Abstract
The mechanisms by which prenatal immune activation increase the risk for neuropsychiatric disorders are unclear. Here, we generated developmental cortical interneurons (cINs)—which are known to be affected in schizophrenia (SCZ) when matured—from induced pluripotent stem cells (iPSCs) derived from healthy controls (HCs) and individuals with SCZ and co-cultured them with or without activated microglia. Co-culture with activated microglia disturbed metabolic pathways, as indicated by unbiased transcriptome analyses, and impaired mitochondrial function, arborization, synapse formation and synaptic GABA release. Deficits in mitochondrial function and arborization were reversed by alpha lipoic acid and acetyl-l-carnitine treatments, which boost mitochondrial function. Notably, activated-microglia-conditioned medium altered metabolism in cINs and iPSCs from HCs but not in iPSCs from individuals with SCZ or in glutamatergic neurons. After removal of activated-microglia-conditioned medium, SCZ cINs but not HC cINs showed prolonged metabolic deficits, which suggests that there is an interaction between SCZ genetic backgrounds and environmental risk factors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The RNA-seq data have been deposited at the GEO (https://www.ncbi.nlm.nih.gov/geo/) at accession numbers are GSE118313, GSE132689 and GSE152438. For the human cDNA ensemble database, we used human cDNA fasta file GRCh38 release 87, which is available at http://www.ensembl.org/info/data/ftp/index.html. The data that support the findings of this study are available from the corresponding author upon request.
References
Knuesel, I. et al. Maternal immune activation and abnormal brain development across CNS disorders. Nat. Rev. Neurol. 10, 643–660 (2014).
Estes, M. L. & McAllister, A. K. Immune mediators in the brain and peripheral tissues in autism spectrum disorder. Nat. Rev. Neurosci. 16, 469–486 (2015).
Hantsoo, L., Kornfield, S., Anguera, M. C. & Epperson, C. N. Inflammation: a proposed intermediary between maternal stress and offspring neuropsychiatric risk. Biol. Psychiatry 85, 97–106 (2019).
Estes, M. L. & McAllister, A. K. Maternal immune activation: implications for neuropsychiatric disorders. Science 353, 772–777 (2016).
Mallard, C., Welin, A. K., Peebles, D., Hagberg, H. & Kjellmer, I. White matter injury following systemic endotoxemia or asphyxia in the fetal sheep. Neurochem. Res. 28, 215–223 (2003).
Kuypers, E. et al. Effects of intra-amniotic lipopolysaccharide and maternal betamethasone on brain inflammation in fetal sheep. PLoS ONE 8, e81644 (2013).
Hutton, L. C., Castillo-Melendez, M., Smythe, G. A. & Walker, D. W. Microglial activation, macrophage infiltration, and evidence of cell death in the fetal brain after uteroplacental administration of lipopolysaccharide in sheep in late gestation. Am. J. Obstet. Gynecol. 198, 117.e1–117.e11 (2008).
Benes, F. M. The GABA system in schizophrenia: cells, molecules and microcircuitry. Schizophr. Res. 167, 1–3 (2015).
Lewis, D. A., Hashimoto, T. & Volk, D. W. Cortical inhibitory neurons and schizophrenia. Nat. Rev. Neurosci. 6, 312–324 (2005).
Volk, D. W. & Lewis, D. A. Early developmental disturbances of cortical inhibitory neurons: contribution to cognitive deficits in schizophrenia. Schizophr. Bull. 40, 952–957 (2014).
Lewis, D. A. et al. Subunit-selective modulation of GABA type A receptor neurotransmission and cognition in schizophrenia. Am. J. Psychiatry 165, 1585–1593 (2008).
Hilker, R. et al. Heritability of schizophrenia and schizophrenia spectrum based on the nationwide Danish twin register. Biol. Psychiatry 83, 492–498 (2018).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Windrem, M. S. et al. Human iPSC glial mouse chimeras reveal glial contributions to schizophrenia. Cell Stem Cell 21, 195–208.e6 (2017).
