Abstract
Thousands of long noncoding RNAs (lncRNAs) have been identified in eukaryotic genomes, many of which are expressed in spatially and temporally restricted patterns. Nonetheless, the roles of the majority of these transcripts are still unknown. One of the mechanisms by which lncRNAs function is through the modulation of chromatin states. To assess the functions of lncRNAs, we developed RNA-DamID, a novel approach that detects lncRNA–genome interactions in a cell-type-specific manner in vivo with high sensitivity and accuracy. Identifying the cell-type-specific genome occupancy of lncRNAs is vital to understanding their mechanisms of action in development and disease. We used RNA-DamID to investigate targeting of the lncRNAs in the Drosophila dosage-compensation complex (DCC) and show that initial targeting is cell-type specific.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Franke, A. & Baker, B. S. The rox1 and rox2 RNAs are essential components of the compensasome, which mediates dosage compensation in D rosophila. Mol. Cell 4, 117–122 (1999).
Southall, T. D. et al. Cell-type-specific profiling of gene expression and chromatin binding without cell isolation: assaying RNA Pol II occupancy in neural stem cells. Dev. Cell 26, 101–112 (2013).
van Steensel, B. & Henikoff, S. Identification of in vivo DNA targets of chromatin proteins using tethered dam methyltransferase. Nat. Biotechnol. 18, 424–428 (2000).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Peabody, D. S. The RNA binding site of bacteriophage MS2 coat protein. EMBO J. 12, 595–600 (1993).
Meller, V. H., Wu, K. H., Roman, G., Kuroda, M. I. & Davis, R. L. roX1 RNA paints the X chromosome of male Drosophila and is regulated by the dosage compensation system. Cell 88, 445–457 (1997).
Quinn, J. J. et al. Rapid evolutionary turnover underlies conserved lncRNA-genome interactions. Genes Dev. 30, 191–207 (2016).
Ho, J. W. K. et al. Comparative analysis of metazoan chromatin organization. Nature 512, 449–452 (2014).
Park, Y., Kelley, R. L., Oh, H., Kuroda, M. I. & Meller, V. H. Extent of chromatin spreading determined by roX RNA recruitment of MSL proteins. Science 298, 1620–1623 (2002).
Straub, T., Grimaud, C., Gilfillan, G. D., Mitterweger, A. & Becker, P. B. The chromosomal high-affinity binding sites for the Drosophila dosage compensation complex. PLoS Genet. 4, e1000302 (2008).
Alekseyenko, A. A. et al. A sequence motif within chromatin entry sites directs MSL establishment on the Drosophila X chromosome. Cell 134, 599–609 (2008).
Keller, C. I. & Akhtar, A. The MSL complex: juggling RNA-protein interactions for dosage compensation and beyond. Curr. Opin. Genet. Dev 31, 1–11 (2015).
Apte, M. S. et al. Generation of a useful roX1 allele by targeted gene conversion. G3 (Bethesda) 4, 155–162 (2014).
Ilik, I. & Akhtar, A. roX RNAs: non-coding regulators of the male X chromosome in flies. RNA Biol. 6, 113–121 (2009).
Ilik, I. A. et al. Tandem stem-loops in roX RNAs act together to mediate X chromosome dosage compensation in D rosophila. Mol. Cell 51, 156–173 (2013).
Villa, R., Schauer, T., Smialowski, P., Straub, T. & Becker, P. B. PionX sites mark the X chromosome for dosage compensation. Nature 537, 244–248 (2016).
Bashaw, G. J. & Baker, B. S. The msl-2 dosage compensation gene of Drosophila encodes a putative DNA-binding protein whose expression is sex specifically regulated by Sex-lethal. Development 121, 3245–3258 (1995).
Zhou, S. et al. Male-specific lethal 2, a dosage compensation gene of Drosophila, undergoes sex-specific regulation and encodes a protein with a RING finger and a metallothionein-like cysteine cluster. EMBO J. 14, 2884–2895 (1995).
Ramírez, F. et al. High-affinity sites form an interaction network to facilitate spreading of the MSL complex across the X Chromosome in D rosophila. Mol. Cell 60, 146–162 (2015).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Luo, L., Liao, Y. J., Jan, L. Y. & Jan, Y. N. Distinct morphogenetic functions of similar small GTPases: Drosophila Drac1 is involved in axonal outgrowth and myoblast fusion. Genes Dev. 8, 1787–1802 (1994).
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003).
Markstein, M., Pitsouli, C., Villalta, C., Celniker, S. E. & Perrimon, N. Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nat. Genet. 40, 476–483 (2008).
Marshall, O. J., Southall, T. D., Cheetham, S. W. & Brand, A. H. Cell-type-specific profiling of protein-DNA interactions without cell isolation using targeted DamID with next-generation sequencing. Nat. Protoc. 11, 1586–1598 (2016).
Marshall, O. J. & Brand, A. H. damidseq_pipeline: an automated pipeline for processing DamID sequencing datasets. Bioinformatics 31, 3371–3373 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Stempor, P. & Ahringer, J. SeqPlots - interactive software for exploratory data analyses, pattern discovery and visualization in genomics. Wellcome Open Res. 1, 14 (2016).
