Abstract
Aberrantly stalled ribosomes initiate the ribosome-associated quality control (RQC) and mRNA surveillance pathways for the degradation of potentially toxic peptides and faulty mRNAs. During RQC, ANKZF1 (yeast Vms1p) releases ubiquitinated nascent proteins from 60S ribosomal subunits for proteasomal degradation. Here, we use a cell-free system to show that ANKZF1 and Vms1p sever polypeptidyl-tRNAs on RQC complexes by precisely cleaving off the terminal 3′CCA nucleotides universal to all tRNAs. This produces a tRNA fragment that cannot be aminoacylated until its 3′CCA end is restored. The recycling of ANKZF1-cleaved tRNAs is intact in the mammalian cytosol via a two-step process that requires the removal of a 2′,3′-cyclic phosphate and TRNT1, the sole CCA-adding enzyme that mediates tRNA biogenesis in eukaryotes. TRNT1 also discriminates between properly folded tRNA substrates and aberrant tRNA substrates, selectively tagging the latter for degradation. Thus, ANKZF1 liberates peptidyl-tRNAs from stalled ribosomes such that the tRNA is checked in an obligate way for integrity before reentry into the translation cycle.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary information files.
References
Chu, J. et al. A mouse forward genetics screen identifies LISTERIN as an E3 ubiquitin ligase involved in neurodegeneration. Proc. Natl Acad. Sci. USA 106, 2097–2103 (2009).
Choe, Y.-J. et al. Failure of RQC machinery causes protein aggregation and proteotoxic stress. Nature 531, 191–195 (2016).
Ishimura, R. et al. Ribosome stalling induced by mutation of a CNS-specific tRNA causes neurodegeneration. Science 345, 455–459 (2014).
Shoemaker, C. J., Eyler, D. E. & Green, R. Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330, 369–372 (2010).
Shoemaker, C. J. & Green, R. Translation drives mRNA quality control. Nat. Struct. Mol. Biol. 19, 594–601 (2012).
Doma, M. K. & Parker, R. Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440, 561–564 (2006).
van Hoof, A., Frischmeyer, P. A., Dietz, H. C. & Parker, R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295, 2262–2264 (2002).
Frischmeyer, P. A. et al. An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295, 2258–2261 (2002).
Brandman, O. et al. A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 151, 1042–1054 (2012).
Shao, S., von der Malsburg, K. & Hegde, R. S. Listerin-dependent nascent protein ubiquitination relies on ribosome subunit dissociation. Mol. Cell 50, 637–648 (2013).
Bengtson, M. H. & Joazeiro, C. A. P. Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 467, 470–473 (2010).
Pisareva, V. P., Skabkin, M. A., Hellen, C. U. T., Pestova, T. V. & Pisarev, A. V. Dissociation by Pelota, Hbs1 and ABCE1 of mammalian vacant 80S ribosomes and stalled elongation complexes. EMBO J. 30, 1804–1817 (2011).
Shao, S., Brown, A., Santhanam, B. & Hegde, R. S. Structure and assembly pathway of the ribosome quality control complex. Mol. Cell 57, 433–444 (2015).
Shen, P. S. et al. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains. Science 347, 75–78 (2015).
Lyumkis, D. et al. Structural basis for translational surveillance by the large ribosomalsubunit-associated protein quality control complex. Proc. Natl Acad. Sci. USA 111, 15981–15986 (2014).
Verma, R., Oania, R. S., Kolawa, N. J. & Deshaies, R. J. Cdc48/p97 promotes degradation of aberrant nascent polypeptides bound to the ribosome. eLife 2, e00308 (2013).
Defenouillère, Q. et al. Cdc48-associated complex bound to 60S particles is required for the clearance of aberrant translation products. Proc. Natl Acad. Sci. USA 110, 5046–5051 (2013).
Defenouillère, Q. et al. Rqc1 and Ltn1 prevent C-terminal alanine-threonine tail (CAT-tail)-induced protein aggregation by efficient recruitment of Cdc48 on stalled 60S subunits. J. Biol. Chem. 291, 12245–12253 (2016).
