Abstract
Rapid eye movement (REM) sleep is a sleep state characterized by skeletal muscle paralysis and cerebral cortical activation. Yet, global cortical dynamics and their role in regulating REM sleep remain unclear. Here we show that in mice, REM sleep is accompanied by highly patterned cortical activity waves, with the retrosplenial cortex (RSC) as a major initiation site. Two-photon imaging of layer 2/3 pyramidal neurons of the RSC revealed two distinct patterns of population activities during REM sleep. These activities encoded two sequential REM sleep substages, characterized by contrasting facial movement and autonomic activity and by distinguishable electroencephalogram theta oscillations. Closed-loop optogenetic inactivation of RSC during REM sleep altered cortical activity dynamics and shortened REM sleep duration via inhibition of the REM substage transition. These results highlight an important role for the RSC in dictating cortical dynamics and regulating REM sleep progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Large raw image data from wide-field imaging and two-photon imaging are available upon request from the authors. Source data are provided with this paper.
Code availability
Links for open-source toolboxes used in this study are provided. Custom-made MATLAB codes for wide-field imaging processing, sleep scoring and mouse facial expression analysis are deposited on GitHub (https://github.com/Yoyo666222/Liulab2022_NN/). Code used for additional analyses is available from the corresponding author upon reasonable request.
References
Scammell, T. E., Arrigoni, E. & Lipton, J. O. Neural circuitry of wakefulness and sleep. Neuron 93, 747–765 (2017).
Liu, D. & Dan, Y. A motor theory of sleep–wake control: arousal–action circuit. Annu. Rev. Neurosci. 42, 27–46 (2019).
Lu, J., Sherman, D., Devor, M. & Saper, C. B. A putative flip-flop switch for control of REM sleep. Nature 441, 589–594 (2006).
Weber, F. et al. Control of REM sleep by ventral medulla GABAergic neurons. Nature 526, 435–438 (2015).
Sanchez-Vives, M. V. & McCormick, D. A. Cellular and network mechanisms of rhythmic recurrent activity in neocortex. Nat. Neurosci. 3, 1027–1034 (2000).
Funk, C. M., Honjoh, S., Rodriguez, A. V., Cirelli, C. & Tononi, G. Local slow waves in superficial layers of primary cortical areas during REM sleep. Curr. Biol. 26, 396–403 (2016).
Valero, M. et al. Sleep down state-active ID2/Nkx2.1 interneurons in the neocortex. Nat. Neurosci. 24, 401–411 (2021).
Vyazovskiy, V. V. & Harris, K. D. Sleep and the single neuron: the role of global slow oscillations in individual cell rest. Nat. Rev. Neurosci. 14, 443–451 (2013).
Krone, L. B. et al. A role for the cortex in sleep–wake regulation. Nat. Neurosci. 24, 1210–1215 (2021).
Morairty, S. R. et al. A role for cortical nNOS/NK1 neurons in coupling homeostatic sleep drive to EEG slow wave activity. Proc. Natl Acad. Sci. USA 110, 20272–20277 (2013).
Aserinsky, E. & Kleitman, N. Regularly occurring periods of eye motility, and concomitant phenomena, during sleep. Science 118, 273–274 (1953).
Jouvet, M. Paradoxical sleep—a study of its nature and mechanisms. Prog. Brain Res. 18, 20–62 (1965).
Siclari, F. et al. The neural correlates of dreaming. Nat. Neurosci. 20, 872–878 (2017).
Cardin, J. A., Crair, M. C. & Higley, M. J. Mesoscopic imaging: shining a wide light on large-scale neural dynamics. Neuron 108, 33–43 (2020).
Makino, H. et al. Transformation of cortex-wide emergent properties during motor learning. Neuron 94, 880–890 (2017).
Kim, T. H. et al. Long-term optical access to an estimated one million neurons in the live mouse cortex. Cell Rep. 17, 3385–3394 (2016).
Aime, M. et al. Paradoxical somatodendritic decoupling supports cortical plasticity during REM sleep. Science 376, 724–730 (2022).
Yuzgec, O., Prsa, M., Zimmermann, R. & Huber, D. Pupil size coupling to cortical states protects the stability of deep sleep via parasympathetic modulation. Curr. Biol. 28, 392–400 (2018).
