Abstract
Serotonin (5-hydroxytryptamine; 5-HT) receptors modulate a variety of physiological processes ranging from perception, cognition and emotion to vascular and smooth muscle contraction, platelet aggregation, gastrointestinal function and reproduction. Drugs that interact with 5-HT receptors effectively treat diseases as diverse as migraine headaches, depression and obesity. Here we present four structures of a prototypical serotonin receptor—the human 5-HT2B receptor—in complex with chemically and pharmacologically diverse drugs, including methysergide, methylergonovine, lisuride and LY266097. A detailed analysis of these structures complemented by comprehensive interrogation of signaling illuminated key structural determinants essential for activation. Additional structure-guided mutagenesis experiments revealed binding pocket residues that were essential for agonist-mediated biased signaling and β-arrestin2 translocation. Given the importance of 5-HT receptors for a large number of therapeutic indications, insights derived from these studies should accelerate the design of safer and more effective medications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Berger, M., Gray, J. A. & Roth, B. L. The expanded biology of serotonin. Annu. Rev. Med. 60, 355–366 (2009).
McCorvy, J. D. & Roth, B. L. Structure and function of serotonin G protein–coupled receptors. Pharmacol. Ther. 150, 129–142 (2015).
Allen, J. A. & Roth, B. L. Strategies to discover unexpected targets for drugs active at G protein–coupled receptors. Annu. Rev. Pharmacol. Toxicol. 51, 117–144 (2011).
Vickers, S. P., Clifton, P. G., Dourish, C. T. & Tecott, L. H. Reduced satiating effect of d-fenfluramine in serotonin 5-HT2C receptor–mutant mice. Psychopharmacology 143, 309–314 (1999).
Roth, B. L. Drugs and valvular heart disease. N. Engl. J. Med. 356, 6–9 (2007).
Rothman, R. B. et al. Evidence for possible involvement of 5-HT2B receptors in the cardiac valvulopathy associated with fenfluramine and other serotonergic medications. Circulation 102, 2836–2841 (2000).
Zanettini, R. et al. Valvular heart disease and the use of dopamine agonists for Parkinson’s disease. N. Engl. J. Med. 356, 39–46 (2007).
Gustafsson, B. I., Hauso, O., Drozdov, I., Kidd, M. & Modlin, I. M. Carcinoid heart disease. Int. J. Cardiol. 129, 318–324 (2008).
Huang, X. P. et al. Parallel functional activity profiling reveals valvulopathogens are potent 5-hydroxytryptamine2B receptor agonists: implications for drug safety assessment. Mol. Pharmacol. 76, 710–722 (2009).
Papoian, T. et al. Regulatory Forum Review*: utility of in vitro secondary pharmacology data to assess risk of drug-induced valvular heart disease in humans: regulatory considerations. Toxicol. Pathol. 45, 381–388 (2017).
Hauso, Ø. et al. Long-term serotonin effects in the rat are prevented by terguride. Regul. Pept. 143, 39–46 (2007).
Wacker, D. et al. Structural features for functional selectivity at serotonin receptors. Science 340, 615–619 (2013).
Wacker, D. et al. Crystal structure of an LSD-bound human serotonin receptor. Cell 168, 377–389 (2017).
Katritch, V., Cherezov, V. & Stevens, R. C. Diversity and modularity of G protein–coupled receptor structures. Trends Pharmacol. Sci. 33, 17–27 (2012).
Latorraca, N. R., Venkatakrishnan, A. J. & Dror, R. O. GPCR dynamics: structures in motion. Chem. Rev. 117, 139–155 (2017).
Wacker, D., Stevens, R. C. & Roth, B. L. How ligands illuminate GPCR molecular pharmacology. Cell 170, 414–427 (2017).
Wang, C. et al. Structural basis for molecular recognition at serotonin receptors. Science 340, 610–614 (2013).
Yin, W. et al. Crystal structure of the human 5-HT1B serotonin receptor bound to an inverse agonist. Cell Discov. 4, 12 (2018).
Peng, Y. et al. 5-HT2C receptor structures reveal the structural basis of GPCR polypharmacology. Cell 172, 719–730 (2018).
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein–coupled receptor. Science 318, 1258–1265 (2007).
Ring, A. M. et al. Adrenaline-activated structure of β2-adrenoceptor stabilized by an engineered nanobody. Nature 502, 575–579 (2013).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure–function relations in G protein–coupled receptors. in Methods in Neurosciences Vol. 25 (ed. Sealfon, S. C.) 366–428 (Academic Press, San Diego, CA, USA, 1995).
