Abstract
Germ cells manifest a unique gene expression program and regain totipotency in the zygote. Here, we perform Hi-C analysis to examine 3D chromatin organization in male germ cells during spermatogenesis. We show that the highly compartmentalized 3D chromatin organization characteristic of interphase nuclei is attenuated in meiotic prophase. Meiotic prophase is predominated by short-range intrachromosomal interactions that represent a condensed form akin to that of mitotic chromosomes. However, unlike mitotic chromosomes, meiotic chromosomes display weak genomic compartmentalization, weak topologically associating domains, and localized point interactions in prophase. In postmeiotic round spermatids, genomic compartmentalization increases and gives rise to the strong compartmentalization seen in mature sperm. The X chromosome lacks domain organization during meiotic sex-chromosome inactivation. We propose that male meiosis occurs amid global reprogramming of 3D chromatin organization and that strengthening of chromatin compartmentalization takes place in spermiogenesis to prepare the next generation of life.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All Hi-C sequencing data used in this study, including processed files for published datasets, have been deposited in the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) under the accession number GSE119805. The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Battulin, N. et al. Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach. Genome Biol. 16, 77 (2015).
Jung, Y. H. et al. Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep. 18, 1366–1382 (2017).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381.e20 (2017).
Hunter, N. Meiotic recombination: the essence of heredity. Cold Spring Harb. Perspect. Biol. 7, a016618 (2015).
Gray, S. & Cohen, P. E. Control of meiotic crossovers: from double-strand break formation to designation. Annu. Rev. Genet. 50, 175–210 (2016).
Kimmins, S. & Sassone-Corsi, P. Chromatin remodelling and epigenetic features of germ cells. Nature 434, 583–589 (2005).
Schultz, N., Hamra, F. K. & Garbers, D. L. A multitude of genes expressed solely in meiotic or postmeiotic spermatogenic cells offers a myriad of contraceptive targets. Proc. Natl Acad. Sci. USA 100, 12201–12206 (2003).
Namekawa, S. H. et al. Postmeiotic sex chromatin in the male germline of mice. Curr. Biol. 16, 660–667 (2006).
Hasegawa, K. et al. Scml2 establishes the male germline epigenome through regulation of histone h2a ubiquitination. Dev. Cell. 32, 574–588 (2015).
Maezawa, S., Yukawa, M., Alavattam, K. G., Barski, A. & Namekawa, S. H. Dynamic reorganization of open chromatin underlies diverse transcriptomes during spermatogenesis. Nucleic Acids Res. 46, 593–608 (2018).
Soumillon, M. et al. Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep. 3, 2179–2190 (2013).
Maezawa, S. et al. Polycomb protein SCML2 facilitates H3K27me3 to establish bivalent domains in the male germline. Proc. Natl Acad. Sci. USA 115, 4957–4962 (2018).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).ʼʼʼʼʼʼʼʼ
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fudenberg, G. & Mirny, L. A. Higher-order chromatin structure: bridging physics and biology. Curr. Opin. Genet. Dev. 22, 115–124 (2012).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Lajoie, B. R., Dekker, J. & Kaplan, N. The Hitchhiker’s guide to Hi-C analysis: practical guidelines. Methods 72, 65–75 (2015).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Zickler, D. & Kleckner, N. Meiotic chromosomes: integrating structure and function. Annu. Rev. Genet. 33, 603–754 (1999).
Scherthan, H. A bouquet makes ends meet. Nat. Rev. Mol. Cell Biol. 2, 621–627 (2001).
Scherthan, H., Schofisch, K., Dell, T. & Illner, D. Contrasting behavior of heterochromatic and euchromatic chromosome portions and pericentric genome separation in pre-bouquet spermatocytes of hybrid mice. Chromosoma. 123, 609–624 (2014).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384.e19 (2016).
Le Dily, F. et al. Distinct structural transitions of chromatin topological domains correlate with coordinated hormone-induced gene regulation. Genes Dev. 28, 2151–2162 (2014).
Wolff, J. et al. Galaxy HiCExplorer: a web server for reproducible Hi-C data analysis, quality control and visualization. Nucleic Acids Res. 46, W11–W16 (2018).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Cao, Y. et al. Accurate loop calling for 3D genomic data with cLoops. bioRxiv https://doi.org/10.1101/465849 (2018).
Lesch, B. J. & Page, D. C. Poised chromatin in the mammalian germ line. Development 141, 3619–3626 (2014).
Turner, J. M. Meiotic sex chromosome inactivation. Development 134, 1823–1831 (2007).
