Abstract
Human genetic data indicate that microglial dysfunction contributes to the pathology of Alzheimer’s disease (AD), exemplified by the identification of coding variants in triggering receptor expressed on myeloid cells 2 (TREM2) and, more recently, in PLCG2, a phospholipase-encoding gene expressed in microglia. Although studies in mouse models have implicated specific Trem2-dependent microglial functions in AD, the underlying molecular mechanisms and translatability to human disease remain poorly defined. In this study, we used genetically engineered human induced pluripotent stem cell-derived microglia-like cells to show that TREM2 signals through PLCγ2 to mediate cell survival, phagocytosis, processing of neuronal debris, and lipid metabolism. Loss of TREM2 or PLCγ2 signaling leads to a shared signature of transcriptional dysregulation that underlies these phenotypes. Independent of TREM2, PLCγ2 also signals downstream of Toll-like receptors to mediate inflammatory responses. Therefore, PLCγ2 activity regulates divergent microglial functions via distinct TREM2-dependent and -independent signaling and might be involved in the transition to a microglial state associated with neurodegenerative disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw and processed RNA sequencing data are available via the National Center for Biotechnology Gene Expression Omnibus repository under accession GSE144120. The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
RNA sequencing data were processed using publicly available open-source software. Sequencing adapters were trimmed using skewer58 (version 0.2.2) with default parameters. UMIs were extracted from each read with the umi2index tool (Lexogen). Quality control of the trimmed reads was performed using FastQC (version 0.11.5; https://www.bioinformatics.babraham.ac.uk/projects/fastqc). Reads were aligned to the human genome version GRCh38_p12 with STAR index (version 2.5.3a)59. The STAR index was built with the --sjdbOverhang=50 argument. Splice junctions from Gencode gene models (release_28) were provided via the --sjdbGTFfile argument. STAR alignments were generated with the following parameters: --outFilterType BySJout, --quantMode TranscriptomeSAM, -- outFilterIntronMotifs RemoveNoncanonicalUnannotated, --outSAMstrandField intronMotif, -- outSAMattributes NH HI AS nM MD XS and --outSAMunmapped Within. Alignments were obtained with the following parameters: --readFilesCommand zcat --outFilterType BySJout -- outFilterMultimapNmax 20 --alignSJoverhangMin 8 --alignSJDBoverhangMin 1 -- outFilterMismatchNmax 999 - outFilterMismatchNoverLmax 0.6 --alignIntronMin 20 -- alignIntronMax 1000000 --alignMatesGapMax 1000000 --quantMode GeneCounts -- outSAMunmapped Within --outSAMattributes NH HI AS nM MD XS –outSAMstrandField intronMotif --outSAMtype BAM SortedByCoordinate --outBAMcompression 6. Alignments sharing the same UMI and genomic coordinate were de-duplicated using the collapse_UMI_bam tool (Lexogen). Gene level counts were obtained using featureCounts from the subread package (version 1.6.2)60. Gene symbols and Entrez gene identifiers were mapped using Ensembl (version 91) via the biomaRt R package (version 2.34.2)61 using R (version 3.4.3). To identify differentially expressed genes, linear models were fit using the limma Bioconductor package62. Only genes with sufficiently large counts, as determined by edgeR’s ‘filterByExpr’ function, were included in the statistical analysis.
References
Efthymiou, A. G. & Goate, A. M. Late onset Alzheimer’s disease genetics implicates microglial pathways in disease risk. Mol. Neurodegener. 12, 43 (2017).
Hansen, D. V., Hanson, J. E. & Sheng, M. Microglia in Alzheimer’s disease. J. Cell Biol. 217, 459–472 (2018).
Condello, C., Yuan, P., Schain, A. & Grutzendler, J. Microglia constitute a barrier that prevents neurotoxic protofibrillar Αβ42 hotspots around plaques. Nat. Commun. 6, 6176 (2015).
