S. 2′-O-methyl-5-hydroxymethylcytidine: a second oxidative derivative of 5-methylcytidine in RNA. J. Am. Chem. Soc. 139, 1766–1769 (2017).
Frye, M. & Blanco, S. Post-transcriptional modifications in development and stem cells. Development 143, 3871–3881 (2016).
Schorn, A. J. & Martienssen, R. Tie-Break: host and retrotransposons play tRNA. Trends Cell Biol. 28, 793–806 (2018).
Jin, S. G. et al. Tet3 reads 5-carboxylcytosine through its CXXC domain and is a potential guardian against neurodegeneration. Cell Reports 14, 493–505 (2016).
Beck, D. B. et al. Delineation of a human mendelian disorder of the DNA demethylation machinery: TET3 deficiency. Am. J. Hum. Genet. 106, 234–235 (2020).
Dunn, D. B. Additional components in ribonucleic acid of rat-liver fractions. Biochim. Biophys. Acta 34, 286–288 (1959).
Klagsbrun, M. An evolutionary study of the methylation of transfer and ribosomal ribonucleic acid in prokaryote and eukaryote organisms. J. Biol. Chem. 248, 2612–2620 (1973).
Schopman, N. C., Heynen, S., Haasnoot, J. & Berkhout, B. A miRNA-tRNA mix-up: tRNA origin of proposed miRNA. RNA Biol. 7, 573–576 (2010).
Maute, R. L. et al. tRNA-derived microRNA modulates proliferation and the DNA damage response and is down-regulated in B cell lymphoma. Proc. Natl Acad. Sci. USA 110, 1404–1409 (2013).
Li, Z. et al. Extensive terminal and asymmetric processing of small RNAs from rRNAs, snoRNAs, snRNAs, and tRNAs. Nucleic Acids Res. 40, 6787–6799 (2012).
Yamasaki, S., Ivanov, P., Hu, G. F. & Anderson, P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J. Cell Biol. 185, 35–42 (2009).
Genenncher, B. et al. Mutations in cytosine-5 tRNA methyltransferases impact mobile element expression and genome stability at specific DNA repeats. Cell Reports 22, 1861–1874 (2018).
Goll, M. G. et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398 (2006).
Hussain, S. et al. NSun2-mediated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs. Cell Reports 4, 255–261 (2013).
Helm, M. Post-transcriptional nucleotide modification and alternative folding of RNA. Nucleic Acids Res. 34, 721–733 (2006).
Agris, P. F. Bringing order to translation: the contributions of transfer RNA anticodon-domain modifications. EMBO Rep. 9, 629–635 (2008).
Anderson, J. T. & Wang, X. Nuclear RNA surveillance: no sign of substrates tailing off. Crit. Rev. Biochem Mol. Biol. 44, 16–24 (2009).
Motorin, Y. & Helm, M. tRNA stabilization by modified nucleotides. Biochemistry 49, 4934–4944 (2010).
Lan, J. et al. Functional role of Tet-mediated RNA hydroxymethylcytosine in mouse ES cells and during differentiation. Nat. Commun. 11, 4956 (2020).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Konig, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Schubert, M., Lindgreen, S. & Orlando, L. AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res. Notes 9, 88 (2016).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Wang, L., Feng, Z., Wang, X., Wang, X. & Zhang, X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26, 136–138 (2010).
Liu, M. Y., DeNizio, J. E. & Kohli, R. M. Quantification of oxidized 5-methylcytosine bases and TET enzyme activity. Methods Enzymol. 573, 365–385 (2016).
Kumar, P., Anaya, J., Mudunuri, S. B. & Dutta, A. Meta-analysis of tRNA derived RNA fragments reveals that they are evolutionarily conserved and associate with AGO proteins to recognize specific RNA targets. BMC Biol. 12, 78 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank R. Martienssen for support, encouragement, and discussions; M. Liu and R. Kohli for the kind gift of recombinant TET2 protein; T. Christopher for technical help; G. Xu for his generous gifts of Tet1/2/3-tKO cells; K. Ingvarsdottir and R. Warneford-Thomson for tissue culture help; and S. Erhardt for helpful discussion. R.B. acknowledges support from the NIH (R01GM127408). C.H. was supported in part by the National Natural Science Foundation of China (32070613, 31800687), the Natural Science Foundation of Hunan Province of China (2020JJ4179) and the Fundamental Research Funds for the Central Universities of China (531107051157). B.A.G. was supported in part by the NIH (R01GM110174, R01AI118891 and P01CA196539). A.J.S. acknowledges assistance from the Cold Spring Harbor Laboratory Shared Resources, which are funded in part by the Cancer Center Support Grant (5PP30CA045508). J.E.W. is a Rita Allen Foundation Scholar and is supported by NIH grant R35-GM119735.