Reidl, J., Starke, J., Omer, D. B., Grinvald, A. & Spors, H. Independent component analysis of high-resolution imaging data identifies distinct functional domains. Neuroimage 34, 94–108 (2007).
Barnett, L. & Seth, A. K. The MVGC multivariate Granger causality toolbox: a new approach to Granger-causal inference. J. Neurosci. Methods 223, 50–68 (2014).
MacDowell, C. J. & Buschman, T. J. Low-dimensional spatiotemporal dynamics underlie cortex-wide neural activity. Curr. Biol. 30, 2665–2680 (2020).
Mackevicius, E. L. et al. Unsupervised discovery of temporal sequences in high-dimensional datasets, with applications to neuroscience. Elife https://doi.org/10.7554/eLife.38471 (2019).
Maciel, R. et al. Is REM sleep a paradoxical state?: Different neurons are activated in the cingulate cortices and the claustrum during wakefulness and paradoxical sleep hypersomnia. Biochem. Pharmacol. 191, 114514 (2021).
Dolensek, N., Gehrlach, D. A., Klein, A. S. & Gogolla, N. Facial expressions of emotion states and their neuronal correlates in mice. Science 368, 89–94 (2020).
Simor, P., van der Wijk, G., Nobili, L. & Peigneux, P. The microstructure of REM sleep: why phasic and tonic? Sleep. Med. Rev. 52, 101305 (2020).
Koroma, M. et al. Sleepers selectively suppress informative inputs during rapid eye movements. Curr. Biol. 30, 2411–2417 (2020).
Patel, A. K., Reddy, V. & Araujo, J. F. Physiology, Sleep Stages (StatPearls, 2022).
Hong, C. C. et al. fMRI evidence for multisensory recruitment associated with rapid eye movements during sleep. Hum. Brain Mapp. 30, 1705–1722 (2009).
Mahn, M. et al. High-efficiency optogenetic silencing with soma-targeted anion-conducting channelrhodopsins. Nat. Commun. 9, 4125 (2018).
de Almeida-Filho, D. G. et al. Hippocampus–retrosplenial cortex interaction is increased during phasic REM and contributes to memory consolidation. Sci. Rep. 11, 13078 (2021).
Renouard, L. et al. The supramammillary nucleus and the claustrum activate the cortex during REM sleep. Sci. Adv. 1, e1400177 (2015).
Koike, B. D. V. et al. Electrophysiological evidence that the retrosplenial cortex displays a strong and specific activation phased with hippocampal theta during paradoxical (REM) sleep. J. Neurosci. 37, 8003–8013 (2017).
Braun, A. R. et al. Regional cerebral blood flow throughout the sleep–wake cycle. An H215O PET study. Brain 120, 1173–1197 (1997).
Kitanishi, T., Umaba, R. & Mizuseki, K. Robust information routing by dorsal subiculum neurons. Sci. Adv. https://doi.org/10.1126/sciadv.abf1913 (2021).
Rasch, B. & Born, J. About sleep’s role in memory. Physiol. Rev. 93, 681–766 (2013).
Walker, M. P. The role of sleep in cognition and emotion. Ann. N. Y. Acad. Sci. 1156, 168–197 (2009).
Rivera-Garcia, A. P., Ramirez-Salado, I., Corsi-Cabrera, M. & Calvo, J. M. Facial muscle activation during sleep and its relation to the rapid eye movements of REM sleep. J. Sleep. Res. 20, 82–91 (2011).
Mikulovic, S. et al. Ventral hippocampal OLM cells control type 2 theta oscillations and response to predator odor. Nat. Commun. 9, 3638 (2018).
Goyal, A. et al. Functionally distinct high and low theta oscillations in the human hippocampus. Nat. Commun. 11, 2469 (2020).
Han, W. et al. Integrated control of predatory hunting by the central nucleus of the amygdala. Cell 168, 311–324 (2017).
Barger, Z., Frye, C. G., Liu, D., Dan, Y. & Bouchard, K. E. Robust, automated sleep scoring by a compact neural network with distributional shift correction. PLoS ONE 14, e0224642 (2019).