Noh, H., Shao, Z. C., Coyle, J. T. & Chung, S. M. Modeling schizophrenia pathogenesis using patient-derived induced pluripotent stem cells (iPSCs). Biochim. Biophys. Acta Mol. Basis Dis. 1863, 2382–2387 (2017).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science https://doi.org/10.1126/science.aat8464 (2018).
Shao, Z. et al. Dysregulated protocadherin-pathway activity as an intrinsic defect in induced pluripotent stem cell-derived cortical interneurons from subjects with schizophrenia. Nat. Neurosci. 22, 229–242 (2019).
Ahn, S., Kim, T. G., Kim, K. S. & Chung, S. Differentiation of human pluripotent stem cells into medial ganglionic eminence vs. caudal ganglionic eminence cells. Methods 101, 103–112 (2016).
Ni, P. et al. iPSC-derived homogeneous populations of developing schizophrenia cortical interneurons have compromised mitochondrial function. Mol. Psychiatry https://doi.org/10.1038/s41380-019-0423-3 (2019).
Ni, P. & Chung, S. Mitochondrial dysfunction in schizophrenia. Bioessays 42, e1900202 (2020).
Choi, G. B. et al. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 351, 933–939 (2016).
Kim, S. et al. Maternal gut bacteria promote neurodevelopmental abnormalities in mouse offspring. Nature 549, 528–532 (2017).
Kim, T. G. et al. Efficient specification of interneurons from human pluripotent stem cells by dorsoventral and rostrocaudal modulation. Stem Cells 32, 1789–1804 (2014).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science https://doi.org/10.1126/science.aat8127 (2018).
Jun, J. I. & Lau, L. F. Taking aim at the extracellular matrix: CCN proteins as emerging therapeutic targets. Nat. Rev. Drug Discov. 10, 945–963 (2011).
McClain, J. A., Phillips, L. L. & Fillmore, H. L. Increased MMP-3 and CTGF expression during lipopolysaccharide-induced dopaminergic neurodegeneration. Neurosci. Lett. 460, 27–31 (2009).
Meyer, U. Prenatal poly(i:C) exposure and other developmental immune activation models in rodent systems. Biol. Psychiatry 75, 307–315 (2014).
Kong, P. et al. Thrombospondin-1 regulates adiposity and metabolic dysfunction in diet-induced obesity enhancing adipose inflammation and stimulating adipocyte proliferation. Am. J. Physiol. Endocrinol. Metab. 305, E439–E450 (2013).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Cai, Q. & Tammineni, P. Mitochondrial aspects of synaptic dysfunction in Alzheimer’s disease. J. Alzheimer’s Dis. 57, 1087–1103 (2017).
Mendoza, E. et al. In vivo mitochondrial inhibition alters corticostriatal synaptic function and the modulatory effects of neurotrophins. Neuroscience 280, 156–170 (2014).
Du, H. et al. Early deficits in synaptic mitochondria in an Alzheimer’s disease mouse model. Proc. Natl Acad. Sci. USA 107, 18670–18675 (2010).
Oishi, Y. et al. SUMOylation of Kruppel-like transcription factor 5 acts as a molecular switch in transcriptional programs of lipid metabolism involving PPAR-δ. Nat. Med. 14, 656–666 (2008).
Drosatos, K. et al. Cardiac myocyte KLF5 regulates Ppara expression and cardiac function. Circ. Res. 118, 241–253 (2016).
Dello Russo, C. et al. The human microglial HMC3 cell line: where do we stand? A systematic literature review. J. Neuroinflammation 15, 259 (2018).
Gumusoglu, S. B. & Stevens, H. E. Maternal inflammation and neurodevelopmental programming: a review of preclinical outcomes and implications for translational psychiatry. Biol. Psychiatry 85, 107–121 (2019).
Tsuyama, T., Tsubouchi, A., Usui, T., Imamura, H. & Uemura, T. Mitochondrial dysfunction induces dendritic loss via eIF2α phosphorylation. J. Cell Biol. 216, 815–834 (2017).
Montalvo, G. B., Artalejo, A. R. & Gilabert, J. A. ATP from subplasmalemmal mitochondria controls Ca2+-dependent inactivation of CRAC channels. J. Biol. Chem. 281, 35616–35623 (2006).