Rasmussen, S. G. et al. Structure of a nanobody-stabilized active state of the β2-adrenoceptor. Nature 469, 175–180 (2011).
Rasmussen, S. G. et al. Crystal structure of the β2-adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
McCorvy, J. D. et al. Structure-inspired design of β-arrestin-biased ligands for aminergic GPCRs. Nat. Chem. Biol. 14, 126–134 (2018).
Stevens, R. C. et al. The GPCR Network: a large-scale collaboration to determine human GPCR structure and function. Nat. Rev. Drug Discov. 12, 25–34 (2013).
Bredberg, U., Eyjolfsdottir, G. S., Paalzow, L., Tfelt-Hansen, P. & Tfelt-Hansen, V. Pharmacokinetics of methysergide and its metabolite methylergometrine in man. Eur. J. Clin. Pharmacol. 30, 75–77 (1986).
Wacker, D. et al. Conserved binding mode of human β2-adrenergic receptor inverse agonists and antagonist revealed by X-ray crystallography. J. Am. Chem. Soc. 132, 11443–11445 (2010).
Nichols, D. E., Monte, A., Huang, X. & Marona-Lewicka, D. Stereoselective pharmacological effects of lysergic acid amides possessing chirality in the amide substituent. Behav. Brain Res. 73, 117–119 (1996).
Hofmann, C. et al. Lisuride, a dopamine receptor agonist with 5-HT2B receptor antagonist properties: absence of cardiac valvulopathy adverse drug reaction reports supports the concept of a crucial role for 5-HT2B receptor agonism in cardiac valvular fibrosis. Clin. Neuropharmacol. 29, 80–86 (2006).
Audia, J. E. et al. Potent, selective tetrahydro-β-carboline antagonists of the serotonin 2B (5HT2B) contractile receptor in the rat stomach fundus. J. Med. Chem. 39, 2773–2780 (1996).
Sato, T. et al. Pharmacological analysis and structure determination of 7-methylcyanopindolol-bound β1-adrenergic receptor. Mol. Pharmacol. 88, 1024–1034 (2015).
Ahuja, S. & Smith, S. O. Multiple switches in G protein–coupled receptor activation. Trends Pharmacol. Sci. 30, 494–502 (2009).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013).
Staus, D. P. et al. Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation. Nature 535, 448–452 (2016).
Che, T. et al. Structure of the nanobody-stabilized active state of the κ opioid receptor. Cell 172, 55–67 (2018).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein–coupled receptors. Structure 20, 967–976 (2012).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–731 (2009).
Roth, C. B., Hanson, M. A. & Stevens, R. C. Stabilization of the human β2-adrenergic receptor TM4-TM3-TM5 helix interface by mutagenesis of Glu1223.41, a critical residue in GPCR structure. J. Mol. Biol. 376, 1305–1319 (2008).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution—from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Jordan, M., Schallhorn, A. & Wurm, F. M. Transfecting mammalian cells: optimization of critical parameters affecting calcium-phosphate precipitate formation. Nucleic Acids Res. 24, 596–601 (1996).
Bourdon, D. M., Wing, M. R., Edwards, E. B., Sondek, J. & Harden, T. K. Quantification of isozyme-specific activation of phospholipase C-β2 by Rac GTPases and phospholipase-C-ε by Rho GTPases in an intact cell assay system. Methods Enzymol. 406, 489–499 (2006).
Acknowledgements
We thank R. Axel (Columbia University) for the HTLA cells expressing TEV-fused β-arrestin2 and R. Fischetti and the staff of APS GM/CA for assistance in the development and use of the minibeam and beam time at GM/CA-CAT beamline 23-ID at the Advanced Photon Source, which is supported by National Cancer Institute grant Y1-CO-1020 and National Institute of General Medical Sciences grant Y1-GM-1104. Use of the Advanced Photon Source was supported by the Office of Science of the US Department of Energy. This work was supported by US National Institutes of Health (NIH) grants R01MH61887 (B.L.R.), R01NS100930 (J.J.), U19MH82441 (J.J. and B.L.R.) and F31-NS093917 (R.H.J.O.), the NIMH Psychoactive Drug Screening Program Contract (B.L.R.) and the Michael Hooker Distinguished Chair of Pharmacology (B.L.R.).