Nugent, A. A. et al. TREM2 regulates microglial cholesterol metabolism upon chronic phagocytic challenge. Neuron 105, 837–854 (2020).
Heppner, F. L., Ransohoff, R. M. & Becher, B. Immune attack: the role of inflammation in Alzheimer disease. Nat. Rev. Neurosci. 16, 358–372 (2015).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Ulrich, J. D., Ulland, T. K., Colonna, M. & Holtzman, D. M. Elucidating the role of TREM2 in Alzheimer’s disease. Neuron 94 (2017).
Kober, D. L. et al. Neurodegenerative disease mutations in TREM2 reveal a functional surface and distinct loss-of-function mechanisms. eLife 5, e20391 (2016).
Sudom, A. et al. Molecular basis for the loss-of-function effects of the Alzheimer’s disease-associated R47H variant of the immune receptor TREM2. J. Biol. Chem. 293, 12634–12646 (2018).
Wang, Y. et al. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 160, 1061–1071 (2015).
Ulland, T. K. & Colonna, M. TREM2 - a key player in microglial biology and Alzheimer disease. Nat. Rev. Neurol. 14, 667–675 (2018).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Jay, T. R. et al. Disease progression-dependent effects of trem2 deficiency in a mouse model of Alzheimer’s disease. J. Neurosci. 37, 637–647 (2017).
Wang, Y. et al. TREM2-mediated early microglial response limits diffusion and toxicity of amyloid plaques. J. Exp. Med. 213, 667–675 (2016).
Krasemann, S. et al. The TREM2–APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Kadamur, G. & Ross, E. M. Mammalian phospholipase C. Annu. Rev. Physiol. 75, 127–154 (2013).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Magno, L. et al. Alzheimer’s disease phospholipase C-gamma-2 (PLCG2) protective variant is a functional hypermorph. Alzheimers Res. Ther. 11, 16 (2019).
Mao, D., Epple, H., Uthgenannt, B., Novack, D. V. & Faccio, R. PLCgamma2 regulates osteoclastogenesis via its interaction with ITAM proteins and GAB2. J. Clin. Invest. 116, 2869–2879 (2006).
Wilde, J. I. & Watson, S. P. Regulation of phospholipase C gamma isoforms in haematopoietic cells: why one, not the other? Cell Signal 13, 691–701 (2001).
Chiang, C. Y., Veckman, V., Limmer, K. & David, M. Phospholipase Cgamma-2 and intracellular calcium are required for lipopolysaccharide-induced Toll-like receptor 4 (TLR4) endocytosis and interferon regulatory factor 3 (IRF3) activation. J. Biol. Chem. 287 (2012).
Zhou, Q. et al. A hypermorphic missense mutation in PLCG2, encoding phospholipase Cgamma2, causes a dominantly inherited autoinflammatory disease with immunodeficiency. Am. J. Hum. Genet. 91, 713–720 (2012).
Friedman, B. A. et al. Diverse brain myeloid expression profiles reveal distinct microglial activation states and aspects of Alzheimer’s disease not evident in mouse models. Cell Rep. 22, 832–847 (2018).
Galatro, T. F. et al. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat. Neurosci. 20, 1162–1171 (2017).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Olah, M. et al. A transcriptomic atlas of aged human microglia. Nat. Commun. 9, 539 (2018).
Abud, E. M. et al. iPSC-derived human microglia-like cells to study neurological diseases. Neuron 94, 278–293 (2017).
Muffat, J. et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nat. Med. 22, 1358–1367 (2016).
Pandya, H. et al. Differentiation of human and murine induced pluripotent stem cells to microglia-like cells. Nat. Neurosci. 20, 753–759 (2017).
McQuade, A. et al. Development and validation of a simplified method to generate human microglia from pluripotent stem cells. Mol. Neurodegener. 13, 67 (2018).
Takahashi, K., Rochford, C. D. & Neumann, H. Clearance of apoptotic neurons without inflammation by microglial triggering receptor expressed on myeloid cells-2. J. Exp. Med. 201, 647–657 (2005).
