Abstract
The molecular function of Atg9, the sole transmembrane protein in the autophagosome-forming machinery, remains unknown. Atg9 colocalizes with Atg2 at the expanding edge of the isolation membrane (IM), where Atg2 receives phospholipids from the endoplasmic reticulum (ER). Here we report that yeast and human Atg9 are lipid scramblases that translocate phospholipids between outer and inner leaflets of liposomes in vitro. Cryo-EM of fission yeast Atg9 reveals a homotrimer, with two connected pores forming a path between the two membrane leaflets: one pore, located at a protomer, opens laterally to the cytoplasmic leaflet; the other, at the trimer center, traverses the membrane vertically. Mutation of residues lining the pores impaired IM expansion and autophagy activity in yeast and abolished Atg9’s ability to transport phospholipids between liposome leaflets. These results suggest that phospholipids delivered by Atg2 are translocated from the cytoplasmic to the luminal leaflet by Atg9, thereby driving autophagosomal membrane expansion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All relevant data are available upon request to the authors. The cryo-EM density maps have been deposited into the Electron Microscopy Data Bank with accession numbers EMD-30535 (hexamer) and EMD-30545 (trimer). Atomic coordinates are deposited into the wwPDB under accession number PDB 7D0I (hexamer). Uncropped gel and blot images are available in Supplementary Fig. 2. Source data are provided with this paper.
Change history
12 November 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Morishita, H. & Mizushima, N. Diverse cellular roles of autophagy. Annu. Rev. Cell Dev. Biol. 35, 453–475 (2019).
Kirkin, V. & Rogov, V. V. A diversity of selective autophagy receptors determines the specificity of the autophagy pathway. Mol. Cell 76, 268–285 (2019).
Nakatogawa, H. Mechanisms governing autophagosome biogenesis. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-020-0241-0 (2020).
Noda, N. N. & Inagaki, F. Mechanisms of autophagy. Annu. Rev. Biophys. 44, 101–122 (2015).
Fujioka, Y. et al. Phase separation organizes the site of autophagosome formation. Nature 578, 301–305 (2020).
Noda, T. et al. Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the Cvt and autophagy pathways. J. Cell Biol. 148, 465–480 (2000).
Mari, M. et al. An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J. Cell Biol. 190, 1005–1022 (2010).
Yamamoto, H. et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J. Cell Biol. 198, 219–233 (2012).
Suzuki, K., Akioka, M., Kondo-Kakuta, C., Yamamoto, H. & Ohsumi, Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J. Cell Sci. 126, 2534–2544 (2013).
Graef, M., Friedman, J. R., Graham, C., Babu, M. & Nunnari, J. ER exit sites are physical and functional core autophagosome biogenesis components. Mol. Biol. Cell 24, 2918–293 (2013).
Kotani, T., Kirisako, H., Koizumi, M., Ohsumi, Y. & Nakatogawa, H. The Atg2-Atg18 complex tethers pre-autophagosomal membranes to the endoplasmic reticulum for autophagosome formation. Proc. Natl Acad. Sci. USA 115, 10363–10368 (2018).
Gomez-Sanchez, R. et al. Atg9 establishes Atg2-dependent contact sites between the endoplasmic reticulum and phagophores. J. Cell Biol. 217, 2743–2763 (2018).
Osawa, T. et al. Atg2 mediates direct lipid transfer between membranes for autophagosome formation. Nat. Struct. Mol. Biol. 26, 281–288 (2019).
Valverde, D. P. et al. ATG2 transports lipids to promote autophagosome biogenesis. J. Cell Biol. 218, 1787–1798 (2019).
Maeda, S., Otomo, C. & Otomo, T. The autophagic membrane tether ATG2A transfers lipids between membranes. Elife 8, e45777 (2019).
Osawa, T., Ishii, Y. & Noda, N. N. Human ATG2B possesses a lipid transfer activity which is accelerated by negatively charged lipids and WIPI4. Genes Cells 25, 65–70 (2020).
Sharom, F. J. Flipping and flopping–lipids on the move. IUBMB Life 63, 736–746 (2011).
Nagata, S., Sakuragi, T. & Segawa, K. Flippase and scramblase for phosphatidylserine exposure. Curr. Opin. Immunol. 62, 31–38 (2020).
Ploier, B. & Menon, A. K. A fluorescence-based assay of phospholipid scramblase activity. J. Vis. Exp. https://doi.org/10.3791/54635 (2016).
Menon, I. et al. Opsin is a phospholipid flippase. Curr. Biol. 21, 149–153 (2011).