Vanden Berghe, P., Kenyon, J. L. & Smith, T. K. Mitochondrial Ca2+ uptake regulates the excitability of myenteric neurons. J. Neurosci. 22, 6962–6971 (2002).
Demaurex, N., Poburko, D. & Frieden, M. Regulation of plasma membrane calcium fluxes by mitochondria. Biochim. Biophys. Acta 1787, 1383–1394 (2009).
Pei, L. & Wallace, D. C. Mitochondrial etiology of neuropsychiatric disorders. Biol. Psychiatry 83, 722–730 (2018).
Devaraju, P. et al. Haploinsufficiency of the 22q11.2 microdeletion gene Mrpl40 disrupts short-term synaptic plasticity and working memory through dysregulation of mitochondrial calcium. Mol. Psychiatry 22, 1313–1326 (2017).
Robicsek, O. et al. Isolated mitochondria transfer improves neuronal differentiation of schizophrenia-derived induced pluripotent stem cells and rescues deficits in a rat model of the disorder. Schizophr. Bull. 44, 432–442 (2018).
Inan, M. et al. Energy deficit in parvalbumin neurons leads to circuit dysfunction, impaired sensory gating and social disability. Neurobiol. Dis. 93, 35–46 (2016).
Fernandez, A. et al. Mitochondrial dysfunction leads to cortical under-connectivity and cognitive impairment. Neuron https://doi.org/10.1016/j.neuron.2019.04.013 (2019).
Lin-Hendel, E. G., McManus, M. J., Wallace, D. C., Anderson, S. A. & Golden, J. A. Differential mitochondrial requirements for radially and non-radially migrating cortical neurons: implications for mitochondrial disorders. Cell Rep. 15, 229–237 (2016).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Vuillermot, S., Luan, W., Meyer, U. & Eyles, D. Vitamin D treatment during pregnancy prevents autism-related phenotypes in a mouse model of maternal immune activation. Mol. Autism 8, 9 (2017).
Glass, C. K., Saijo, K., Winner, B., Marchetto, M. C. & Gage, F. H. Mechanisms underlying inflammation in neurodegeneration. Cell 140, 918–934 (2010).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Hong, S. et al. Functional analysis of various promoters in lentiviral vectors at different stages of in vitro differentiation of mouse embryonic stem cells. Mol. Ther. 15, 1630–1639 (2007).
Fitzmaurice, G. M., Laird, N. M. & Ware, J. H. Applied Longitudinal Analysis 2nd edn (John Wiley & Sons, 2011).
Laird, N. M. & Ware, J. H. Random-effects models for longitudinal data. Biometrics 38, 963–974 (1982).
Jensen, K. P. et al. Alcohol-responsive genes identified in human iPSC-derived neural cultures. Transl. Psychiatry 9, 96 (2019).
Acknowledgements
This study was supported by MH107884 (S.C.) and NYSTEM C32607GG (S.C.). We thank K. F. Berman and J. Apud at the National Institute of Mental Health for their contribution in providing human fibroblast samples. We thank D. R. Weinberger at the Lieber Institute for Brain Development for supplying human fibroblast samples and for critical review of the manuscript.
Author information
Authors and Affiliations
Contributions
G.-H.P., H.N., Z.S., P.N., Y.Q., B.M.C., J.T.C. and S.C. designed the experiments. G.-H.P., H.N., Z.S., P.N., Y.Q. and D.L. differentiated the cINs and analyzed them by qPCR and immunocytochemistry. G.-H.P., H.N. and P.N. analyzed oxidative phosphorylation of the cINs using a Seahorse Analyzer. G.-H.P., H.N., C.P.B., J.S.P., C.P.A., J.M.P., D.T.L., S.Z.G. and Y.G. performed arborization analyses. G.-H.P., H.N., C.P.A. and J.M.P. analyzed inhibitory synapses in cIN organoids. G.-H.P. performed and analyzed the synaptic GABA release analysis. B.M.C. and D.L.M. provided the HC and SZC cell lines and reviewed data interpretation and manuscript content. T.A.L., H.S.X., C.Y. and W.H. did the RNA-seq analysis. H.-Y.K. performed statistical analyses. G.-H.P. and S.C. wrote the manuscript. S.C. financially supported this study.