Ulland, T. K. et al. TREM2 maintains microglial metabolic fitness in Alzheimer’s disease. Cell 170, 649–663 (2017).
Chae, J. J. et al. Connecting two pathways through Ca2+ signaling: NLRP3 inflammasome activation induced by a hypermorphic PLCG2 mutation. Arthritis Rheumatol. 67, 563–567 (2015).
Poliani, P. L. et al. TREM2 sustains microglial expansion during aging and response to demyelination. J. Clin. Invest. 125, 2161–2170 (2015).
Ghosh, S. Macrophage cholesterol homeostasis and metabolic diseases: critical role of cholesteryl ester mobilization. Expert Rev. Cardiovasc. Ther. 9, 329–340 (2011).
Cheng-Hathaway, P. J. et al. The Trem2 R47H variant confers loss-of-function-like phenotypes in Alzheimer’s disease. Mol. Neurodegener. 13, 29 (2018).
Yang, J. et al. Pathological axonal death through a MAPK cascade that triggers a local energy deficit. Cell 160, 161–176 (2015).
Bae, Y. S., Lee, H. Y., Jung, Y. S., Lee, M. & Suh, P. G. Phospholipase Cgamma in Toll-like receptor-mediated inflammation and innate immunity. Adv. Biol. Regul. 63, 92–97 (2017).
Zhu, L., Jones, C. & Zhang, G. The role of phospholipase C signaling in macrophage-mediated inflammatory response. J. Immunol. Res. 2018 (2018).
Marschallinger, J. et al. Lipid-droplet-accumulating microglia represent a dysfunctional and proinflammatory state in the aging brain. Nat. Neurosci. 23, 194–208 (2020).
Kober, D. L. & Brett, T. J. TREM2-ligand interactions in health and disease. J. Mol. Biol. 429, 1607–1629 (2017).
Cantoni, C. et al. TREM2 regulates microglial cell activation in response to demyelination in vivo. Acta Neuropathol. 129, 429–447 (2015).
Pluvinage, J. V. et al. CD22 blockade restores homeostatic microglial phagocytosis in ageing brains. Nature 568, 187–192 (2019).
Heneka, M. T. et al. NLRP3 is activated in Alzheimer’s disease and contributes to pathology in APP/PS1 mice. Nature 493, 674–678 (2013).
Heneka, M. T., Kummer, M. P. & Latz, E. Innate immune activation in neurodegenerative disease. Nat. Rev. Immunol. 14, 463–477 (2014).
Foley, P. Lipids in Alzheimer’s disease: a century-old story. Biochim. Biophys. Acta 1801, 750–753 (2010).
Lue, L. F. et al. Inflammatory repertoire of Alzheimer’s disease and nondemented elderly microglia in vitro. Glia 35, 72–79 (2001).
Schlepckow, K. et al. Enhancing protective microglial activities with a dual function TREM2 antibody to the stalk region. EMBO Mol. Med. 12, e11227 (2020).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Huntwork-Rodriguez, S. et al. JNK-mediated phosphorylation of DLK suppresses its ubiquitination to promote neuronal apoptosis. J. Cell Biol. 202, 747–763 (2013).
Safaiyan, S. et al. Age-related myelin degradation burdens the clearance function of microglia during aging. Nat. Neurosci. 19, 995–998 (2016).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Kucukural, A., Yukselen, O., Ozata, D. M., Moore, M. J. & Garber, M. DEBrowser: interactive differential expression analysis and visualization tool for count data. BMC Genomics 20, 6 (2019).
Babicki, S. et al. Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res. 44, W147–153 (2016).
Liao, Y., Wang, J., Jaehnig, E. J., Shi, Z. & Zhang, B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–W205 (2019).
Jiang, H., Lei, R., Ding, S. W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinforma. 15, 182 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Durinck, S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Liu, R. et al. Why weight? Modelling sample and observational level variability improves power in RNA-seq analyses. Nucleic Acids Res. 43, e97 (2015).