Ichijima, Y., Sin, H. S. & Namekawa, S. H. Sex chromosome inactivation in germ cells: emerging roles of DNA damage response pathways. Cell. Mol. Life Sci. 69, 2559–2572 (2012).
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome. Biol. 16, 152 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Minajigi, A. et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
Wang, C. Y., Jegu, T., Chu, H. P., Oh, H. J. & Lee, J. T. SMCHD1 merges chromosome compartments and assists formation of super-structures on the inactive X. Cell 174, 406–421.e25 (2018).
Namekawa, S. H., Payer, B., Huynh, K. D., Jaenisch, R. & Lee, J. T. Two-step imprinted X inactivation: repeat versus genic silencing in the mouse. Mol. Cell. Biol. 30, 3187–3205 (2010).
Sin, H. S., Ichijima, Y., Koh, E., Namiki, M. & Namekawa, S. H. Human postmeiotic sex chromatin and its impact on sex chromosome evolution. Genome Res. 22, 827–836 (2012).
Liang, Z. et al. Chromosomes progress to metaphase in multiple discrete steps via global compaction/expansion cycles. Cell 161, 1124–1137 (2015).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Schalbetter, S. A., Fudenberg, G., Baxter, J., Pollard, K. S. & Neale, M. J. Principles of meiotic chromosome assembly. bioRxiv https://doi.org/10.1101/442038 (2018).
Muller, H. et al. Characterizing meiotic chromosomes’ structure and pairing using a designer sequence optimized for Hi-C. Mol. Syst. Biol. 14, e8293 (2018).
Duan, Z. et al. A three-dimensional model of the yeast genome. Nature 465, 363–367 (2010).
Mizuguchi, T. et al. Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe. Nature 516, 432–435 (2014).
Tjong, H., Gong, K., Chen, L. & Alber, F. Physical tethering and volume exclusion determine higher-order genome organization in budding yeast. Genome Res. 22, 1295–1305 (2012).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Haarhuis, J. H. I. et al. The cohesin release factor wapl restricts chromatin loop extension. Cell 169, 693–707.e14 (2017).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
McNicoll, F., Stevense, M. & Jessberger, R. Cohesin in gametogenesis. Curr. Top. Dev. Biol. 102, 1–34 (2013).
Loukinov, D. I. et al. Boris, a novel male germ-line-specific protein associated with epigenetic reprogramming events, shares the same 11-zinc-finger domain with CTCF, the insulator protein involved in reading imprinting marks in the soma. Proc. Natl Acad. Sci. USA 99, 6806–6811 (2002).
Bergeron-Sandoval, L. P., Safaee, N. & Michnick, S. W. Mechanisms and consequences of macromolecular phase separation. Cell 165, 1067–1079 (2016).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Larson, A. G. et al. Liquid droplet formation by HP1alpha suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Holliday, R. The biological significance of meiosis. Symp. Soc. Exp. Biol. 38, 381–394 (1984).
Bellve, A. R. Purification, culture, and fractionation of spermatogenic cells. Methods Enzymol. 225, 84–113 (1993).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Belaghzal, H., Dekker, J. & Gibcus, J. H. Hi-C 2.0: An optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation. Methods 123, 56–65 (2017).
Selvaraj, S., J, R. D., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nat. Biotechnol. 31, 1111–1118 (2013).
Abdennur, N., Goloborodko, A., Imakaev, M., & Mirny, L. mirnylab/cooler: v0.7. 10. https://doi.org/10.5281/zenodo.1243296 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kaplan, N. & Dekker, J. High-throughput genome scaffolding from in vivo DNA interaction frequency. Nat. Biotechnol. 31, 1143–1147 (2013).
Virkar, Y. & Clauset, A. Power-law distributions in binned empirical data. Ann. Appl. Stat. 8, 89–119 (2014).
Oliphant, T. E. Python for Scientific Computing. Comput. Sci. Eng. 9, 10–20 (2007).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Zhang, Y. et al. Spatial organization of the mouse genome and its role in recurrent chromosomal translocations. Cell 148, 908–921 (2012).
Ramirez, F. et al. High-affinity sites form an interaction network to facilitate spreading of the MSL complex across the X chromosome in Drosophila. Mol. Cell 60, 146–162 (2015).
Khan, A. & Mathelier, A. Intervene: a tool for intersection and visualization of multiple gene or genomic region sets. BMC Bioinformatics 18, 287 (2017).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Kobayashi, H. et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet. 8, e1002440 (2012).