Vehring, S. et al. Flip-flop of fluorescently labeled phospholipids in proteoliposomes reconstituted with Saccharomyces cerevisiae microsomal proteins. Eukaryot. Cell 6, 1625–1634 (2007).
Kihara, A., Noda, T., Ishihara, N. & Ohsumi, Y. Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152, 519–530 (2001).
Cheng, J. et al. Yeast and mammalian autophagosomes exhibit distinct phosphatidylinositol 3-phosphate asymmetries. Nat. Commun. 5, 3207 (2014).
Lai, L. T. F. et al. Subnanometer resolution cryo-EM structure of Arabidopsis thaliana ATG9. Autophagy 16, 575–583 (2020).
Suzuki, S. W. et al. Atg13 HORMA domain recruits Atg9 vesicles during autophagosome formation. Proc. Natl Acad. Sci. USA 112, 3350–3355 (2015).
Tian, W., Chen, C., Lei, X., Zhao, J. & Liang, J. CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res. 46, W363–W367 (2018).
Welter, E., Thumm, M. & Krick, R. Quantification of nonselective bulk autophagy in S. cerevisiae using Pgk1-GFP. Autophagy 6, 794–797 (2010).
Harding, T. M., Morano, K. A., Scott, S. V. & Klionsky, D. J. Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J. Cell Biol. 131, 591–602 (1995).
Guardia, C. M. et al. Structure of human ATG9A, the only transmembrane protein of the core autophagy machinery. Cell Rep. 31, 107837 (2020).
Schutter, M., Giavalisco, P., Brodesser, S. & Graef, M. Local fatty acid channeling into phospholipid synthesis drives phagophore expansion during autophagy. Cell 180, 135–149.e14 (2020).
Brunner, J. D., Schenck, S. & Dutzler, R. Structural basis for phospholipid scrambling in the TMEM16 family. Curr. Opin. Struct. Biol. 39, 61–70 (2016).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Drew, D. et al. GFP-based optimization scheme for the overexpression and purification of eukaryotic membrane proteins in Saccharomyces cerevisiae. Nat. Protoc. 3, 784–798 (2008).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Hamasaki, M., Noda, T. & Ohsumi, Y. The early secretory pathway contributes to autophagy in yeast. Cell Struct. Funct. 28, 49–54 (2003).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Barad, B. A. et al. EMRinger: side chain–directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Grinkova, Y. V., Denisov, I. G. & Sligar, S. G. Engineering extended membrane scaffold proteins for self-assembly of soluble nanoscale lipid bilayers. Protein Eng. Des. Sel. 23, 843–848 (2010).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Ritchie, T. K. et al. in Methods in Enzymology Vol. 464 (ed. Düzgünes, N.) 211–231 (Elsevier, 2009).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Gietz, R. D. & Schiestl, R. H. Large-scale high-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 38–41 (2007).
Noda, T. Viability assays to monitor yeast autophagy. Methods Enzymol. 451, 27–32 (2008).
Ngo, K. X., Kodera, N., Katayama, E., Ando, T. & Uyeda, T. Q. Cofilin-induced unidirectional cooperative conformational changes in actin filaments revealed by high-speed atomic force microscopy. Elife 4, e04806 (2015).
Lazaridis, T. Effective energy function for proteins in lipid membranes. Proteins 52, 176–192 (2003).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jung, J. et al. GENESIS: a hybrid-parallel and multi-scale molecular dynamics simulator with enhanced sampling algorithms for biomolecular and cellular simulations. Wiley Interdiscip. Rev. Comput Mol. Sci. 5, 310–323 (2015).
Mori, T. & Sugita, Y. Implicit micelle model for membrane proteins using superellipsoid approximation. J. Chem. Theory Comput. 16, 711–724 (2020).
Rostislavleva, K. et al. Structure and flexibility of the endosomal Vps34 complex reveals the basis of its function on membranes. Science 350, aac7365 (2015).