Acknowledgements
We thank members of Lewcock laboratory and Discovery Biology at Denali Therapeutics for reading of the manuscript and helpful discussions. We thank E. Chow and the UCSF Center for Advanced Technology for their expertise and assistance in generating the QuantSeq data.
Author information
Authors and Affiliations
Contributions
Conceptualization: B.J.A, L.P., G.D.P. and J.W.L.; methodology: B.J.A., L.P., A.R., J.S. and K.M.M.; investigation: B.J.A., L.P., C.L., A.R., S.S.D., B.v.L., Y.M. and K.M.M.; formal analysis: B.J.A., L.P., K.L., T.S. and G.A.; writing—original draft: B.J.A and L.P.; writing—review and editing: B.J.A, L.P., J.W.L., T.S. and G.D.P.
Corresponding authors
Ethics declarations
Competing interests
All authors are paid employees and shareholders of Denali Therapeutics Inc. Denali has filed patent applications related to the subject matter of this paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation and validation of wildtype and KO iPSC-derived microglia (iMG).
a, Timeline and strategy for robust generation of iMG that accurately model primary human microglia. b, Immunoblot shows NANOG protein expression in iPSCs, but absence in human astrocytes and iMG. Blots are cropped to show indicated protein bands. n = 3 biologically independent experiments, with similar results. c, Representative phase images show similar microglia morphology between primary human microglia and iMG. n = 3 biologically independent experiments, with similar results. Scale bar, 50 µm. d, Immunostaining shows expression of several microglia markers in iMG. n = 3 biologically independent experiments, with similar results. Scale bar, 100 µm. e, Image showing live iMG stained with CD11b-647 cultured under conditions that induce a homeostatic, ramified morphology. n = 3 biologically independent experiments, with similar results. Scale bar, 25 µm. f, Sanger sequencing traces show successful indel generation in the TREM2 locus via CRISPR/Cas9. n = 12 biologically independent samples, with similar results. g, Karyotyping analysis of undifferentiated iPSC parental line and TREM2 KO clone. n = 3 biologically independent experiments. h, TREM2 immunostaining shows protein expression in WT iMG, but absence in TREM2 KO iMG. n = 4 biologically independent experiments, with similar results. Scale bar, 25 µm. i, Fraction of PU.1-positive cells in WT and TREM2 KO iMG cultures, quantified via PU.1 and DAPI immunofluorescence. n = 4 biologically independent experiments. j, Sanger sequencing traces show successful indel generation in the PLCG2 locus via CRISPR/Cas9. n = 12 biologically independent samples, with similar results. k, Karyotyping analysis of undifferentiated iPSC parental line and PLCG2 KO clone. n = 3 biologically independent experiments. l, Full immunoblot shows PLCγ2 protein expression (red arrow) in WT iMG, but absence in PLCG2 KO iMG. n = 4 biologically independent experiments, with similar results. m, Fraction of PU.1-positive cells in WT and PLCG2 KO iMG cultures, quantified via PU.1 and DAPI immunofluorescence. PLCG2 KO, n = 3 biologically independent experiments; WT, n = 4 biologically independent experiments. All data are mean ± s.e.m.
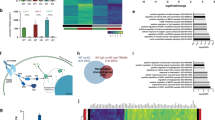
Extended Data Fig. 2 Expression differences between iMG genotypes in baseline conditions.
a, Gene ontology analysis of genes altered in both TREM2 and PLCG2 KO iMG compared to wildtype iMG, sorted by P-value ranking. Pathways were identified using Over-Representation Analysis using the Benjamini-Hochberg procedure to calculate FDR. n = 3 biologically independent experiments. b, Fluidigm mRNA analysis of indicated iMG genotypes in baseline culture conditions. Heatmap and mRNA expression fold changes of genes identified to be downregulated in both TREM2 and PLCG2 KO iMG. n = 3 biologically independent experiments. Adjusted P-values are presented; one-way ANOVA, post-hoc Tukey test for multiple comparisons. c, Regression plot showing the PLCG2 KO/WT ratio (x-axis) compared to the TREM2 KO/WT ratio (y-axis) of gene expression level measured via Fluidigm. Each dot represents the –log2 transformed fold change values for a single gene. n = 3 biologically independent experiments.