Osuna, B. A., Howard, C. J., Kc, S., Frost, A. & Weinberg, D. E. In vitro analysis of RQC activities provides insights into the mechanism and function of CAT tailing. eLife 6, e27949 (2017).
Kuroha, K., Zinoviev, A., Hellen, C. U. T. & Pestova, T. V. Release of ubiquitinated and non-ubiquitinated nascent chains from stalled mammalian ribosomal complexes by ANKZF1 and Ptrh1. Mol. Cell 72, 286–302.e8 (2018).
Kostova, K. K. et al. CAT-tailing as a fail-safe mechanism for efficient degradation of stalled nascent polypeptides. Science 357, 414–417 (2017).
Izawa, T., Park, S.-H., Zhao, L., Hartl, F. U. & Neupert, W. Cytosolic protein Vms1 links ribosome quality control to mitochondrial and cellular homeostasis. Cell 171, 890–903.e18 (2017).
Yonashiro, R. et al. The Rqc2/Tae2 subunit of the ribosome-associated quality control (RQC) complex marks ribosome-stalled nascent polypeptide chains for aggregation. eLife 5, e11794 (2016).
Heo, J.-M. et al. A stress-responsive system for mitochondrial protein degradation. Mol. Cell 40, 465–480 (2010).
Verma, R. et al. Vms1 and ANKZF1 peptidyl-tRNA hydrolases release nascent chains from stalled ribosomes. Nature 557, 446–451 (2018).
Zurita Rendón, O. et al. Vms1p is a release factor for the ribosome-associated quality control complex. Nat. Commun. 9, 2197 (2018).
Xiong, Y. & Steitz, T. A. A story with a good ending: tRNA 3′-end maturation by CCA-adding enzymes. Curr. Opin. Struct. Biol. 16, 12–17 (2006).
Wilusz, J. E., Whipple, J. M., Phizicky, E. M. & Sharp, P. A. tRNAs marked with CCACCA are targeted for degradation. Science 334, 817–821 (2011).
Shao, S. & Hegde, R. S. Reconstitution of a minimal ribosome-associated ubiquitination pathway with purified factors. Mol. Cell 55, 880–890 (2014).
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011).
Yang, W. Nucleases: diversity of structure, function and mechanism. Q. Rev. Biophys. 44, 1–93 (2011).
Honda, S., Morichika, K. & Kirino, Y. Selective amplification and sequencing of cyclic phosphate-containing RNAs by the cP-RNA-seq method. Nat. Protoc. 11, 476–489 (2016).
Schürer, H., Lang, K., Schuster, J. & Mörl, M. A universal method to produce in vitro transcripts with homogeneous 3′ ends. Nucleic Acids Res. 30, e56 (2002).
Czech, A., Wende, S., Mörl, M., Pan, T. & Ignatova, Z. Reversible and rapid transfer-RNA deactivation as a mechanism of translational repression in stress. PLoS Genet. 9, e1003767 (2013).
Zuo, Y. & Deutscher, M. P. The physiological role of RNase T can be explained by its unusual substrate specificity. J. Biol. Chem. 277, 29654–29661 (2002).
Nagaike, T. et al. Identification and characterization of mammalian mitochondrial tRNA nucleotidyltransferases. J. Biol. Chem. 276, 40041–40049 (2001).
Feng, Q. & Shao, S. In vitro reconstitution of translational arrest pathways. Methods 137, 20–36 (2018).
Zinder, J. C. & Lima, C. D. Targeting RNA for processing or destruction by the eukaryotic RNA exosome and its cofactors. Genes Dev. 31, 88–100 (2017).
Shigematsu, M., Kawamura, T. & Kirino, Y. Generation of 2′,3′-cyclic phosphate-containing RNAs as a hidden layer of the transcriptome. Front. Genet. 9, 562 (2018).
Jilani, A. et al. Molecular cloning of the human gene, PNKP, encoding a polynucleotide kinase 3′-phosphatase and evidence for its role in repair of DNA strand breaks caused by oxidative damage. J. Biol. Chem. 274, 24176–24186 (1999).
Karimi-Busheri, F. et al. Molecular characterization of a human DNA kinase. J. Biol. Chem. 274, 24187–24194 (1999).