Acknowledgements
We thank H. Yamamoto at the University of Tokyo and Y. Ohashi and R.L. Williams at the MRC Laboratory of Molecular Biology for providing plasmids and strains for yeast experiments; Y. Jin for useful advice regarding QF-FRL analyses; T. Ando and Y. Ishii for assistance with protein preparation; Y. Nakada-Nakura, K. Liu and T. Uemura for technical assistance in the generation of antibodies; and A. I. May for proofreading. This work was supported in part by JSPS KAKENHI grant nos. 25111004, 18H03989, 19H05707 (to N.N.N.), 15K21608, 18K06097 (to K.M.), 19K16071 (to T.K.), 19K07265 (to T.T.), (19K06532 to T.M.), 19K16344 (to D.N), 26119006, 19H05645 (to Y.S.), 16H06375, 19H05708 (to Y.O.), 15H05902, 18H04023 (to T.F.), 17H01430, 19H05708 (to H.N.), JST CREST grant no. JPMJCR13M7 (to N.N.N. and H.N.), JPMJCR14M1 (to M.K.), grants from the Takeda Science Foundation (to N.N.N.) and from the Naito Foundation (to N.N.N.), and by Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under grant no. JP19am0101001 (support no. 0053) and 19am0101079 (support no. 0739), and from Research on Development of New Drugs from the AMED.
Author information
Authors and Affiliations
Contributions
K.M. and N.N.N. conceived the project. K.M. purified recombinant proteins and prepared proteoliposomes. K.M., A.T. and M.K. performed cryo-EM experiments. K.M., T.K., Y.O. and H.N. performed yeast experiments. T.T. and T.F. performed quick-freezing and freeze-fracture replica labeling method experiments. D.N. performed HS-AFM observation. T.M. and Y.S. performed MD simulation. N.N. and S.I. prepared antibodies against Atg9. All authors analyzed the data. K.M. and N.N.N. wrote the manuscript with input from all other authors. N.N.N. supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Anke Sparmann and Inês Chen were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
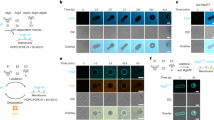
Extended Data Fig. 1 In vitro analysis of lipid scramblase activity of Atg9.
a, b, Dithionite assay. Assay was performed using proteoliposomes (green) in the existence of ATP/Mg (red) or calcium ion (cyan) (a) and proteoliposomes containing NBD-C6-PC, NBD-C6-PE, or NBD-C6-PS (b). FI(t) and FI(0) indicate the fluorescence intensity at each time and 0, respectively. c, SDS-PAGE of the purified PI3K complex I stained by Coomassie Brilliant Blue. d, Left, results of the floatation assay of liposomes with either 5% PI3P (PC/PI/PS/PE/PI3P) or 18% PI (PC/PI/PS/PE) detected by p40phox PX-EYFP. Fluorescence intensity (FI) of p40phox PX-EYFP was detected at Fraction 1 only when PI3P liposomes were used, indicating that p40phox PX-EYFP specifically recognizes PI3P liposomes but not PI liposomes. Right, results of the floatation assay of 18% PI liposomes treated with PI3K and various concentrations of ATP. As a positive control, 5% PI3P liposomes were also analyzed. The graph shows the fluorescence intensity at Fraction 1. Fluorescence intensity was proportional to the ATP concentration, indicating that catalytic conversion of PI to PI3P proceeded in liposomes.
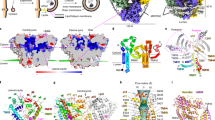
Extended Data Fig. 2 Cryo-EM analysis of Atg9.
a, SEC profiles of SpAtg9-GFP fusion (black and red) or SpAtg9 alone (blue). b, Summary of cryo-EM analysis of SpAtg9 hexamer. A representative micrograph of SpAtg9 (upper left), local resolution map (lower left), workflow of cryo-EM (upper middle), Fourier shell correlation curves (lower middle), and 2D class average images (upper right), where selected particle class was surrounded by a red line. Lower right, representative density map of some regions with a structural model. Each amino-acid sequence is also provided. c, Summary of cryo-EM analysis of SpAtg9 trimer. Workflow of cryo-EM (left), Fourier shell correlation curves (middle), and 2D class average images (right). d, Superimposed cryo-EM map of hexamer (resolution 3.0 Å, colored yellow) and trimer (resolution 4.7 Å, colored grey). Protomer boundaries are shown with blue, red, and black lines. e, comparison of the cryo-EM map between plant Atg9 (EMD-9681) (top) and yeast trimer (resolution 4.7 Å) (bottom). f, DISOPRED result of SpAtg9 (upper) and schematic drawing of the domain organization of Atg9 homologs (bottom), where structured and disordered regions predicted by DISOPRED are shown with a square and a black line, respectively. Arrow line (cyan) indicates the region whose model was built by cryo-EM experiment.