Adams, S. R. et al. RNF8 and SCML2 cooperate to regulate ubiquitination and H3K27 acetylation for escape gene activation on the sex chromosomes. PLoS Genet. 14, e1007233 (2018).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Dowen, J. M. et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 159, 374–387 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome. Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome. Biol. 9, R137 (2008).
Kartashov, A. V. & Barski, A. BioWardrobe: an integrated platform for analysis of epigenomics and transcriptomics data. Genome. Biol. 16, 158 (2015).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Quinlan, A. R. BEDTools: the Swiss-Army tool for genome feature analysis.Curr. Protoc. Bioinformatics 47, 11.12.1–11.12.34 (2014).
Hadley, W. Ggplot2 (Springer Science + Business Media, LLC, New York, NY, 2016).
Acknowledgements
We thank J. Dekker, J. Gibcus, J. Taylor, M. Sauria, C. Price, J. Wolff, M. Yukawa, W. Deng, and R. Perea for advice and/or support during various stages of this project. We thank members of the Namekawa and Kaplan laboratories for discussion and helpful comments regarding the manuscript. Funding sources: Albert J. Ryan Fellowship to K.G.A.; National Institute of Health (NIH) R01 GM098605, R01 GM122776, and R21 ES027117, Research Grant (FY13-510) from the March of Dimes Foundation to S.H.N.; Azrieli Faculty Fellowship and Taub Fellowship to N.K.; NIH DP2 GM119134 to A.B.
Author information
Authors and Affiliations
Contributions
The manuscript was written by K.G.A., N.K., and S.H.N., with critical feedback from all other authors, and K.G.A and S.H.N. designed the Hi-C experiments. K.G.A and S.M. performed the Hi-C experiments. K.G.A., A.S., H.K., A.B., N.K., and S.H.N. designed and interpreted the computational analyses. K.G.A., A.S., H.K., and N.K. performed the computational analyses. N.K and S.H.N jointly supervised this work.
Corresponding authors
Ethics declarations
Competing interests
A.B. is a cofounder of Datirium, LLC.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
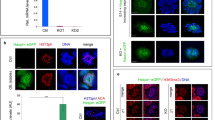
Supplementary Figure 1 Purity of germ cells isolated for Hi-C libraries.
a, Box-and-whisker plots in the style of Tukey showing the distributions of percent purity of cell fractions obtained from sedimentation velocity at unit gravity (Methods) for the following: pachytene spermatocyte (PS) Hi-C library replicates 1 and 2, and round spermatid (RS) Hi-C library replicates 1 and 2. Numbers (n) along the top indicate the numbers of fractions used to prepare the corresponding library replicates below. Means and standard deviations for the purities of each cell fraction comprising the pachytene spermatocyte libraries: 92% ± 5.6% (replicate 1) and 91% ± 4.7% (replicate 2); for the round spermatid libraries: 94% ± 1.5% (replicate 1) and 95% ± 2.1% (replicate 2). b, Fluorescence wide-field microscopy images of representative cell fractions for pachytene spermatocytes (top) and round spermatids (bottom). Scale bars: 20 µm. See Supplementary Dataset 1.
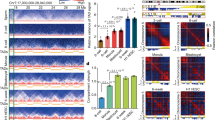
Supplementary Figure 2 Comparison of 3D chromatin organization in pachytene spermatocytes versus mitotic chromosomes.
a, Heat maps showing normalized Hi-C interaction frequencies (100-kb bins, chromosome 2) in pachytene spermatocytes (PS), metaphase meiosis II oocytes (MII oocyte), non-synchronized human foreskin fibroblasts (HFF1-non-synchronized), and synchronized prometaphase mitosis human foreskin fibroblasts (HFF1-mitosis; HFF1-non-synchronized is a control for HFF1-mitosis). b, Pearson’s correlation for Hi-C interaction frequencies (100-kb bins, chromosome 2) along with eigenvector 1 (EV1) from principle component analysis. See Supplementary Dataset 3.
Supplementary Figure 3 Genomic compartment strength calibration.
A random sample from the sperm Hi-C matrix, which has the highest genomic compartment strength, was mixed with a random sample from a Hi-C matrix which does not have genomic compartments (Methods). These were mixed at different ratios such that 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% of the total reads were sampled from the sperm matrix, and this was repeated 10 times. Then, average genomic compartment strength was calculated at each mixing ratio to construct calibration curves for intrachromosomal compartments (grey circles) and interchromosomal compartments (grey triangles). Standard deviations are not shown as they were all smaller than 0.005. Finally, pachytene spermatocyte (PS) and round spermatid (RS) genomic compartment strength values were plotted on the calibration curves in order to estimate whether it is plausible that the observed genomic compartment strength is due only to contamination by ~10% (dashed line) cells that have genomic compartments. See Supplementary Dataset 3.