Mayer, M., Schiffer, S. & Marchfelder, A. tRNA 3′ processing in plants: nuclear and mitochondrial activities differ. Biochemistry 39, 2096–2105 (2000).
Phizicky, E. M., Schwartz, R. C. & Abelson, J. Saccharomyces cerevisiae tRNA ligase. Purification of the protein and isolation of the structural gene. J. Biol. Chem. 261, 2978–2986 (1986).
Popow, J. et al. HSPC117 is the essential subunit of a human tRNA splicing ligase complex. Science 331, 760–764 (2011).
Phizicky, E. M. & Hopper, A. K. tRNA biology charges to the front. Genes Dev. 24, 1832–1860 (2010).
Thompson, D. M., Lu, C., Green, P. J. & Parker, R. tRNA cleavage is a conserved response to oxidative stress in eukaryotes. RNA 14, 2095–2103 (2008).
Fu, H. et al. Stress induces tRNA cleavage by angiogenin in mammalian cells. FEBS Lett. 583, 437–442 (2009).
Yamasaki, S., Ivanov, P., Hu, G.-F. & Anderson, P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J. Cell Biol. 185, 35–42 (2009).
Cole, S. E., LaRiviere, F. J., Merrikh, C. N. & Moore, M. J. A convergence of rRNA and mRNA quality control pathways revealed by mechanistic analysis of nonfunctional rRNA decay. Mol. Cell 34, 440–450 (2009).
Brown, A., Shao, S., Murray, J., Hegde, R. S. & Ramakrishnan, V. Structural basis for stop codon recognition in eukaryotes. Nature 524, 493–496 (2015).
Acknowledgements
We thank A.E. Johnson, Y. Miao and Y. Shao for reagents and advice for producing tRNAs; J. Bridgers for help with the initial tRNA experiments; T. Guettler and D. Görlich for the SuperTEV expression plasmid; and R.S. Hegde, T.A. Rapoport, J.W. Harper, A. Brown, S. Juszkiewicz and Shao Laboratory members for useful discussions. This work was supported by Harvard Medical School startup funds, a Richard and Susan Smith Family Award for Excellence in Biomedical Research, a Charles H. Hood Foundation Child Health Research Award, and the Vallee Scholars Program (to S.S.), a NDSEG predoctoral fellowship (V.C.), the Jane Coffin Childs Memorial Fund for Medical Research (61-1681 to Q.F.), and an American Heart Association postdoctoral fellowship (19POST34400009 to M.J.M.).
Author information
Authors and Affiliations
Contributions
M.C.J.Y. and S.S. designed, performed and analyzed the experiments with help from A.F.A.K, Q.F., V.C. and M.J.M. S.S. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Reconstitution of ANKZF1 activity with purified factors.
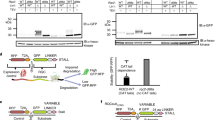
Coomassie stain of the SDS-PAGE gel analyzed by autoradiography in a, Fig. 1b and b, Fig. 1c. Individual recombinant proteins are indicated. c, Full-sized autoradiography (top) and Coomassie stained (bottom) SDS-PAGE gel in Fig. 1d, with cropped area indicated, demonstrating the effect of RNase A on Listerin-ubiquitinated substrates treated without or with ANKZF1. split. fact. – ribosome splitting factors (50 nM Hbs1L, 50 nM Pelota, 100 nM ABCE1); ubiq. fact. – ubiquitination factors (75 nM E1, 250 nM E2, 5 nM Listerin, 10 μM ubiquitin); CK – creatine kinase, part of an energy regenerating system; Ub – ubiquitin; NC-tRNA – nascent chain-tRNA; NC* - ANKZF1-cleaved nascent chain; NC – nascent chain; poly-Ub – poly-ubiquitinated nascent chain.
Supplementary Fig. 2 ANKZF1 releases nascent proteins from ribosomes.