Author information
Authors and Affiliations
Contributions
J.D.M. conceived the project, designed the mutant constructs and experiments, performed pharmacological assays, analyzed the data and wrote the manuscript; D.W. expressed protein, purified the receptor, optimized crystallization conditions, grew crystals for data collection, collected and processed diffraction data, supervised structure determination and assisted with preparing the manuscript; S.W. expressed protein, purified the receptor, optimized crystallization conditions, grew crystals for data collection, collected and processed diffraction data and assisted with preparing the manuscript; B.A. expressed protein, purified receptor, optimized crystallization conditions and grew crystals for data collection; J.L. designed and synthesized LY266097 analogs and performed analytical chemical analysis; K.L. assisted with performing PI hydrolysis signaling studies and analyzed the data; A.R.T. assisted with performing β-arrestin recruitment experiments; R.H.J.O. assisted with performing BRET experiments; T.C. assisted with binding studies; J.J. supervised ligand synthesis and edited the manuscript; B.L.R. was responsible for the overall project strategy and management and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Activation mechanisms and 5-HT2BR–methylergonovine structure.
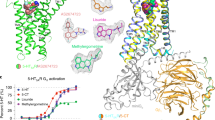
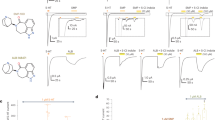
a, Additional 5-HT2BR ergoline SAR showing Gq-mediated calcium flux by N(1)-H-containing ergolines, LSD (red, EC50 = 26 nM, Emax = 82%) and ergotamine (ERG, black, EC50 = 334 nM, Emax = 85%) and antagonism by N(1)-alkyl ergolines, including LY215840, which contains N(1)-isopropyl (blue, IC50 = 2.0 nM). Data are expressed as percent 5-HT response and represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate. b, Alignment of the 5-HT2BR ERG (gray), LSD (purple) and methylergonovine (light blue) crystal structures reveals slight differences in positioning of the indole N(1)-H with respect to TM5. c, 5-HT2BR–methylergonovine structure Fo – Fc omit map of ligand (left), 2Fo – Fc regular map of ligand (middle) and binding pocket residues (right). d, Surface expression measured by ELISA reveals similar expression levels of the T1403.37 and A2255.46 mutants as for wild-type 5-HT2BR. Data represent means ± s.e.m. from quadruplicate replicates from two independent experiments (n = 2). e, Fo – Fc omit density map of residues T1403.37 and L3627.35 in the 5-HT2BR–methylergonovine structure. f, 5-HT Gq-mediated calcium flux agonist potency at T140A3.37 (red, EC50 = 73 nM) and T140V3.37 (blue, EC50 = 93 nM) compared to T140S3.37 (green, EC50 = 5.7 nM) and wild-type 5-HT2BR (black, EC50 = 5.2 nM). Data are expressed as fold-over-basal and represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate. g, 5-HT Gq-mediated calcium flux agonist potency at A225S5.46 (red, EC50 = 1.4 nM) and A225G5.46 (blue, EC50 = 1.5 nM) as compared to wild-type 5-HT2BR (black, EC50 = 3.2 nM). Data are expressed as fold-over-basal and represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate.
Supplementary Figure 2 Activation mechanism and 5-HT2BR(A225G5.46)–methysergide structure.
a, 5-HT2BR Gq-mediated PI hydrolysis showing similar basal levels of IP accumulation and Gq agonist activity by methysergide for A225G5.46 (green, EC50 = 0.7 nM) compared to no measured agonist activity for wild-type 5-HT2BR (black). Data are expressed as counts per minute (CPM) of [3H]IP and represent means ± s.e.m. from three independent experiments (n = 3) performed in duplicate. b, 5-HT2BR(A225G5.46)–methysergide structure Fo – Fc omit map of ligand (left), 2Fo – Fc regular map of ligand (middle) and binding pocket residues (right). c, Fo – Fc omit density map of residues T1403.37 and L3627.35 in the 5-HT2BR(A225G5.46)–methysergide structure. d, Alignment of the structure for the β2AR–ICI-118,551 complex (blue) with the nanobody-stabilized active state of β2AR in complex with epinephrine (green) indicates that the methyl of ICI-118,551 precludes inward TM5 movement.
Supplementary Figure 3 5-HT2BR–lisuride structure and OBP mutant L3627.35 surface expression.
a, 5-HT2BR–lisuride structure Fo – Fc omit map of ligand (left), 2Fo – Fc regular map of ligand (middle) and binding pocket residues (right). b, Fo – Fc omit density map of residues T1403.37 and L3627.35 in the structure of lisuride-bound 5-HT2BR. c, Alignment of the structures for the 5-HT2BR–LSD and 5-HT2BR–lisuride complexes reveals similar positioning of the indole N(1)-H with respect to TM5. d, Surface expression measured by ELISA reveals similar expression levels of L3627.35 mutants as for wild-type 5-HT2BR. Data represent means ± s.e.m. from quadruplicate replicates from two independent experiments (n = 2).