Corresponding author
Ethics declarations
Competing interests
T.A.L. and H.S.X. were employees of Pfizer, Inc. at the time this work was performed. J.T.C. reports holding a patent on the use of d-serine to treat serious mental disorders that is owned by Massachusetts General Hospital and consulting with Concert Pharm.
Additional information
Peer review information Nature Neuroscience thanks Stewart Anderson, Robert Friedlander, Urs Meyer, and the other, anonymous, reviewer for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation of cINs from human iPSCs.
a, Table of subjects analyzed in pilot study in Fig. 1. HC refers to healthy control subjects and SCZ refers to people with SCZ. b, Differentiation scheme of cINs from human iPSCs. SRM: serum replacement media, LDN: 100 nM LDN193189, SB: 10 μM SB431542, SAG: 0.1 μM Smoothened agonist, and IWP2: 5 μM Inhibitor of Wnt production-2. c, ELISA analysis of mouse IL1β release from activated microglia BV2. Data are presented as mean±SEM. One-way ANOVA followed by posthoc analysis using Tukey’s multiple comparisons test was used for analysis (p=0.00000489 and n=3 batches, f=173.8, df=2). d, Immunocytochemistry and cell counting analysis of differentiated cINs, demonstrating MGE-derived cIN phenotype with GAD1 and SOX6 expression. Scale bar=50 μm. Percentages of cells positive for each marker were quantified in relation to DAPI-stained total nuclei. For GAD1, Sample sizes (n) as the number of independent differentiations for each line are as follows; 272:4, 190:5, 367:5, 226:4, 67:4, 128:5, 212:4, and 162:5. For SOX6, sample sizes (n) as the number of independent differentiations for each line are as follows; 272:4, 190:4, 367:4, 226:3, 67:4, 128:5, 212:4, and 162:5. At least 500 cells were counted for each line. Data are presented as mean ±SEM. e, Immunocytochemistry and cell counting analysis of OLIG2+, GFAP+, CHAT+, TH+, 5-HT+ and VIP+ cells. Scale bar= 25 or 50 μm. Proportions of OLIG2+ neurons (p=0.782, two-sided chi-square test), GFAP+ neurons (Non-detectable for both HC and SCZ), CHAT+ neurons (Non-detectable for both HC and SCZ), TH+ neurons (p=0.700, two-sided chi-square test), 5-HT+ neurons (p=0.911, two-sided chi-square test) and VIP+ neurons (non-detectable for both HC and SCZ) between groups. Analysis was repeated at least three times with comparable results.
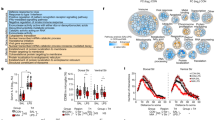
Extended Data Fig. 2 RNA-seq analysis of cINs co-cultured with activated microglia.
a, Total identified gene numbers in RNA-seq in Fig. 1. b, PCA analysis of RNA-seq analysis in Fig. 1 (from 4 HC lines and 4 SCZ lines with 3 independent differentiations from each line as shown in Extended Data Fig. 2a). c, Various phenotype marker expression in cINs with or without co-culture with activated microglia, analyzed by RNA-seq. Gene expression is shown as RPKM, obtained from STAR-featureCount. Differentially expressed genes were analyzed by Kallisto-Sleuth (Wald test for two-sided significance testing, n=24 batches from 4 HC lines and 4 SCZ lines, each line with 3 independent differentiations). Error bars are SEM. d, qPCR analysis of inflammatory response gene expression in cINs with or without activated microglia co-culture. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-Level Hierarchical Linear Mixed Effect Model after log-transformation was used for analysis (n=24 batches from 4 HC lines and 4 SCZ lines, each line with 3 independent differentiations). e, Differential expression of inflammatory response genes in HC vs SCZ postmortem PFC from Wang et al. 16 f, Differential expression of THBS1 in HC vs SCZ postmortem PFC from Wang et al. 16.