5 nM of purified 80S ribosome-nascent protein complexes (RNCs, as in Fig. 1a) containing radiolabeled nascent protein were incubated with an energy regenerating system, ribosome splitting factors (50 nM Hbs1L, 50 nM Pelota, 100 nM ABCE1), 10 nM NEMF, 75 nM E1, 250 nM E2, 10 μM ubiquitin, and the indicated combinations of 5 nM Listerin and/or 25 nM ANKZF1. 20 μL reactions were size fractionated on 200 μL 10-50% sucrose gradients. Eleven 20 μL fractions (fract.) collected from the top of the gradients were analyzed directly by SDS-PAGE and autoradiography. Migration of radiolabeled nascent polypeptide chain (NC), nascent chain-tRNA (NC-tRNA), ANKZF1-cleavage product (NC*; red arrow) and ubiquitinated nascent proteins (poly-Ub) are indicated. In the absence of ANKZF1, Listerin-mediated ubiquitination occurs on 60S ribosomal subunits (blue arrow), while ANKZF1-cleaved products are released from ribosomal to soluble (sol.) fractions (right gels).
Supplementary Fig. 3 ANKZF1 selectively acts on 60S–RQC complexes.
a, 5 nM 80S RNCs were incubated with energy, 50-100 nM splitting factors, 10 nM NEMF, 12.5 nM wildtype (WT) or the indicated ANKZF1 mutants without (left) or with (right) ubiquitination reagents. Reactions were analyzed by SDS-PAGE and autoradiography (top), and a portion of the Coomassie stained gel is shown (bottom). This demonstrates that mutations in the putative catalytic loop of ANKZF1 abolishes nascent protein release (red arrow). b, 80S RNCs were incubated with energy, splitting factors, and either NEMF or eIF6. Schemes of how NEMF and eIF6 prevent 40S rejoining are shown. 200 μL of these splitting reactions were separated by size into 25 fractions on a 4.8 mL 10–30% sucrose gradient. Fractions containing 60S or 80S ribosomal subunits were pooled and incubated without (left) or with (right) 25 nM ANKZF1. Note that only nascent polypeptidyl-tRNA on 60S ribosomal subunits, but not 80S ribosomes, are cleaved by ANKZF1. eIF6, which does not bind directly to peptidyl-tRNA at the 60S ribosomal subunit interface as NEMF does, is less effective at mediating ANKZF1 activity. c, Cell-free translation reactions of model substrates without (∆VHP) or with the autonomously folding villin-headpiece domain (VHP) were separated on a 10–50% sucrose gradient. Short nascent proteins without folded domains ‘slip out’ of ribosomes upon splitting. Soluble (fractions 1-3) and ribosomal (fractions 5–8) fractions were pooled and incubated with 25 nM ANZKF1. Reactions were analyzed by SDS-PAGE and autoradiography (top) and Coomassie staining (bottom) This demonstrates that ANKZF1 does not act on soluble NC-tRNA or ribosome-associated NC-tRNA in the absence of ribosome splitting.
Supplementary Fig. 4 ANKZF1 and Vms1p cleave peptidyl-tRNA.
a, 5 nM RNCs were incubated with energy, 50–100 nM splitting factors, 10 nM NEMF, and 25 nM ANKZF1 or 25 nM (1X) or 125 nM (5X) of wildtype (WT) or mutant Vms1p, the yeast homolog of ANKZF1. Reactions were analyzed by SDS-PAGE (top) or NuPAGE (bottom) and autoradiography. Note that yeast Vms1p generates the same cleavage product as ANKZF1 and also relies on conserved residues in the catalytic loop. In addition, the lower pH of NuPAGE gels better preserves peptidyl-tRNA bonds which are susceptible to hydrolysis in basic Tris-tricine gels. Thus, for experiments to analyze ANKZF1- and Vms1p-generated products (Figs. 2–4), we switched primarily to NuPAGE to facilitate detection of tRNA-associated products, and to using 125 nM Vms1p, which we were able to obtain at higher concentrations and purities than ANKZF1, to optimize cleavage efficiency. b, Coomassie stain of purified Vms1p proteins. c, 5 nM purified 80S RNCs were incubated with the indicated components (12.5 nM ANKZF1, 50 μg/mL RNase A, 1 mM puromycin, 1 μM eRF1). Red arrow depicts ANKZF1 cleavage product (NC*), which requires ribosome splitting and NEMF. NC* is larger than the expected size of nascent chain (NC) produced by RNase A or puromycin (puro.) treatment. NC-tRNA—nascent polypeptidyl-tRNA, eRF1—eukaryotic release factor 1. Note that the translation termination factor eRF1 is not expected to act catalytically on these substrates due to the lack of a stop codon.