Extended Data Fig. 3 Generation and characterization of independent KO clones for TREM2 and PLCG2.
a Sanger sequencing traces show successful indel generation in the TREM2 locus via CRISPR/Cas9 in KO clone 2. n = 12 biologically independent samples, with similar results. b, TREM2-APC flow cytometry plot shows protein expression in WT iMG, but absence in TREM2 KO clone 2 iMG. n = 3 biologically independent experiments, with similar results. c, Sanger sequencing traces show successful indel generation in the PLCG2 locus via CRISPR/Cas9 in KO clone 2. n = 12 biologically independent samples, with similar results. d, Immunoblot shows PLCγ2 protein expression in WT iMG, but absence in PLCG2 KO clone 2 iMG. Blots are cropped to show indicated protein bands. n = 3 biologically independent experiments, with similar results. e, Relative mRNA expression levels in indicated iMG populations. Each sample was normalized to an internal GAPDH control and all samples are normalized to WT iMG. n = 4 biologically independent experiments. f, iMG survival in indicated iMG genotypes after 7 days of MCSF withdrawal, quantified as the ratio of Cell-Titer Glo luminescence at day 7 to day 0. TREM2 KO clone 2, n = 3 biologically independent experiments; all other genotypes, n = 4 biologically independent experiments. All data are mean ± s.e.m. For (e and f), one-way ANOVA, post-hoc Tukey test for multiple comparisons.
Extended Data Fig. 4 Activity of SYK inhibitor and gene epistasis profiling.
a, Levels of SYK phosphorylated at Y525/526 in WT iMG stimulated with 70% PC/30% PS liposomes (1 mg/ml, 5 min) in the presence or absence of pretreatment with SYK inhibitor (1 μM, 1 hour). n = 3 biologically independent experiments. b, Relative mRNA expression levels in WT iMG nucleofected with indicated siRNA pools. Each sample was normalized to an internal GAPDH control and all samples are normalized to WT iMG treated with a non-targeting control (NTC). PLIN2, LIPA, APOC1, FABP5, n = 3 biologically independent experiments; ITGAM, TREM2, PLCG2, LPL, CD52, n = 4 biologically independent experiments. c, Relative mRNA expression levels in an independent parental line of WT iMG nucleofected with indicated siRNA pools. Each sample was normalized to an internal GAPDH control and all samples are normalized to the corresponding parental WT iMG treated with a non-targeting control (NTC). n = 3 biologically independent experiments. All data are mean ± s.e.m. For (a), two-tailed, unpaired Student’s t-test. For (b-c), one-way ANOVA, post-hoc Tukey test for multiple comparisons.
Extended Data Fig. 5 Multiple TREM2 and PLCG2 KO iMG clones display shared lipid accumulation defects upon myelin challenge.