Extended Data Fig. 3 Co-culture with activated microglia results in dysregulated metabolic gene pathways in developmental cINs.
a, Expression of CTGF in HC cINs and SCZ cINs with or without co-culture with activated microglia, analyzed by qPCR analysis. Paired data from the same line are connected by a solid line. b, Expression of THBS1 in HC cINs and SCZ cINs with or without co-culture with activated microglia, analyzed by qPCR analysis. Paired data from the same line are connected by a solid line. c, Expression of KLF5 in HC cINs and SCZ cINs with or without co-culture with activated microglia in the pilot cohort, analyzed by qPCR analysis. Paired data from the same line are connected by a solid line. d, Expression of KLF5 in HC cINs and SCZ cINs with or without co-culture with activated microglia in the replication cohort, analyzed by qPCR analysis. Paired data from the same line are connected by a solid line.
Extended Data Fig. 4 Generation of cINs from human replication cohort iPSCs.
a, Table of subjects analyzed as a replication cohort. HC refers to healthy control subjects and SCZ refers to people with SCZ. b, Immunocytochemistry and cell counting analysis of generated cINs for expression of GAD1 and SOX6, analyzed after 8 weeks’ differentiation. Scale bar= 50 μm. Percentages of cells positive for each marker were quantified in relation to DAPI-stained total nuclei. For GAD1, Sample sizes (n) as the number of independent differentiations for each line are as follows; PYAUM:4, 317:3, L9:3, L7:4, L5:4, 292:3, 689:4, 285:4, 282:4, 58:4, L10:3, and L8:3. For SOX6, sample sizes (n) as the number of independent differentiations for each line are as follows; PYAUM:5 317:3, L9:3, L7:4, L5:3, 292:3, 689:4, 285:4, 282:4, 58:3, L10:3, and L8:3. At least 500 cells were counted for each line. Data are presented as mean ±SEM. c, qPCR analysis of replication cohort cINs with or without activated microglia co-culture. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-Level Hierarchical Linear Mixed Effect Model after log-transformation was used for analysis (n=30 batches from 5 HC lines and 5 SCZ lines, each line with 3 independent differentiations).
Extended Data Fig. 5 Metabolic analysis of cINs cultured in activated microglia-conditioned media.
a, Schematic diagram of Seahorse analysis OCR data. Basal Respiration=baseline OCR-Rotenone/antimycin A OCR, ATP production=Baseline OCR-Oligomycin OCR and Maximum Respiration=FCCP OCR-Rotenone/antimycin A OCR. b, Schematic diagram of Seahorse analysis of ECAR data. Glycolytic Reserve= Oligomycin ECAR- Baseline ECAR. c, Overexpression and siRNA knock down of CTGF and THBS1, probed by qPCR analysis. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed Ratio paired t-test was used for analysis (n=3 batches, O.E. of CTGF: t=19.35, df=2, O.E. of THBS1: t=47.92, df=2, siRNA of CTGF: t=18.89, df=2, siRNA of THBS1: t=17.86, df=2). d, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) of cINs cultured with or without activated microglia-conditioned media using a Seahorse Analyzer. Data are presented as mean±SEM. Two-tailed paired t-test after log transformation was used for analysis of ATP Production and Baseline Glycolysis and Two-tailed paired t-test was used for analysis of Glycolytic Reserve (n=15 lines, ATP Production: t=3.978, df=14, Baseline Glycolysis: t=0.06106, df=14, Glycolytic Reserve: t=1.534, df=14). e, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) with or without overexpression of CTGF and THBS1 using a Seahorse Analyzer. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis of ATP Production and Two-tailed paired t-test after log transformation was used for analysis of Baseline Glycolysis and Glycolytic Reserve (n=12 lines, ATP Production: t=4.186, df=11, Baseline Glycolysis: t=0.9984, df=11, Glycolytic Reserve: t=0.9361, df=11). f, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) with or without siRNA-mediated knockdown of CTGF and THBS1 using a Seahorse Analyzer. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis of ATP Production and Two-tailed paired t-test after log transformation was used for analysis of Baseline Glycolysis and Glycolytic Reserve (n=12 lines, ATP Production: t=3.164, df=11, Baseline Glycolysis: t=1.293, df=11, Glycolytic Reserve: t=1.300, df=11). g, Analysis of oxidative phosphorylation (Basal Respiration and ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in HC cINs using a Seahorse Analyzer one week after removal of activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis of Basal Respiration, ATP Production and Baseline Glycolysis. Two-tailed paired t-test after log transformation was used for analysis of Glycolytic Reserve (n=8 lines; Each data point is averaged from 1-4 independent differentiations, Basal Respiration: t=8.404, df=2, ATP Production: t=0.7116, df=7, Baseline Glycolysis: t=0.6444, df=7, Glycolytic Reserve: t=1.495, df=7). h, Analysis of oxidative phosphorylation (Basal respiration and ATP production) and glycolysis (Baseline glycolysis and glycolysis reserve) in SCZ cINs using a Seahorse Analyzer one week after removal of activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n=7 lines, Each data point is averaged from 1-2 independent differentiations, Basal Respiration: t=5.622, df=6, ATP Production: t=3.750, df=6, Baseline Glycolysis: t=0.2902, df=6, Glycolytic Reserve: t=0.1759, df=6).