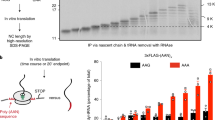
Supplementary Fig. 5 tRNALeu is selectively incorporated into RNCs.
a, TBE-urea-PAGE and SYBR Gold comparison of transcribed leucyl-tRNA (tRNALeu) containing either the UAA (tRNALeu(UAA)) or the CAG (tRNALeu(CAG)) anticodon, and total mammalian liver tRNAs. b, Cell-free translation reactions of radiolabeled protein containing three consecutive rare leucine UUA codons37 without or with total mammalian liver tRNAs or leucyl-tRNAs with the indicated anticodon. Translation stalls at the rare leucine codons, generating a truncated nascent chain (sNC), unless reactions are supplemented with leucyl-tRNA with the cognate UAA codon (which is also present in total liver tRNAs) to rescue translation of the full-length nascent chain (FL NC). Note that tRNALeu(UAA), but not tRNALeu(CAG), rescues translation of FL NC. c, The indicated radiolabeled tRNAs were transcribed and analyzed by TBE-urea-PAGE and autoradiography. d, Truncated mRNAs ending in the indicated leucine codon (UUA or CUG) were translated without additional tRNA in the presence of 35S-methionine (S), or in the presence of 33P-radiolabeled leucyl-tRNA (P) with the indicated anticodon. Reactions were directly analyzed by NuPAGE and autoradiography (left) or first subjected to denaturing immunoprecipitations against an N-terminal Flag epitope tag (Flag IPs) encoded in the nascent protein sequence (right). This demonstrates that radiolabeled leucyl-tRNA specifically incorporates into ribosome-nascent protein complexes (RNCs) stalled at the cognate codon. e, Cell-free translation reactions of a truncated mRNA ending in two consecutive rare UAA codons (as in Fig. 2a) in an endogenous rabbit reticulocyte translation system supplemented with no additional tRNA, 0.2 mg/mL pig liver tRNAs, or 0.5% (v/v) of nonradiolabeled or 33P-labeled tRNALeu. Reactions lacking radiolabeled tRNALeu contained 35S-methionine to radiolabel the nascent protein. Reactions were directly analyzed before or after RNase A treatment by NuPAGE and autoradiography to compare labeling efficiencies of the nascent chain-tRNA (NC-tRNA, red arrow). sNC-tRNA refers to the stalled products on the rare UAA codons in the absence of tRNA supplementation. f, Cell-free translation reactions as in a containing 35S-methionine and nonradiolabeled tRNALeu (top) or cold methionine and radiolabeled tRNALeu (bottom) were separated on a 10-50% sucrose gradient. Eleven fractions were collected from the top and analyzed by NuPAGE and autoradiography. This demonstrates that radiolabeled NC-tRNA products are ribosome-associated, while free tRNA remains soluble.
Supplementary Fig. 6 Isolation and Vms1p cleavage of RNCs with radiolabeled peptidyl-tRNALeu.
a, TBE-urea-PAGE and SYBR Gold (top) or autoradiography (bottom) analysis of tRNALeu transcribed with either radiolabeled CTP or UTP. b, Purification of stalled 80S ribosome-nascent chain complexes (RNCs) from cell-free translation reactions of the truncated mRNA depicted in Fig. 2a with either C- or U-labeled tRNALeu. IVT—total in vitro translation reaction; HS-RNCs—high-salt-washed RNCs; FT—flow-through; Elu—elution. c, 35S-methionine labeled RNCs generated using the same model mRNA as in b with total liver tRNAs were incubated with energy, 50–100 nM ribosome splitting factors, and 10 nM NEMF and 125 nM wildtype (WT) or R288A (mut.) Vms1p as indicated. Reactions were directly analyzed before or after RNase A treatment by NuPAGE and autoradiography. This demonstrates that the requirements for Vms1p activity on RNCs containing peptidyl-tRNALeu are identical to those containing peptidyl-tRNAVal (Fig. 1). d, Coomassie stain of the SDS-PAGE gel analyzed in Fig. 2b. Note that 20-fold less of the reactions containing 35S-methionine-labeled RNCs were loaded to roughly normalize the autoradiography signals. e, Radiolabeled tRNALeu at approximately one (1X) or five (5X) times the concentration isolated from RNCs (for example Fig. 2c) was incubated with 125 nM Vms1p without or with an energy regenerating system (ERS). Without being incorporated into RNCs, there is no noticeable Vms1p cleavage of free tRNAs. Transcribed tRNALeu with (FL) or without (ΔCCA) the 3′CCA nucleotides are loaded as size markers.