Author information
Authors and Affiliations
Contributions
C.H. and R.B. conceived the project and designed the experiments. C.H. generated the ESC lines and performed most of the experiments. J.B. performed rescue experiments and small RNA sequencing. K.A.J. performed all MS under the supervision of B.A.G. A.J.S. sequenced and analyzed small RNAs. J.E.W. helped with RNA biochemistry. C.H. and R.B. wrote the manuscript with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation of epitope-tagged Tet2 alleles and CLIP quantification.
a, Genotype validation for 6xHis–HA knock-in at the Tet2 locus by Sanger sequencing. The targeted allele scheme (top), expected protein and DNA sequence (middle), and sequencing traces (bottom) for the two clones used in subsequent experiments are shown. b, Quantification of Fig. 1D; fluorescence signal (crosslinked RNA) was normalized to WB signal (protein). Bars represent the mean + s.e.m. P-value is from a Student’s t-test.
Extended Data Fig. 2 Additional analyses on TET2 CLIP-seq.
a, Fluorescence image of two CLIP replicates from two cell lines (#65 and #75) used for CLIP-seq library construction. The dashed red boxes indicate the position of the excised bands. Bottom panel, Western blot for HA was used as a loading control (bottom). Uncropped blot images are shown in Supplementary Fig. 1. b, Percentage of transcripts from the indicated classes enriched > 2-fold in TET2 CLIP compared to input (black bars). The non-enriched portion of each class is shown in gray. Only transcripts detected (> 1 read) in at least one replicate were considered. snRNA, small nuclear RNAs; rRNA, ribosomal RNAs; miRNA, micro RNAs; lncRNA, long noncoding RNAs; misc, RNAs not included in the other displayed categories; pseudo, pseudogene-derived RNAs; snoRNA, small nucleolar RNAs; scaRNA, small Cajal body RNAs; mRNA, protein-coding messenger RNAs. c, Same as (B) but considering as enriched only RNAs containing a peak (see text for details) with FDR-corrected P-value < 10−5.
Extended Data Fig. 3 Replicate consistency for the hm5C RIP experiment.
Clustered heatmap showing the Pearson’s correlation coefficients between RPKMs calculated on all annotated genes in input and hm5C RIP for two biological replicates each.
Extended Data Fig. 4 RNA fractionation and mass spectrometry.
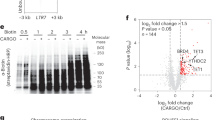
a, Representative gel (6% polyacrylamide, 7 M urea) showing the three RNA fractions analyzed in Fig. 4: total RNA (no fractionation), small RNAs < 200 nts (column-based size selection), and tRNAs (gel purification). The band corresponding to tRNAs (~70 nts) is indicated by the arrow. b, Mass spectrometry chromatograms of nucleosides fragmented into nucleobases. Data were acquired by isolating a precursor ion (nucleoside), fragmenting the precursor ion, and then isolating and detecting a known fragment ion (nucleobase). The top two chromatograms show C (244.093 → 112.050 m/z) and a spiked-in heavy C standard. The bottom two chromatograms show hm5C (256.103 → 142.061 m/z) and a spiked-in heavy hm5C standard (277.103 → 145.061 m/z). The representative chromatograms shown were obtained from the same run on tRNAs from Tet1/2/3 tKO cells transiently transfected with TET2 WT (Fig. 4D). Only one known nucleoside, 5-aminomethyluridine (nm5U), is isobaric with hm5C, and can be easily distinguished from hm5C by retention time. c, Western blot for TET2 comparing WT (+/+) and presumptive KO (–/–) clones as determined by PCR screening and Sanger sequencing. Tubulin is shown as loading control. d, Mass spectrometry quantification of hm5C in size-selected small RNAs < 200 nts from WT (left) Tet2 single KO (middle) and Tet1/2/3 triple KO (right) mESCs. Bars represent mean + s.e.m. ***, P < 0.001. P-values are from one-way ANOVA followed by Holm-Sidak test. Uncropped gel and blot images for a and c are shown in Supplementary Fig. 1.