Extended Data Fig. 6 Functional analysis of cINs with activated microglia-conditioned media.
a, Tracing of cINs without activated microglia-conditioned media, with activated microglia-conditioned media or with activated microglia-conditioned media + ALA/ALC treatment. Scale bar= 50 µm. Analysis was repeated at least three times with comparable results. b, Analysis of mitochondrial function using a Seahorse Analyzer in cINs cultured in activated microglia-conditioned media with or without ALA/ALC treatment. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n=10 lines, Basal Respiration: t=3.741, df=9, Maximum Respiration: t=2.987, df=9 ATP Production: t=5.281, df=9). c, Negative control for synapse staining in the absence of primary antibodies for synaptic puncta detection. Scale bar= 5 µm. Analysis was repeated at least three times with comparable results. d, Synapse analysis of cIN organoids with or without treatment with activated microglia conditioned media. Scale bar= 5 µm. Analysis was repeated at least three times with comparable results.
Extended Data Fig. 7 Gene expression analysis and functional analysis of HC and SCZ cINs with or without activated microglia co-culture.
a, qPCR analysis of CTGF and THBS1 expression in HC cINs vs. SCZ cINs with or without activated microglia co-culture. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed Unpaired t-test after log transformation was used for analysis of CTGF and THBS1 for control conditions. Two-tailed Unpaired t-test was used for analysis of CTGF and THBS1 for those with activated microglia co-culture (n=10 lines for HC and n=10 lines for SCZ, Each data point is averaged from 3 independent differentiations, CTGF of Control: t=0.7972, df=18, THBS1 of Control: t=0.7871, df=18, CTGF of Act.MG: t=0.3137, df=8, THBS1 of Act.MG: t=0.1880, df=18). b, Analysis of oxidative phosphorylation in HC cINs vs. SCZ cINs cultured with or without activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test was used for analysis of Basal Respiration and Maximum Respiration for control conditions. Two-tailed Unpaired t-test after log transformation was used for analysis of Basal Respiration for those cultured in activated microglia-conditioned media. Two-tailed Unpaired t-test was used for analysis of Maximum Respiration for those cultured in activated microglia-conditioned media (n=7 lines for HC and n=7 lines for SCZ, Basal Respiration of Control: t=2.428, df=12, Maximum Respiration of Control: t=2.517, df=12, Basal Respiration of Act.MG: t=5.424, df=12, Maximum Respiration of Act.MG: t=4.262, df=12). c, qPCR analysis of KLF5 expression in HC cINs vs. SCZ cINs with or without co-culture with activated microglia one week after the removal of activated microglia co-culture. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed Unpaired t-test was used for analysis of KLF5 qPCR for control conditions and Two-tailed Unpaired t-test after log transformation was used for analysis of KLF5 qPCR for those with activated microglia co-culture (n=10 lines for HC and n=10 lines for SCZ; Each data point is averaged from 3 independent differentiations, Control: t=0.1021, df=18, Act.MG: t=1.435, df=18). d, Analysis of oxidative phosphorylation in HC cINs vs. SCZ cINs cultured with or without activated microglia-conditioned media one week after removal of activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test after log transformation was used for analysis of Maximum Respiration for control conditions and Two-tailed Unpaired t-test was used for analysis of Maximum Respiration for those cultured in activated microglia-conditioned media (n=7 lines for HC and n=7 lines for SCZ; Each data point is averaged from 1-4 independent differentiations, Maximum Respiration of Control: t=4.646, df=12, Maximum Respiration of Act.MG: t=3.256, df=12). e, Arborization analysis of cINs infected with a limiting titer of GFP-expressing lentivirus and cultured without activated microglia-conditioned media, with activated microglia-conditioned media or with activated microglia-conditioned media + ALA/ALC treatment. Data are presented as mean ± SEM. One-way ANOVA, followed by posthoc analysis using Dunnett’s multiple comparisons test was used for analysis (n=8 lines consisting of 4 HC lines and 4 SCZ lines, Neurite Length: f=6.116, df=2, Branch Number: f=6.465, df=2).