Supplementary Fig. 7 Reconstitution of TRNT1-mediated tRNA recycling.
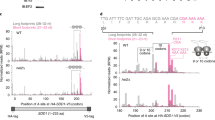
a, Full-length (FL) tRNALeu or tRNALeu lacking three (ΔCCA, red) or four (ΔACCA, dark blue) 3′ terminal nucleotides were transcribed in vitro and treated with 0.05 U/μL Exonuclease T (Exo T), followed by 100 nM recombinant TRNT1 as indicated. Reactions were analyzed by TBE-urea-PAGE and SYBR Gold staining. Note that Exonuclease T removes a single nucleotide from FL (lane 2, light blue arrow) and ΔCCA tRNALeu (lane 7, dark blue arrow), consistent with its activity as a single-stranded 3′ to 5′ exonuclease that is inhibited by consecutive cytidine nucleotides. The two C nucleotides of the 3′CCA prevent further digestion of FL tRNALeu, while basepairing (Fig. 2a) prevents further digestion of ΔCCA tRNALeu. TRNT1 repairs tRNAs lacking one (compare lanes 2 and 4) or three (compare lanes 6 and 8) 3′ nucleotides, but not those lacking four (lanes 9, 13, and 14, dark blue arrows). The slight repair seen in lane 13 is from the small amount of product lacking three 3′ terminal nucleotides (red arrow, lane 11) generated by heterogeneous transcription of the ΔACCA tRNALeu construct without Exonuclease T digestion. b, Vms1p cleavage reactions (total) of non-radiolabeled RNCs ending in valyl-tRNA (tRNAVal) as in Supplementary Fig. 4a were separated on a 10–50% sucrose gradient. Soluble (sol.) and ribosomal (ribo.) fractions were pooled and analyzed by immunoblotting against the Flag-tagged nascent protein (top) or subjected to RNA extraction, TBE-urea-PAGE, and SYBR Gold staining (middle). The extracted RNAs were also used in TRNT1 repair reactions (as in Fig. 3b) with T4 PNK and radiolabeled CTP, and analyzed by TBE-urea-PAGE and autoradiography (bottom). Note that the tRNA fragment generated by Vms1p is exclusively in the soluble fraction, indicating that it is released from RNCs after cleavage. c, HEK293T cells were harvested (total) and sequentially fractionated with 100 μg/mL digitonin to isolate cytosol and 1% Triton X-100 to solubilize organellar membranes. Individual fractions were analyzed by SDS-PAGE and immunoblotting for the indicated components. TOM20 is a mitochondrial outer membrane protein. d, Coomassie stain of purified TRNT1 lacking the N-terminal mitochondrial transit peptide. e, The indicated nonradiolabeled transcribed tRNALeu constructs were incubated with TRNT1 in the presence of radiolabeled CTP or UTP. This demonstrates that cytidine is specifically incorporated into TRNT1-repaired products. f, Non-radiolabeled RNCs ending in tRNAVal were subjected to cleavage and TRNT1 repair reactions exactly as in Fig. 3c, except with mammalian ANKZF1 instead of yeast Vms1p. g, Side-by-side cleavage reactions of 35S-radiolabeled RNCs ending in leucyl-tRNA (tRNALeu) with mammalian ANKZF1 or yeast Vms1p, demonstrating that both generate the identical NC* product sensitive to RNase A (left). RNAs were extracted from these reactions (right) and analyzed directly (top) or subject to TRNT1 repair reactions in the presence of radiolabeled CTP with or without T4 PNK (bottom). Note that the Vms1p and ANKZF1 lanes are switched between the protein (left) and RNA (right) gels, but that both generate the same tRNA fragment that is selectively repaired by TRNT1 only after the removal of the 2′,3′-cyclic phosphate by T4 PNK. h, Coomassie stain of SDS-PAGE gels analyzed in Fig. 3e.