a, Lipidomic analysis of free cholesterol (left), cholesteryl ester (middle), and ceramide (right) species in second, independent TREM2 and PLCG2 KO iMG clones treated with vehicle or myelin (25 µg/ml) for 48 hours. Mean lipid abundance value for each species is normalized to cell number and presented on a log2 transformed scale. n = 4 biologically independent experiments, except PLCG2 KO conditions for ceramide species, n = 3 biologically independent experiments. b, Lipidomic analysis of hexosylceramide (left) and diacylglycerol (right) species in indicated iMG genotypes and clones treated with vehicle or myelin (25 µg/ml) for 48 hours. Mean lipid abundance value for each species is normalized to cell number and presented on a log2 transformed scale. n = 3 biologically independent experiments, except WT and TREM2 KO clone 2 conditions, n = 4 biologically independent experiments. c, Relative TREM2 mRNA expression levels in indicated iMG populations. n = 4 biologically independent experiments. d, Flow cytometry histogram shows TREM2 surface expression in TREM2(R47H) iMG. n = 3 biologically independent experiments, with similar results. e, Relative fraction of cells with pHrodo signal remaining 96 hours after treatment with pHrodo-myelin in the indicated iMG populations, normalized to WT as 1. TREM2(R47H), n = 2 biologically independent experiments; TREM2 HET, n = 3 biologically independent experiments; WT, TREM2 KO, n = 4 biologically independent experiments. f, Relative PLCG2 mRNA expression levels in indicated iMG populations. n = 4 biologically independent experiments. All data are mean ± s.e.m. For (a-b), two-way ANOVA, genotype effect between treatment populations, post-hoc Tukey test for multiple comparisons. All statistical tests were performed on log2 transformed abundance values. OE, transgene expression.
Extended Data Fig. 6 iMG exhibit lipid accumulation when co-cultured with post-axotomy neurons.
a, Composite images of DRG neuronal explants before (left) and after (right) axotomy. n = 4 biologically independent experiments, with similar results. Scale bar, 500 µm. b, mCherry (yellow) fluorescence in neurons co-cultured with WT (top), PLCG2 KO (middle), or TREM2 KO (bottom) iMG without axotomy shows no neuronal degeneration at 0, 9, and 15 hours. n = 4 DRG explants from independent animals per condition, with similar results. Scale bar, 50 µm. c, Timecourse of neuronal debris accumulation within iMG in indicated co-culture conditions, quantified as percentage of mCherry+ iMG. n = 4 DRG explants from independent animals per condition. Data are mean ± s.e.m.; two-way repeated measures ANOVA, post-hoc Dunnett’s test for multiple comparisons. d, BODIPY staining (green) of neutral lipids shows accumulation in iMG co-cultured with DRG neurons from 0 to 18 hours post-axotomy. n = 4 biologically independent experiments, with similar results. Scale bar, 50 µm.
Extended Data Fig. 7 Expression differences between iMG genotypes upon zymosan challenge.
Fluidigm mRNA analysis of indicated iMG genotypes treated with vehicle or zymosan (50 µg/ml) for 24 hours. Heatmap and mRNA expression fold changes of cytokine and chemokine genes identified to be responsive to zymosan challenge in WT iMG. Fold change values of downregulated genes appear in red. n = 3 biologically independent experiments. Adjusted P-values are presented; one-way ANOVA, post-hoc Tukey test for multiple comparisons.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Table 1
Supplementary Tables 1–4.
Supplementary Video 1
Live capture of WT iMG stained with AF647-linked CD11b antibody and cultured under homeostatic conditions, demonstrating dynamic ramified morphology. Field was imaged every 30 min for 11 h. n = 3 biologically independent experiments, with similar results.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 1
Unprocessed Western Blots
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Unprocessed Western Blots
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 1
Unprocessed Western Blots
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed Western Blots
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Rights and permissions
About this article
Cite this article
Andreone, B.J., Przybyla, L., Llapashtica, C. et al. Alzheimer’s-associated PLCγ2 is a signaling node required for both TREM2 function and the inflammatory response in human microglia. Nat Neurosci 23, 927–938 (2020). https://doi.org/10.1038/s41593-020-0650-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0650-6
This article is cited by
-
An integrated toolkit for human microglia functional genomics
Stem Cell Research & Therapy (2024)
-
Updates on mouse models of Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
BHLHE40/41 regulate microglia and peripheral macrophage responses associated with Alzheimer’s disease and other disorders of lipid-rich tissues
Nature Communications (2024)
-
Advancements in Single-Cell RNA Sequencing Research for Neurological Diseases
Molecular Neurobiology (2024)
-
APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia
Nature (2024)