Meng, Q. et al. Tracking eye movements during sleep in mice. Front. Neurosci. 15, 616760 (2021).
Vollmer, M. A robust, simple and reliable measure of heart rate variability using relative RR intervals. in Computing in Cardiology 42, 609–612 (2015).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. Elife https://doi.org/10.7554/eLife.12727 (2016).
Liu, D. et al. A common hub for sleep and motor control in the substantia nigra. Science 367, 440–445 (2020).
Song, J. H. et al. Precise mapping of single neurons by calibrated 3D reconstruction of brain slices reveals topographic projection in mouse visual cortex. Cell Rep. 31, 107682 (2020).
Vesuna, S. et al. Deep posteromedial cortical rhythm in dissociation. Nature 586, 87–94 (2020).
Cardoso, J. F. High-order contrasts for independent component analysis. Neural Comput. 11, 157–192 (1999).
Franklin, K. B. J. & Paxinos, G. Paxinos and Franklin’s the Mouse Brain in Stereotaxic Coordinates, Compact: The Coronal Plates and Diagrams (Academic Press, 2008).
Pnevmatikakis, E. A. & Giovannucci, A. NoRMCorre: an online algorithm for piecewise rigid motion correction of calcium imaging data. J. Neurosci. Methods 291, 83–94 (2017).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Zhou, P. et al. Efficient and accurate extraction of in vivo calcium signals from microendoscopic video data. Elife https://doi.org/10.7554/eLife.28728 (2018).
Acknowledgements
We thank M. M. Poo, Y. Yang and S. Xu for helpful comments; J. Tang and Y. Pan for tracing and histology; L. Kang for obtaining and breeding transgenic mice; S. Gong for mouse drawings; and X. Ge for useful information on the wide-field imaging system and discussion on the wide-field imaging data. This work was supported by the National Science and Technology Innovation 2030 grant (2022ZD0206100), the Shanghai Municipal Science and Technology Major Project (2018SHZDZX05 and 20JC1419500), a Lingang Laboratory grant (LG-QS-202203-01) and a Lingang Laboratory & National Key Laboratory of Human Factors Engineering joint grant.
Author information
Authors and Affiliations
Contributions
Y.D. and D.L. designed the experiments and wrote the manuscript. Y.D. performed most of the experiments and data analysis. Y.D. developed the deep learning algorithm for online sleep scoring and established methods for facial expression analysis. J.L. performed part of the optogenetic experiments and histology. M.Z. performed part of two-photon imaging experiments, retrograde tracing and whole-brain mapping. D.L. supervised all aspects of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Pierre-Hervé Luppi, Yueqing Peng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Activity of cortical areas across brain states.
a, Raw fluorescence signals of RSC and S1BF excited by sequential 410 nm or 470 nm light. Shown are separated ∆F/F Ca2+ signals for F410 and F470, together with EEG spectrogram, EMG trace and brain states. b, Example F410-corrected Ca2+ traces (∆F/F) for 6 cortical areas together with EEG spectrogram, EMG trace, EEG theta/delta ratio and color-coded brain states, during a same recording period as data shown in Fig. 1d. c, Total facial movements during active wake (aWake), quiet wake (qWake), NREM sleep or REM sleep (n = 6 sessions from 5 mice). The red box denotes facial area used for calculating facial movements. For each mouse the amplitude of facial movements was normalized by that of NREM state before averaged across mice. P(aWake vs NREM) = 0.0015; P(qWake vs NREM) = 0.36; P(REM vs NREM) = 0.0063. *P < 0.05, two-sided paired t test. d, Average Ca2+ activity during aWake and qWake state (n = 6 sessions from 5 mice). e, Activity difference between REM sleep and aWake state, with statistical significance (two-sided paired t test) plotted against difference in average Ca2+ ∆F/F. Error bars, SEM.