Supplementary Fig. 8 tRNA recycling in the mammalian cytosol.
a, TBE-urea-PAGE and SYBR Gold (left) or autoradiography (right) analysis of transcription reactions of FL-HDV and ΔCCA tRNALeu (ΔCCA-HDV) constructs containing a 3′ hepatitis delta virus (HDV) ribozyme (see Fig. 4a). The unprocessed, 5′ tRNA fragments containing a 2′,3′-cyclic phosphate, and 3′ HDV fragments are indicated. b, Radiolabeled ΔCCA-HDV was treated with 0.12 U/μL T4 PNK followed by 100 nM TRNT1 as indicated in the presence of 0.1 mM NTPs. Note that, like Vms1p-cleaved tRNAs, CCA addition by TRNT1 only occurs after T4 PNK treatment. This contrasts with transcribed tRNAs, which are directly modified by TRNT1 (see Fig. 3a and Supplementary Fig. 7a). c, Radiolabeled ΔCCA-HDV was incubated with increasing amounts of hypotonic cytosolic lysate isolated from HEK293 cells and analyzed by TBE-urea-PAGE and autoradiography, demonstrating that recycling of ΔCCA-HDV to full-length (FL) tRNALeu is intact in these cytosolic lysates. d, SYBR Gold stain of the TBE-urea-PAGE gels analyzed by autoradiography in Fig. 4c. e, Cell-free translation reactions of an mRNA containing three rare UUA leucine codons in endogenous rabbit reticulocyte lysate supplemented with the indicated tRNAs, demonstrating that FL-HDV and ΔCCA-HDV rescue translation similarly as transcribed tRNAs with 3′-hydroxyl groups. f, Cell-free translation reactions of a truncated mRNA substrate ending in two rare UUA leucine codons (as in Fig. 2a). Reactions were in an endogenous rabbit reticulocyte lysate system with 35S-methionine and no exogenous tRNAs, or with cold methionine and the indicated radiolabeled tRNAs. Reactions were directly analyzed by NuPAGE and autoradiography before (left) or after (right) immunoprecipitations (Flag IPs) against the N-terminal Flag tag encoded in the nascent protein. This demonstrates that the radiolabeled tRNAs are incorporated into ribosome-nascent protein complexes (RNCs). g, FL-HDV was incubated with the indicated 293 lysates at physiological or high pH (by addition of NaOH) to observe additional modifications. The high pH, which would hydrolyze amino acid-tRNA ester bonds, rules out the possibility that the higher molecular weight bands are due to aminoacylation. h, ΔCCA-HDV was incubated with rabbit reticulocyte lysate (RRL) or HEK293 cytosol and directly analyzed by TBE-urea-PAGE and autoradiography, demonstrating a higher rate of additional modifications by HEK293 lysate. i, Immunoblotting and Ponceau stain analysis of cytosolic lysates isolated from HEK293T cells knocked down for the indicated proteins used in Fig. 4e. APLF is a control protein.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Supplementary Dataset 1
Rights and permissions
About this article
Cite this article
Yip, M.C.J., Keszei, A.F.A., Feng, Q. et al. Mechanism for recycling tRNAs on stalled ribosomes. Nat Struct Mol Biol 26, 343–349 (2019). https://doi.org/10.1038/s41594-019-0211-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0211-4
This article is cited by
-
Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Nature Communications (2022)
-
eIF6 rebinding dynamically couples ribosome maturation and translation
Nature Communications (2022)
-
Identification of a novel trigger complex that facilitates ribosome-associated quality control in mammalian cells
Scientific Reports (2020)
-
Structure and function of Vms1 and Arb1 in RQC and mitochondrial proteome homeostasis
Nature (2019)
-
tRNA recycling on stalled ribosomes
Nature Structural & Molecular Biology (2019)