Extended Data Fig. 8 Gene expression analysis of cINs cultured with or without activated HMC3-conditioned media.
a, qPCR analysis of inflammatory gene expression in cINs with and without culturing in activated microglia HMC3-conditioned media. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed paired t-test was used for analysis of NF-κBIZ and TNFRSF12A and Two-tailed paired t-test after log transformation was used for analysis of TNFAIP3 (n=7 lines, Each data point is averaged from 3 independent differentiations, NF-κBIZ: t=4.406, df=6, TNFRSF12A: t=3.351, df=6, TNFAIP3: t=7.189, df=6). b, qPCR analysis of CTGF and THBS1 expression in HC cINs or SCZ cINs cultured with or without activated microglia HMC3-conditioned media. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed Ratio paired t-test was used for analysis (n=5 lines for HC and n=5 lines for SCZ; Each data point is averaged from 3 independent differentiations, CTGF of HC: t=4.018, df=4, THBS1 of HC: t=3.027, df=4, CTGF of SCZ: t=4.131, df=4, THBS1 of SCZ: t=3.638, df=4). c, qPCR analysis of KLF5 expression in HC cINs or SCZ cINs cultured with or without activated microglia HMC3-conditioned media one week after the removal of activated microglia conditioned media. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed paired t-test after log transformation was used for analysis of HC KLF5 qPCR and Two-tailed paired t-test was used for analysis of SCZ KLF5 qPCR (n=4 lines from HC and n=4 lines from SCZ; Each data point is averaged from 3 independent differentiations, HC: t=0.6992, df=3, SCZ: t=4.364, df=3).
Extended Data Fig. 9 Metabolic analysis of cINs with or without activated HMC3-conditioned media.
a, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in HC cINs cultured with or without activated microglia HMC3-conditioned media using a Seahorse Analyzer. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n=6 lines, ATP Production: t=3.028, df=5, Baseline Glycolysis: t=2.064, df=5, Glycolytic Reserve: t=0.02005, df=5). b, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in SCZ cINs cultured with or without activated microglia HMC3-conditioned media using a Seahorse Analyzer. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n=7 lines, ATP Production: t=4.421, df=6, Baseline Glycolysis: t=0.8454, df=6, Glycolytic Reserve: t=0.8159, df=6). c, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in HC cINs using a Seahorse Analyzer one week after removal of activated microglia HMC3-conditioned media. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis of ATP Production and Baseline Glycolysis and Two-tailed paired t-test after log transformation was used for analysis of Glycolytic Reserve (n=6 lines; Each data point is averaged from 1-4 independent differentiations, ATP Production: t=0.05286, df=5, Baseline Glycolysis: t=1.294, df=5, Glycolytic Reserve: t=0.1876, df=5). d, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in SCZ cINs using a Seahorse Analyzer one week after removal of activated microglia HMC3-conditioned media. Data are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n=7 lines; Each data point is averaged from 1-2 independent differentiations, ATP Production: t=4.051, df=6, Baseline Glycolysis: t=2.008, df=6, Glycolytic Reserve: t=0.7266, df=6). e, Analysis of oxidative phosphorylation in HC cINs vs. SCZ cINs cultured with or without activated microglia HMC3-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test was used for analysis (n=6 lines, Basal Respiration of Control: t=3.743, df=10, Maximum Respiration of Control: t=2.646, df=10, Basal Respiration of Act.MG: t=5.004, df=10, Maximum Respiration of Act.MG: t=3.842, df=10). f, Analysis of oxidative phosphorylation in HC cINs vs. SCZ cINs culture with or without activated microglia HMC3-conditioned media one week after removal of the activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test was used for analysis of both Basal Respiration and Maximum Respiration for control conditions. Two-tailed