Extended Data Fig. 5 Additional small RNA sequencing dataset comparing Tet2−/− with E14 ESCs.
a, Coverage of tRNA genes by non-CCA (left) and CCA-containing (right) reads in small RNAs purified from control (E14) or Tet2−/− cells. Plots show the average RPMs. Position of the three types of tRFs discussed in the text are indidated (tRF5, tRF3a, and tRF3b). b, Quantification of (a) but only considering size-filtered reads; 28–35 for tRF5, 17–19 (inclusive of CCA) for tRF3a, and 22 (inclusive of CCA) for tRF3b. c, Differential expression analysis for individual tRFs in E14 and Tet2−/− cells. Estimated (DESeq2) fold changes are plotted on the x axis and the log-converted P-value on the y axis. Blue and red dots highlight individual tRFs that pass an adjusted P-value cutoff of 0.1 and are downregulated or upregulated in the KO, respectively. d, Overlap of TET2-bound tRNAs as determined by CLIP (Fig. 2), and the tRF3a significantly upregulated in Tet2−/− cells as determined in (c). The TET2-bound tRNAs were grouped according to the predicted sequence of the tRF3 produced from them. The P-value was calculated based on the hypergeometric distribution. e, Same as (d) but showing the overlap with tRNAs enriched by hm5C RIP (Fig. 3). f, Comparison of estimated log2(fold-changes) for all tRF3a significantly enriched in Tet2−/− cells in two independent experiments (exp 1, Fig. 5; and exp 2, Extended Data Fig. 5). Replicates are from three independent cell cultures and RNA purifications per genotype.
Extended Data Fig. 6 Examples of tRNAs methylated by NSUN2 and regulated by TET2.
a, Heatmap for the % of unconverted BS-seq reads on tRNAs as reported by Legrand et al.10 (GEO series GSE81825). b, Overlap of tRF3a (top) or tRF3b (bottom) detected as significantly upregulated in Tet2-/− cells compared to WT in Fig. 5 with highly methylated targets of NSUN2 (>75% m5C at NSUN2 sites). P-values are from Fisher’s test comparing overlaps with methylated and unmethylated tRNAs. c, Levels (RPMs) for a tRF3a from LeuCAA tRNAs in WT and Tet2−/− cells. The two plots show data from two independent experimental replicates, corresponding to Fig. 5 (left) and Extended Fig. 5 (right). Mean ± s.e.m. are shown. d, Genomic browser snapshot for CLIP and hm5C at the chr11.tRNA1911-LeuCAA locus. Matching inputs are shown. The y axis represents RPMs. e, Schematic depiction of methylation patterns on chr11.tRNA1911-LeuCAA as determined by BS-seq in Legrand et al.10. The position of m5C in the anticodon and after the variable loop (VL) is indicated by thicker circles and the % of uncovered reads is shown using the same color scale used in (a). f–h, Same as (c–e) but for chr13.tRNA988-SerGCT. i–k, Same as (c–e) but for chr13.tRNA112-SerTGA.
Supplementary information
Supplementary Information
Supplementary Figure 1 (uncropped blots).
Supplementary Table 1
Read numbers for the CLIP experiment.
Supplementary Table 2
Read numbers for the RIP experiment.
Supplementary Table 3
List of tRNAs recovered by TET2 CLIP (Fig. 2) and hm5C RIP (Fig. 3).
Supplementary Table 4
Synthetic nucleotide sequences.
Rights and permissions
About this article
Cite this article
He, C., Bozler, J., Janssen, K.A. et al. TET2 chemically modifies tRNAs and regulates tRNA fragment levels. Nat Struct Mol Biol 28, 62–70 (2021). https://doi.org/10.1038/s41594-020-00526-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-00526-w
This article is cited by
-
Epitranscriptomic modifications in mesenchymal stem cell differentiation: advances, mechanistic insights, and beyond
Cell Death & Differentiation (2024)
-
Roles and regulation of tRNA-derived small RNAs in animals
Nature Reviews Molecular Cell Biology (2024)
-
Hypoxia induces alterations in tRNA modifications involved in translational control
BMC Biology (2023)
-
Coordination of RNA modifications in the brain and beyond
Molecular Psychiatry (2023)
-
Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Nature Communications (2023)