Supplementary Figure 4 Association of histone post-translational modifications and genomic compartments in late spermatogenesis.
ChIP-seq data for H3K27ac, H3K4me3, and H3K27me3 to examine enrichment with respect to A and B compartment interiors and exteriors ± 1 kb (128-kb bins, all chromosomes) in pachytene spermatocytes (PS), round spermatids (RS), sperm, and embryonic stem cells (ESC). See Supplementary Dataset 3.
Supplementary Figure 5 Interchromosomal interactions in late spermatogenesis.
a-d, Heat maps showing genome-wide normalized Hi-C interaction frequencies (250-kb bins) in pachytene spermatocytes (PS; a), round spermatids (RS; b), sperm (c), and embryonic stem cells (ESC; d).
Supplementary Figure 6 TADs in late spermatogenesis.
a, Numbers of TADs called with the software package HiCExplorer (Methods) along with derived statistics for TAD size distributions for pachytene spermatocytes (PS), round spermatids (RS), sperm, and embryonic stem cells (ESC). SE: standard error. b, Size distribution histograms for TADs. See Supplementary Dataset 4.
Supplementary Figure 7 A subset of weak TAD boundaries apparent in pachytene spermatocytes is maintained in sperm.
a, Average observed/expected interaction frequencies at pachytene spermatocyte TAD boundaries ± 2 Mb (20-kb bins, chromosome 2) for pachytene spermatocytes (PS), round spermatids (RS), sperm, and embryonic stem cells (ESC). b, Schematic for interpretation of 2D matrix visualizations of observed/expected interaction frequencies at pachytene spermatocyte TAD start and stop boundaries. c, 2D matrix visualizations of log2 observed/expected interaction frequencies at pachytene spermatocyte TAD start and stop boundaries ± 0.5 Mb for all cell types (20-kb bins, all chromosomes). See Supplementary Dataset 4.
Supplementary Figure 8 Pairwise point interactions and active transcription during late spermatogenesis.
a, Average read enrichments in log2 counts per million (CPM) from ChIP-seq data for H3K27ac, H3K4me3, and H3K27me3, and RNA-seq data at sites of pachytene spermatocyte pairwise point interaction anchors (“anchors”) and all sequenced regions of the genome excluding anchor regions (“other”). b, Box-and-whisker plots in the style of Tukey showing the distributions of log2 counts per million for the datasets at “anchor” and “other” regions. Numbers along the top indicate the adjusted p values from Wilcoxon rank sum tests, with Bonferroni post corrections, between “anchor” and “other” regions. Statistics were derived from n = 1 sample pooled from 2 biologically independent samples. c, Hierarchical clusters of the “anchor,” ChIP-, and RNA-seq datasets after Pearson (top) and Spearman (bottom) correlation calculations. d, Hi-C interaction heat maps (20-kb bins, chromosome 2, 48–55 ± Mb) for pachytene spermatocytes (PS), round spermatids (RS), sperm, and embryonic stem cells (ESC) showing the dynamics of local interactions of active gene loci together with RNA- and ChIP-seq data. y axis: RPKM. Solid bars: TADs called with the software package HiCExplorer (Methods). Green and grey highlights, arrows, and dashed circles indicate localized pairwise point interactions and related features of interest. See Supplementary Dataset 5.
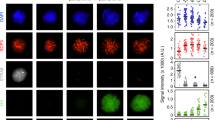
Supplementary Figure 9 Chromosome X lacks higher-order structure in late spermatogenesis.
a, Heat maps showing normalized Hi-C interaction frequencies (100-kb bins, chromosome X) in pachytene spermatocytes (PS), metaphase meiosis II oocytes (MII oocyte), non-synchronized human foreskin fibroblasts (HFF1-non-synch.), and synchronized prometaphase mitosis human foreskin fibroblasts (HFF1-mitosis; HFF1-non-synch. is a control for HFF1-mitosis). b, Pearson’s correlation for Hi-C interaction frequencies (100-kb bins, chromosome X) along with eigenvector 1 (EV1) from principle component analysis. c, Hi-C interaction heat maps (20-kb bins, chromosome X, 80–92 Mb) showing the dynamics of local interactions of gene loci together with RNA-seq data and ChIP-seq data for H3K27ac, H3K4me3, and H3K27me3. See Supplementary Dataset 3.