Extended Data Fig. 3 Detailed interactions observed in the cryo-EM structure.
a, Close−up view of H3 and H6. Prolines responsible for kinking helices are indicated. b-e, Close−up view of the interactions observed in ARCH. f, H4 and H5 bound to adjacent protomers within a trimer. One protomer is shown in a ribbon model whereas the other two protomers are shown in a surface model. g, Schematic drawing of the trimer structure. h, Close-up view of the interactions observed between H1, H3, and H4. i, Close-up view of the trimer-trimer interface observed in the hexamer structure (left) and proposed transmembrane regions (TM) within the hexamer (right).
Extended Data Fig. 4 Characterization of Atg9 trimer.
a, Thickness of lipid bilayer membrane on mica measured by HS-AFM. b, 2D class average images of Atg9-nanodiscs recognized by Fab. Scale bar, 50 Å. c, Density map of LMNG at the LP. Two LMNG molecules are colored differently. d, Comparison of the VP radius between cryo-EM and MD models. e, Comparison of different classes of trimer Atg9 structure. Left and right show density map at 5.3 and 6.2 Å resolution, on which structural model of SpAtg9 is superimposed by real-space refinement. The distance is between Cα atoms of Leu539 located at VP. f, Comparison of the SEC profiles between WT and Hexamer mutant (I641D, V643A, V654A, and F655A) of SpAtg9 using a Superose 6 increase 10/300GL column.
Extended Data Fig. 5 Mutational analysis of Atg9.
a, Circular dichroism (CD) spectra of WT and mutant forms of SpAtg9. b, SEC profiles of WT and mutant forms of SpAtg9. Blue bar indicates the fraction used for in vitro analysis. c, d, Fluorescence images of colocalization of Atg17 puncta with Atg9-GFP mutants (c) and with Atg2-mNeongreen in cells expressing Atg9 mutants (d).
Extended Data Fig. 6 Random mutational analysis of Atg9.
a, Ape1 maturation assay of atg9Δ cells expressing Atg9 with random mutations. b, Left table shows the random mutation sites in ScAtg9 that partially or severely impaired Ape1 maturation and their equivalent residues in SpAtg9. Right figure shows the location of residues at LP and VP that were shown to be important for autophagy by random mutagenesis experiments.
Supplementary information
Supplementary Information
Supplementary Figures 1 and 2, Supplementary Notes 1 and 2 and Supplementary Tables 1 and 2.
Supplementary Video 1
Visualization of the cryo-EM map of the SpAtg9 hexamer with the final atomic model.
Supplementary Video 2
HS-AFM observation of the cytoplasmic side of Atg9 embedded in the lipid bilayer membrane. Z-scale, 0–6.5 nm.
Supplementary Video 3
HS-AFM observation of the luminal side of Atg9 embedded in the lipid bilayer membrane. Z-scale, 0–1.9 nm.
Supplementary Video 4
Density map of LMNG bound to Atg9, related to Fig. 3a and Extended Data Fig. 4c.
Supplementary Video 5
Mutation sites in VP, related to Fig. 3b.
Supplementary Video 6
Vacuolar protease−deficient cells expressing Atg9 mutants were treated with rapamycin for 24 h before autophagic body accumulation in these cells was observed by phase contrast microscopy. Time-lapse images were taken at 32.65 ms per frame.
Supplementary Video 7
Observation of Atg9 vesicles in yeast cells. The numbers correspond to mutant numbers (mutant number 72 corresponds to the Wall mutant).
Source data
Source Data Fig. 1
Graph source data
Source Data Fig. 2
Graph source data
Source Data Fig. 3
Graph source data
Source Data Fig. 4
Graph source data
Source Data Fig. 5
Graph source data
Rights and permissions
About this article
Cite this article
Matoba, K., Kotani, T., Tsutsumi, A. et al. Atg9 is a lipid scramblase that mediates autophagosomal membrane expansion. Nat Struct Mol Biol 27, 1185–1193 (2020). https://doi.org/10.1038/s41594-020-00518-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-00518-w
This article is cited by
-
Optineurin provides a mitophagy contact site for TBK1 activation
The EMBO Journal (2024)
-
Complete set of the Atg8–E1–E2–E3 conjugation machinery forms an interaction web that mediates membrane shaping
Nature Structural & Molecular Biology (2024)
-
Unexpected inhibition of the lipid kinase PIKfyve reveals an epistatic role for p38 MAPKs in endolysosomal fission and volume control
Cell Death & Disease (2024)
-
Lipid droplets and cellular lipid flux
Nature Cell Biology (2024)
-
Experimental determination and mathematical modeling of standard shapes of forming autophagosomes
Nature Communications (2024)