Extended Data Fig. 2 Cortex-wide neuronal activity dynamics across brain states.
a-d, Matrix of Granger causality between each pair of 11 cortical areas (left) and summary of causality between RSC and other areas (right), during late REM sleep (a), aWake state (b), early NREM sleep (c) and late NREM sleep (d). Data from 6 sessions from 5 mice were pooled for these analysis. Note that the uppermost row is the causality from RSC to other 10 areas. Causality with P < 0.01 (Granger’s F-test) was shown after correction for multiple comparisons by false discovery rate. Red arrows, from RSC to other modules; Blue arrows, from other modules to RSC. e, Common spatiotemporal motifs of cortex-wide Ca2+ activity. Time courses of common motifs 5 to 7 as shown in (Fig. 2c–e) showing spatiotemporal patterns of neuronal Ca2+ activity across dorsal cortex. The intensity scale is normalized separately for each motif.
Extended Data Fig. 3 Two-photon imaging of cortical neuronal activities across brain states.
a, Top: fluorescence image of RSC in a RBP4Cre mouse injected with AAV-DIO-GCaMP6s. Scale bar: 200 μm. Bottom: example field of view for two-photon imaging. Scale bar: 50 μm. b, Example Ca2+ traces of RSC L5 neurons together with color-coded brain states. c, Average Ca2+ activity of RSC L5 neurons across sleep states. Each line indicates averaged Ca2+ activity for all identified neurons from one recording session (n = 9 sessions from 4 mice). P(Wake vs NREM) = 0.72; P(Wake vs REM) = 0.0023; P(NREM vs REM) = 0.0012; **P < 0.005, two-sided paired t test. d-f, Similar to a-c, but for L2/3 neurons in the M2 of Thy1-GCaMP6s mice. Data were from n = 6 sessions in 4 mice. P > 0.1 for all comparisons. P(Wake vs NREM) = 0.22; P(Wake vs REM) = 0.85; P(NREM vs REM) = 0.10. Scale bar in d: 50 μm. g-h, Scatter (g) or line (h) plots showing the distribution of the activity difference between REM sleep and other states, for RSC L2/3 neurons (n = 1238 neurons from 6 mice), RSC L5 neurons (n = 289 from 4 mice) and M2 L2/3 neurons (n = 193 from 4 mice). i, Percentages of REM-active, REM-inactive and non-modulated neurons in RSC L2/3 (n = 12 sessions from 6 mice), RSC L5 (n = 9 sessions from 4 mice) or M2 L2/3 (n = 6 sessions from 4 mice). * P < 0.05, ** P < 0.01, *** P < 0.001, two-sided unpaired t test. j, Averaged Ca2+ activities for REM-active (n = 706 neurons), REM-inactive (n = 188) and non-modulated (n = 344) neurons in the L2/3 of RSC, at NREM → REM (top) or REM → wake (bottom) transitions (from 6 mice). The same dataset were used as the top panel in g. Vertical lines, transition time points. Dotted lines above the averaged traces indicate time points that are significantly different from baseline (−30s to −6s before NREM → REM, or 6 s to 30 s after REM → wake). Closed dots: P < 0.05 for REM-active neurons; open dots: P < 0.05 for REM-inactive neurons. P > 0.1 for non-modulated neurons. Kruskal-Wallis tests with Dunn’s correction for multiple comparison. Note that the latency for significant activity increase of REM-active neurons was 2.3 s. Error bars or shadings, SEM.
Extended Data Fig. 4 RSC population neuronal activity encodes REM substages.
a, Example EEG spectrogram, EMG trace, EEG theta/delta ratio, low theta/high theta ratio and color-coded brain states. Blue shadings show three REM episodes in a same recording session. b, Visualization of the two CC clusters of RSC neuronal activity in the same three REM episodes as shown in a. REM-active neurons (n = 63 in a recording session with 115 identified neurons) were used in this analysis. c, The ratio of cell numbers for type I and type II neuron (n = 27 episode from 6 mice). * P = 0.026, two-sided paired t test. Error bars, SEM. d, Averaged neuronal activities of type I and type II neurons in the three REM episodes, together with color-coded two REM substages. Shadings, SEM. e, Maps of recorded RSC cells in the three REM episodes as shown in a,b,d. Cells defined as type I were color-coded in cyan, and cells defined as type II were in purple. Note that neurons with stable identity across REM episodes were color-filled. Bottom right, comparison of the percentage of neurons with stable identity in observed data to that estimated from random sampling, for each pair of REM episodes (n = 28 pairs from 6 mice). ***P = 0.0004, two-sided paired t test. Error bars, SEM. f, Averaged Ca2+ activity of RSC L2/3 (top, n = 6 mice), RSC L5 neuron (middle, n = 4 mice) from two-photon imaging and total RSC activity from wide-field imaging (bottom, n = 5 mice) at brain state transitions. Each qREM substage was temporally compressed to one unit length, and aREM substage compressed to three unit length, before averaged over multiple episodes. Shadings, SEM. g, Distribution of correlation coefficient (CC: Ca2+ activity vs. binary labelling of eye movements) in type I&II RSC populations. The distribution was calculated for each episode before averaging across 27 REM episodes. Each line represents CC distribution for one episode. P > 0.7 for comparison of each CC distribution with zero (PTypeI = 0.88, PTypeII = 0.77), two-tail z test. Error bars, SEM.