Supplementary Figure 4 Divergent function by OBP versus EBP mutations.
a, Left, 5-HT Gq-mediated calcium flux comparing T140A3.37 (red, EC50 = 130 nM) to wild-type 5-HT2BR (black, EC50 = 5.2 nM). Right, 5-HT β-arrestin2 recruitment comparing T140A3.37 (red, EC50 = 178 nM) to wild-type 5-HT2BR (black, EC50 = 3.6 nM). b, Left, 5-HT Gq-mediated calcium flux comparing A225G5.46 (green, EC50 = 1.5 nM) to wild-type 5-HT2BR (black, EC50 = 2.2 nM). Right, 5-HT β-arrestin2 recruitment comparing A225G5.46 (green, EC50 = 4.6 nM) to wild-type 5-HT2BR (black, EC50 = 3.6 nM). Data in a and b are expressed as fold-over-basal and represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate. c, LSD β-arrestin2 recruitment for the L362A7.35 (red), L362N7.35 (orange), L362Y7.35 (blue) and L362F7.35 (green) 5-HT2BR mutants. Data are expressed as luminescent counts per second (LCPS), indicating β-arrestin2 recruitment as measured by Tango, and represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate. d, Mutant 5-HT2BR L362F7.35 Gq-mediated PI hydrolysis by 5-HT (black, EC50 = 1.8 nM) and lisuride (blue, EC50 = 0.65 nM). Data are expressed as counts per minute (CPM) of [3H]IP and represent means ± s.e.m. from two independent experiments (n = 2) performed in duplicate. e, Mutant 5-HT2BR L362F7.35 Gq/ γ1 dissociation as measured by BRET2 showing 5-HT (black, EC50 = 1.3 nM) and lisuride (blue, EC50 = 1.2 nM) Gq agonist activity. Data are represented as means ± s.e.m. from two independent experiments (n = 2) and are expressed as NET BRET ratio (Methods). f, Mutant 5-HT2BR L362F7.35 LSD β-arrestin2 recruitment as measured by BRET1. Data are represented as means ± s.e.m. from two independent experiments (n = 2) and are expressed as NET BRET ratio (Methods).
Supplementary Figure 5 5-HT2BR structure and function by LY266097.
a,5-HT2BR–LY266097 structure Fo – Fc omit map of ligand (left), 2Fo – Fc regular map of ligand (middle) and binding pocket residues (right). b, Fo – Fc omit density map of residues T1403.37 and L3627.35 in the 5-HT2BR–LY266097 structure. c, LY266097 5-HT2BR Gq-mediated PI hydrolysis (EC50 = 3.6 nM, Emax = 52%). Data are expressed as percent 5-HT response and represent means ± s.e.m. from three independent experiments (n = 3) performed in duplicate. d, LY266097 β-arrestin2 recruitment as measured by BRET1 over 5, 30, 60 and 120 min. Data represent means ± s.e.m. from two independent experiments (n = 2) performed in duplicate and are expressed as percent 5-HT NET BRET ratio response. e, β-arrestin2 recruitment as measured by Tango comparing agonist activity of 5-HT (black, EC50 = 11 nM) to LY266097 (blue, closed circles), where LY266097 exhibits β-arrestin2 recruitment antagonism (blue, open circles, IC50 = 2.2 nM) of 5-HT. Data represent means ± s.e.m. from two independent experiments (n = 2) performed in triplicate and are expressed as percent 5-HT response.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Table 1 and Supplementary Note
Rights and permissions
About this article
Cite this article
McCorvy, J.D., Wacker, D., Wang, S. et al. Structural determinants of 5-HT2B receptor activation and biased agonism . Nat Struct Mol Biol 25, 787–796 (2018). https://doi.org/10.1038/s41594-018-0116-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0116-7
This article is cited by
-
Uncovering the unique characteristics of different groups of 5-HT5AR ligands with reference to their interaction with the target protein
Pharmacological Reports (2024)
-
AlphaFold2 versus experimental structures: evaluation on G protein-coupled receptors
Acta Pharmacologica Sinica (2023)
-
Molecular mechanism of biased signaling at the kappa opioid receptor
Nature Communications (2023)
-
Identification of 5-HT2A receptor signaling pathways associated with psychedelic potential
Nature Communications (2023)
-
The serotonin transporter sustains human brown adipose tissue thermogenesis
Nature Metabolism (2023)