Unpaired t-test after log transformation was used for analysis of Basal Respiration for those cultured in activated microglia HMC3-conditioned media. Two-tailed Unpaired t-test was used for analysis of Maximum Respiration for those cultured in activated microglia HMC3-conditioned media (n=6 lines; Each data point is averaged from 1-4 independent differentiations, Basal Respiration of Control: t=2.295, df=10, Maximum Respiration of Control: t=2.409, df=10, Basal Respiration of Act.MG: t=4.066, df=10, Maximum Respiration of Act.MG: t=7.071, df=10). g, Analysis of action potential firing-dependent synaptic GABA release in HC cINs vs. SCZ cINs cultured with or without activated microglia HMC3-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test was used for analysis (n=6 lines; Each data point is averaged from 1-3 independent differentiations, Control: t=4.038, df=10, Act.MG: t=5.253, df=10). h, Analysis of action potential firing-dependent synaptic GABA release in HC cINs vs. SCZ cINs cultured with or without activated microglia HMC3-conditioned media one week after removal of activated microglia-conditioned media. Data are presented as mean±SEM. Two-tailed Unpaired t-test after log transformation was used for control conditions and Two-tailed Unpaired t-test was used for those cultured in activated microglia HMC3-conditioned media (n=6 lines; Each data point is averaged from 1-3 independent differentiations, Control: t=2.379, df=10, Act.MG: t=3923, df=10).
Extended Data Fig. 10 Metabolic analysis of developmental glutamatergic neurons cultured in activated microglia-conditioned media.
a, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in HC glutamatergic neurons using a Seahorse Analyzer. Data are presented as mean±SEM. Two-Level Hierarchical Linear Mixed Effect Model was used for analysis (n=4 independent differentiations from 2 lines, each line with 2 independent differentiations). b, Analysis of oxidative phosphorylation (ATP Production) and glycolysis (Baseline Glycolysis and Glycolytic Reserve) in SCZ glutamatergic neurons using a Seahorse Analyzer. Data are presented as mean±SEM. Two-Level Hierarchical Linear Mixed Effect Model after log transformation was used for analysis of ATP Production and Glycolytic Reserve and Two-Level Hierarchical Linear Mixed Effect Model was used for analysis of Baseline Glycolysis (n=4 independent differentiations from 2 lines, each line with 2 independent differentiations). c, Analysis of oxidative phosphorylation in HC glutamatergic neurons vs. SCZ glutamatergic neurons cultured with or without activated microglia HMC3-conditioned media. Data are presented as mean±SEM. One-Level Hierarchical Linear Mixed Effect Model was used for analysis (n=4 independent differentiations from 2 lines, each line with 2 independent differentiations).
Supplementary information
Supplementary Information
Supplementary Tables 1–13.
Rights and permissions
About this article
Cite this article
Park, GH., Noh, H., Shao, Z. et al. Activated microglia cause metabolic disruptions in developmental cortical interneurons that persist in interneurons from individuals with schizophrenia. Nat Neurosci 23, 1352–1364 (2020). https://doi.org/10.1038/s41593-020-00724-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-00724-1
This article is cited by
-
Systemic inflammatory biomarkers in Schizophrenia are changed by ECT administration and related to the treatment efficacy
BMC Psychiatry (2024)
-
Mast cell stabilizer, an anti-allergic drug, reduces ventricular arrhythmia risk via modulation of neuroimmune interaction
Basic Research in Cardiology (2024)
-
Noteworthy perspectives on microglia in neuropsychiatric disorders
Journal of Neuroinflammation (2023)
-
Regulation of synaptic connectivity in schizophrenia spectrum by mutual neuron-microglia interaction
Communications Biology (2023)
-
Current advancements of modelling schizophrenia using patient-derived induced pluripotent stem cells
Acta Neuropathologica Communications (2022)