Extended Data Fig. 5 Comparison of EEG power spectra and autonomic activity between the two REM substages.
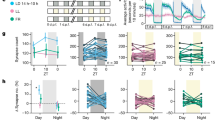
a, Normalized EEG power spectra in qREM and aREM, averaged across 10 sessions from 7 mice. Red dots indicate frequency bins that are significantly different between the two substages (at P < 0.05, two-sided paired t test with Bonferroni multiple comparison correction). Note that qREM has a significant higher power from 6.5 to 7.5 Hz, while aREM has significant higher power from 8.2 to 11 Hz. b, Averaged EEG spectrogram, EEG delta, theta power, low theta (6–8 Hz) to high theta (8–10 Hz) ratio, eye movements, facial movements and neck EMG power at brain state transitions. Each qREM substage was temporally compressed to one unit length. For each trace, data were normalized by that of NREM sleep before averaging across mice. Vertical lines represent transition points. Shadings, SEM. c, EEG theta/delta ratio (left) and low theta/high theta ratio (right) in qREM and aREM substages (n = 10 sessions from 7 mice). P(theta/delta)=0.13; P(Low/high theta)=0.04. d, Normalized eye movements (measured by pupil movements, left), and facial movements (right) in qREM and aREM substages (n = 10 sessions from 7 mice). The red box denotes facial area used for calculating facial movements. P(Eye movement)=0.0033; P(Facial movement)= 0.000024. e, Normalized neck EMG power in qREM and aREM substages (n = 10 sessions from 7 mice). P = 0.42. f, Heart rate variability (HRV), measured by relative RR intervals, in qREM and aREM substages (n = 7 session from 5 mice). Note that lower HRV indicates higher sympathetic activation. P = 0.0083. g, Respiration rate in qREM and aREM substages (n = 6 sessions from 3 mice). P = 0.00025. h, Cortical hemodynamics in qREM and aREM substages, reflected by decrease of fluorescence signal excited by 410 nm light (n = 6 sessions from 5 mice). For each session, the decrease in F410 was normalized by that of NREM sleep before averaged across mice (P = 0.0044). *P < 0.05, **P < 0.01, ***P < 0.001, two-sided paired t test. Error bars and shadings, SEM.
Extended Data Fig. 6 Facial EMG recording revealed two REM substages in freely-moving mice.
a, Schematics for EEG, neck EMG and facial EMG recording in a freely-moving mouse. MS, masseter muscle. b, Example EEG spectrogram, EEG theta (6–10 Hz) to delta (1–4.5 Hz) ratio, low theta (6–8 Hz) to high theta (8–10 Hz) ratio, neck EMG, facial EMG trace and color-coded four brain states. c, Averaged EEG spectrogram, EEG theta/delta ratio, low theta/high theta ratio, neck EMG, and facial EMG power at NREM → REM sleep, and REM sleep→Wake transitions, averaged across 6 recording sessions from 4 mice. For each trace, data were normalized by that of NREM sleep before averaging across sessions. d, Top: example scatter plot for theta/delta ratio and neck EMG power. Bottom: example scatter plot for low theta/high theta ratio and facial EMG power. Each dot represents data in a 5-s bin in the example recording session in (b). Data from NREM sleep in 100 s preceding the two REM episodes were also shown. e, Normalized EEG power spectra during qREM and aREM, averaged from 4 mice. Red dots indicate frequency bins that are significantly different between the two substages, at P < 0.05 (two-sided paired t test with Bonferroni multiple comparison correction). f, Normalized neck EMG (left) or facial EMG (right) in qREM and aREM. The EMG amplitude was normalized by that of NREM sleep before averaged across recording sessions (n = 6 sessions from 4 mice). Each line represents one recording session. P(Neck EMG) = 0.85; P(Facial EMG) = 0.0064. g, Heart rate variability (HRV), measured by relative RR intervals, in qREM and aREM substages (n = 6 sessions from 4 mice, P = 0.033). Each line represents one recording session. *P < 0.05, two-sided paired t test. Error bars or shadings, SEM.