Acknowledgements
We thank M. Kuroda (Harvard University), V. Meller (Wayne State University), T. Megraw (Florida State University) and P. Amaral (Gurdon Institute) for reagents, T. Leonardi for advice on data visualization and P. Amaral, T. Southall, O. Marshall and members of the Brand Lab for advice and discussion. This work was funded by the Royal Society Darwin Trust Research Professorship, a Wellcome Trust Senior Investigator Award 103792 and Wellcome Trust Programme Grant 092545 to A.H.B. S.W.C. was funded by a Herchel Smith Research Studentship. A.H.B acknowledges core funding to the Gurdon Institute from the Wellcome Trust (092096) and CRUK (C6946/A14492).
Author information
Authors and Affiliations
Contributions
S.W.C. and A.H.B. designed the experiments. S.W.C. performed the experiments. S.W.C. and A.H.B. analyzed the data. S.W.C. and A.H.B. wrote the manuscript. Both authors reviewed the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Constructs used for RNA-DamID.
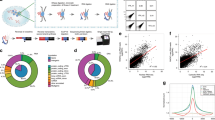
(a) An MCP tandem dimer fused to two nuclear localisation signals and Dam is expressed under the control of UAS. Expression is reduced by an upstream open reading frame (uORF, also called LT3) followed by two stop codons upstream of the Dam-fusion gene. Mini-white is used a selectable marker to identify transgenic flies. An attB site allows site-specific integration in attP containing fly lines using the ϕC31 integrase. (b) A lncRNA fused to the 3 tandem 5’ MS2 tags is expressed under the control of UAS. Mini-white is used a selectable marker to identify transgenic flies. An attB site allows site-specific integration in attP containing fly lines using the ϕC31 integrase. (c) 3 tandem 5’ MS2 tags are expressed under the control of UAS. This construct is used as a negative control to eliminate any non-specific signal caused by the association of the MS2 tags with specific regions of chromatin. Mini-white is used a selectable marker to identify transgenic flies. An attB site allows site-specific integration in attP containing fly lines using the ϕC31 integrase.
Supplementary Figure 2 Normalization of RNA-DamID data.
(a) RNA-DamID data is normalised by comparing enrichment of an MS2-tagged lncRNA, co-expressed with Dam-MCP over an 3xMS2 transcript co-expressed with Dam-MCP. (b) Genome browser tracks of negative control (3xMS2, Dam-MCP) and RNA-DamID (3xMS2-roX2, Dam-MCP) displayed in reads per million mapped. The negative control predominantly methylates open chromatin regions. RNA-DamID preferentially methylates roX2 targets on the X chromosome. Generating a ratio of the test over control reduces autosomal peaks. (c) roX2 RNA-DamID signal normalised to the negative control is highly correlated to roX2 RNA-DamID normalised to Dam-alone. Data is plotted as log2 normalised RNA-DamID signal enrichment over negative control.
Supplementary Figure 3 RNA-DamID is highly reproducible.
roX2 RNA-DamID biological replicates are highly correlated on the X chromosome but not on the autosomes where little signal is detected. Data is plotted as log2 normalized RNA-DamID signal enrichment over negative control.
Supplementary Figure 4 RNA-DamID identifies 563 novel roX2-binding sites.
RNA-DamID enables the identification of novel roX2 binding sites. These novel sites roX2 RNA-DamID co-localise with roX2 ChIRP, Msl3 TaDa and H4K16ac ChIP. RNA-DamID scale represents log2 fold change of 3xMS2-roX2, Dam-MCP compared to 3xMS2, Dam-MCP (average of two biological replicates). Msl3 TaDa scale is log2 fold change of Msl3-Dam fusion compared to Dam-alone. ChIRP signal is log2 transformed. H4K16ac represents log2 fold change of H4K16ac ChIP over input.
Supplementary Figure 5 roX2 does not spread in cis.
(a) roX2 does not spread from the integration site into flanking chromatin on chromosome 3L. Data is shown as reads mapped per million for roX2 and the negative control mapped separately from replicate 1. (b) Regions upstream and downstream of the site of transcription are methylated in both the negative control and 3xMS2-roX2. The roX2 transgene is highly methylated when co-expressed with Dam-MCP (this sequence is absent from the negative control).
Supplementary Figure 6 roX1 and roX2 occupancy are highly correlated.
roX1 and roX2 binding is whole larvae is highly correlated across the X chromosome. Data is plotted as log2 normalised RNA-DamID signal enrichment over negative control.
Supplementary information
Rights and permissions
About this article
Cite this article
Cheetham, S.W., Brand, A.H. RNA-DamID reveals cell-type-specific binding of roX RNAs at chromatin-entry sites. Nat Struct Mol Biol 25, 109–114 (2018). https://doi.org/10.1038/s41594-017-0006-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-017-0006-4
This article is cited by
-
A micropeptide JunBP regulated by TGF-β promotes hepatocellular carcinoma metastasis
Oncogene (2023)
-
Non-coding RNAs in chromatin folding and nuclear organization
Cellular and Molecular Life Sciences (2021)
-
Overcoming challenges and dogmas to understand the functions of pseudogenes
Nature Reviews Genetics (2020)
-
Enhanced nucleotide chemistry and toehold nanotechnology reveals lncRNA spreading on chromatin
Nature Structural & Molecular Biology (2020)
-
Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells
Nature Communications (2018)