Supplementary information
Supplementary Information
Supplementary Figures 1–9
Supplementary Dataset 1
Purity of germ cells isolated for Hi-C libraries. Each of the four sheets represents data for Hi-C libraries. From left to right, those libraries are for pachytene spermatocyte (PS) replicate 1, pachytene spermatocyte replicate 2, round spermatid (RS) replicate 1, and round spermatid replicate 2. Within each sheet, the columns represent, from left to right, the name of the image of the isolated germ cell fraction from sedimentation velocity at unit gravity (Methods), the population cell type, the library (rep: replicate), and the mean purity (Methods).
Supplementary Dataset 2
Details and metrics for Hi-C datasets used in this study. The columns represent, from left to right, the dataset’s full name, its abbreviated name, replicate information (if applicable), PubMed reference number for initial study, database for initial study, accession number for initial study, URL for database query, database for this study, accession number for this study, restriction enzyme used when generating the dataset, paired-end read length for sequencing, and the genome to which the reads were aligned. The remaining 13 columns represent dataset sequencing results and mapping statistics; for detailed information on the metrics, please see Methods 72, 65–75 (2015).
Supplementary Dataset 3
Genomic compartment strength calibration and first eigenvectors from principle component analyses of pooled Hi-C datasets. The first sheet represents genomic compartment strength measurements for mixtures of the sperm Hi-C matrix and a Hi-C matrix that does not have genomic compartments (Methods); the genomic compartment strength measurements are for the means and standard deviations of intrachromosomal (cis) interactions, and the means and standard deviations of interchromosomal (trans) interactions. The next four sheets contain first eigenvectors from principle component analyses of the following Hi-C datasets binned in 128-kb windows (Methods): pachytene spermatocyte (PS), round spermatid (RS), sperm, and embryonic stem cell (ESC); EV1: first eigenvector. The final four sheets contain first eigenvectors from principle component analyses of the following Hi-C datasets binned in 100-kb windows (Methods): pachytene spermatocyte, metaphase meiosis II oocyte (MII oocyte), non-synchronized human foreskin fibroblast (HFF1-non-synchronized), and synchronized prometaphase mitosis human foreskin fibroblast (HFF1-mitosis; HFF1-non-synchronized is a control for HFF1-mitosis).
Supplementary Dataset 4
Information for topologically associating domains (TADs): boundaries, boundary intersections between datasets, sizes, and derived statistics. The first three sheets contain the genomic locations of TAD boundaries ± 30 kb (60 kb centered on each boundary; Methods) for the following datasets: pachytene spermatocyte (PS), round spermatid (RS), and sperm. The fourth sheet contains the results from the evaluation of TAD boundary intersections via the program UpSetR (Methods). The fifth through eighth sheets contain information for the genomic locations of TAD start and stop positions, the initial separation scores for TADs (Methods), and individual TAD sizes for the following datasets: pachytene spermatocyte, round spermatid, sperm, and embryonic stem cell (ESC). The final sheet contains derived statistics for TAD sizes for the datasets.
Supplementary Dataset 5
Pairwise point interactions in pachytene spermatocytes. The first sheet contains genomic location information for anchors of pairwise point interactions; each point interaction has two anchors, and each anchor has a start and stop position. The sixth column contains the binomial p value calculated by the point interaction calling program cLoops (Methods). The second sheet contains genomic location information for the center of each anchor of each point interaction (Methods).
Rights and permissions
About this article
Cite this article
Alavattam, K.G., Maezawa, S., Sakashita, A. et al. Attenuated chromatin compartmentalization in meiosis and its maturation in sperm development. Nat Struct Mol Biol 26, 175–184 (2019). https://doi.org/10.1038/s41594-019-0189-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0189-y
This article is cited by
-
Rewiring of the 3D genome during acquisition of carboplatin resistance in a triple-negative breast cancer patient-derived xenograft
Scientific Reports (2023)
-
Tcf1 preprograms the mobilization of glycolysis in central memory CD8+ T cells during recall responses
Nature Immunology (2022)
-
Alcohol induced increases in sperm Histone H3 lysine 4 trimethylation correlate with increased placental CTCF occupancy and altered developmental programming
Scientific Reports (2022)
-
3D chromatin remodelling in the germ line modulates genome evolutionary plasticity
Nature Communications (2022)
-
Meiotic sex chromosome inactivation and the XY body: a phase separation hypothesis
Cellular and Molecular Life Sciences (2022)