Extended Data Fig. 7 Optogenetic inactivation altered cortical neuronal dynamics.
a, Images showing expression of AAV-CaMKIIα -NpHR3.0-mCherry in the RSC of Thy1-GCaMP6s mouse. Left: top view of wide-field imaging through a cortex-wide transparent window, excited by 593 nm light. Scale bar: 500 μm. Middle: a fluorescence image of a coronal brain slice showing mCherry expression. Scale bar: 2 mm. Right: image with higher magnification of the region in red box. Scale bar: 200 μm. b, Average Ca2+ activity across brain states in RSCREM inactivation mice (6 sessions from 4 mice). c, Activity difference between REM sleep and wake state, with statistical significance (two-sided paired t test) plotted against difference in average Ca2+ ∆F/F. Error bars, SEM. d, Time courses of 2 new motifs in RSCREM inactivation mice showing spatiotemporal patterns of neuronal Ca2+ activity across dorsal cortex. The intensity scale is normalized separately for each motif. e, Matrix of Granger causality between each pair of 11 cortical areas during REM sleep, in NPHR-expressing mice without light stimulation. Noted that the uppermost row is causality from RSC to other 10 modules. Causality with P < 0.01 (Granger’s F-test) was shown after correction for multiple comparisons by false discovery rate. f, Left, same as (e), but for Granger causality in mice with NPHR-mediated RSCREM inactivation. Right, Summary of causality from FrA in RSCREM inactivation mice.
Extended Data Fig. 8 Effects of optogenetic inactivation of RSC during REM sleep, aREM substage, or NREM sleep.
a, Schematic for close-loop optogenetic inactivation during REM sleep based on pupil size change. b, Left: schematic for optogenetic inactivation through a transparent window covering RSC bilaterally. Right: top view wide-field image and fluorescence image showing GtACR2-EGFP expression in RSC. Scale bar: 200 μm. c-d, Duration of REM substages (c), and episode duration or episode number of total REM sleep (d) in light control (6 sessions from 4 mice) and RSCREM inactivation group (8 sessions from 3 mice). Each dot represents averaged data from one recording session. P(qREM duration)= 0.00047; P(aREM duration)= 0.00071; P(REM duration)=0.027; P(REM episode)=0.07. *P < 0.05, ***P < 0.001, two-sided unpaired t test. e, Schematic for close-loop optogenetic inactivation during aREM substage. f, EEG spectrogram, EMG trace and color-coded brain states from an example RSCaREM inactivation recording session. Blue shadings show laser period. g-h, similar to c-d, but for RSCaREM inactivation. GtACR2 control: 10 sessions from 4 mice, RSCaREM inactivation: 8 sessions from 3 mice. P(qREM duration)=0.98; P(aREM duration)=0.00042; P(REM duration)=0.00053; P(REM episode number)=0.0037. **P < 0.01, ***P < 0.001, two-sided unpaired t test. i, Color-coded brain states for all NREM trials (with the onset of 1-min blue laser fell in NREM sleep) in light control (14 sessions from 7 mice) and RSCNREM inactivation group (14 sessions from 5 mice). Vertical lines represent the laser onset and offset. j, Cumulative probability for wake generation or REM sleep generation during 60-s laser stimulation. P > 0.8, bootstrap, 10000 iteration. Blue shading represents laser period. Error bars and shadings, SEM.
Extended Data Fig. 9 Optogenetic inactivation of M2 or ACC during REM sleep.
a, Schematic for REM sleep-specific optogenetic inactivation. b, Top: schematic for optogenetic inactivation through a transparent window covering M2 bilaterally. Bottom: fluorescence image showing GtACR2-EGFP expression in M2. c, Top, schematic for optogenetic inactivation through optic fibers; Bottom, fluorescence image showing GtACR2-EGFP expression. Scale bars: 200 μm. d-e, Duration of REM substages (d), and episode duration or episode number of total REM sleep (e) in GtACR2 control (10 sessions from 6 mice), ACCREM inactivation (10 sessions from 3 mice), and M2REM inactivation group (12 sessions from 5 mice). Each dot represents averaged data from one recording session. P > 0.05 for all comparisons between the inactivation and control groups, two-sided unpaired t test. Error bars, SEM.
Extended Data Fig. 10
Whole-brain mapping of inputs to the RSC and M2 a, Schematic for injection of retrograde tracer in RSC, and fluorescence image showing the injection site. Inset shows higher magnification of the region in red box. Scale bar: 200 μm. b, Similar to a, but for M2. c, Example of whole-brain reconstruction of retrograde-labelled cells (also referred to as ‘inputs’) from RSC (red) or M2 (blue). Injection site is excluded. d, Percentages of labelled cells (referred to as ‘inputs’) in the thalamocortical areas (top) and subcortical areas (bottom), normalized by the total number of labelled cells for each mouse. Each dot represents data from one mouse (RSC: n = 3; M2: n = 3). Orb., orbital area; Olf., olfaction areas; Insu., insula area; Som.M. somatomotor areas; Som. S., somatosensory areas; Gus., gustatory areas; Aud., auditory areas; Vis., visual areas; Thal., thalamus; Hipp., hippocampal region; Retrohipp., retrohippocampal region; Cort. Subplate, cortical subplate; Hypoth., hypothalamus. Error bars, SEM. e, Summary showing the preference for each region projecting to the RSC. In each pair of tracing experiments (n = 3 pairs), the RSC-preferring index for each region was calculated as (input to RSC-input to M2) / (input to RSC + input to M2). *PACC = 0.034, PPons = 0.017, PRetrohipp. = 0.025, PInsu = 0.045, **PSom.S. = 0.0043, ***PGus.= 0.00015, two-sided paired t test. Error bars, SEM. f, Fluorescence images showing retrograde-labelled neurons projecting to M2 (top) or RSC (bottom). Cc: corpus callosum; PCG: pontine central gray; V4: fourth ventricle; mev: midbrain trigeminal nucleus; PB: parabrachial nucleus; Subd: dorsal part of the subiculum; CLA: claustrum. Scale bar: 100 μm.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5
Supplementary Video 1
Mouse facial behaviors during REM substages: Video recording of mouse facial movements, color-coded sleep states and CC matrix for HOGs of video frames (similar as in Fig. 3g). Text above the mouse video represents the state of the current video frame. The blue light in the video recording depicts excitation light for wide-field imaging. The video is shown at five times the original speed.
Supplementary Video 2
Optogenetic inactivation of RSC neurons inhibits aREM initiation: Video recording of mouse facial movements, color-coded sleep states and CC matrices for HOGs of video frames in a GtACR2 control mouse (top) and in a RSCREM inactivation mouse (bottom). The scales of the CC color bar are the same for control and inactivation mice. Text represents the state of the current video frame. The onset of light was ~9 s before REM sleep and can be visualized in the video frames. The video is shown at five times the original speed.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, Y., Li, J., Zhou, M. et al. Cortical regulation of two-stage rapid eye movement sleep. Nat Neurosci 25, 1675–1682 (2022). https://doi.org/10.1038/s41593-022-01195-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01195-2
This article is cited by
-
Diverse and asymmetric patterns of single-neuron projectome in regulating interhemispheric connectivity
Nature Communications (2024)
-
40 Hz light flickering promotes sleep through cortical adenosine signaling
Cell Research (2024)
-
SlumberNet: deep learning classification of sleep stages using residual neural networks
Scientific Reports (2024)
-
Prefrontal cortical regulation of REM sleep
Nature Neuroscience (2023)
-
Sleep fMRI with simultaneous electrophysiology at 9.4 T in male mice
Nature Communications (2023)