Abstract
Recent reports have revealed that oligodendrocyte precursor cells (OPCs) are heterogeneous. It remains unclear whether such heterogeneity reflects different subtypes of cells with distinct functions or instead reflects transiently acquired states of cells with the same function. By integrating lineage formation of individual OPC clones, single-cell transcriptomics, calcium imaging and neural activity manipulation, we show that OPCs in the zebrafish spinal cord can be divided into two functionally distinct groups. One subgroup forms elaborate networks of processes and exhibits a high degree of calcium signaling, but infrequently differentiates despite contact with permissive axons. Instead, these OPCs divide in an activity- and calcium-dependent manner to produce another subgroup, with higher process motility and less calcium signaling and that readily differentiates. Our data show that OPC subgroups are functionally diverse in their response to neurons and that activity regulates the proliferation of a subset of OPCs that is distinct from the cells that generate differentiated oligodendrocytes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw sequence data, gene expression data and cell type annotation tables have been deposited in the Gene Expression Omnibus under accession number GSE132166. A web resource is available at https://ki.se/en/mbb/oligointernode. The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Bergles, D. E. & Richardson, W. D. Oligodendrocyte development and plasticity. Cold Spring Harb. Perspect. Biol. 8, a020453 (2015).
Hughes, E. G., Orthmann-Murphy, J. L., Langseth, A. J. & Bergles, D. E. Myelin remodeling through experience-dependent oligodendrogenesis in the adult somatosensory cortex. Nat. Neurosci. 21, 696–706 (2018).
Gibson, E. M. et al. Neuronal activity promotes oligodendrogenesis and adaptive myelination in the mammalian brain. Science 344, 1252304 (2014).
McKenzie, I. A. et al. Motor skill learning requires active central myelination. Science 346, 318–322 (2014).
Zawadzka, M. et al. CNS-resident glial progenitor/stem cells produce Schwann cells as well as oligodendrocytes during repair of CNS demyelination. Cell Stem Cell 6, 578–590 (2010).
Emery, B. & Lu, Q. R. Transcriptional and epigenetic regulation of oligodendrocyte development and myelination in the central nervous system. Cold Spring Harb. Perspect. Biol. 7, a020461 (2015).
Liu, J., Moyon, S., Hernandez, M. & Casaccia, P. Epigenetic control of oligodendrocyte development: adding new players to old keepers. Curr. Opin. Neurobiol. 39, 133–138 (2016).
Foerster, S., Hill, M. F. E. & Franklin, R. J. M. Diversity in the oligodendrocyte lineage: plasticity or heterogeneity? Glia 67, 1797–1805 (2019).
Viganò, F. & Dimou, L. The heterogenic nature of NG2-glia. Brain Res. 1638, 129–137 (2016).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Emery, B. & Barres, B. A. Unlocking CNS cell type heterogeneity. Cell 135, 596–598 (2008).
Marques, S. et al. Transcriptional convergence of oligodendrocyte lineage progenitors during development. Dev. Cell 46, 504–517 (2018).
Richardson, W. D., Young, K. M., Tripathi, R. B. & McKenzie, I. NG2-glia as multipotent neural stem cells: fact or fantasy? Neuron 70, 661–673 (2011).
Hill, R. A., Patel, K. D., Medved, J., Reiss, A. M. & Nishiyama, A. NG2 cells in white matter but not gray matter proliferate in response to PDGF. J. Neurosci. 33, 14558–14566 (2013).
Viganò, F., Möbius, W., Götz, M. & Dimou, L. Transplantation reveals regional differences in oligodendrocyte differentiation in the adult brain. Nat. Neurosci. 16, 1370–1372 (2013).
Falcão, A. M. et al. Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat. Med. 24, 1837–1844 (2018).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Spitzer, S. O. et al. Oligodendrocyte progenitor cells become regionally diverse and heterogeneous with age. Neuron 101, 459–471 (2019).
Sakry, D. et al. Oligodendrocyte precursor cells modulate the neuronal network by activity-dependent ectodomain cleavage of glial NG2. PLoS Biol. 12, e1001993 (2014).
Bergles, D. E., Roberts, J. D., Somogyi, P. & Jahr, C. E. Glutamatergic synapses on oligodendrocyte precursor cells in the hippocampus. Nature 405, 187–191 (2000).
De Biase, L. M., Nishiyama, A. & Bergles, D. E. Excitability and synaptic communication within the oligodendrocyte lineage. J. Neurosci. 30, 3600–3611 (2010).
Kukley, M., Capetillo-Zarate, E. & Dietrich, D. Vesicular glutamate release from axons in white matter. Nat. Neurosci. 10, 311–320 (2007).
Orduz, D. et al. Interneurons and oligodendrocyte progenitors form a structured synaptic network in the developing neocortex. eLife 4, e06953 (2015).
Káradóttir, R., Cavelier, P., Bergersen, L. H. & Attwell, D. NMDA receptors are expressed in oligodendrocytes and activated in ischaemia. Nature 438, 1162–1166 (2005).
Chittajallu, R., Aguirre, A. & Gallo, V. NG2-positive cells in the mouse white and grey matter display distinct physiological properties. J. Physiol. 561, 109–122 (2004).
Barres, B. A. & Raff, M. C. Proliferation of oligodendrocyte precursor cells depends on electrical activity in axons. Nature 361, 258–260 (1993).
Makinodan, M., Rosen, K. M., Ito, S. & Corfas, G. A critical period for social experience-dependent oligodendrocyte maturation and myelination. Science 337, 1357–1360 (2012).
Xiao, L. et al. Rapid production of new oligodendrocytes is required in the earliest stages of motor-skill learning. Nat. Neurosci. 19, 1210–1217 (2016).
Schebesta, M. & Serluca, F. C. olig1 expression identifies developing oligodendrocytes in zebrafish and requires hedgehog and notch signaling. Dev. Dyn. 238, 887–898 (2009).
Auer, F., Vagionitis, S. & Czopka, T. Evidence for myelin sheath remodeling in the CNS revealed by in vivo imaging. Curr. Biol. 28, 549–559 (2018).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Chen, Y. et al. The oligodendrocyte-specific G protein-coupled receptor GPR17 is a cell-intrinsic timer of myelination. Nat. Neurosci. 12, 1398–1406 (2009).
Viganò, F. et al. GPR17 expressing NG2-glia: oligodendrocyte progenitors serving as a reserve pool after injury. Glia 64, 287–299 (2015).
Emery, B. et al. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell 138, 172–185 (2009).
Early, J. J. et al. An automated high-resolution in vivo screen in zebrafish to identify chemical regulators of myelination. eLife 7, e35136 (2018).
Kirischuk, S., Scherer, J., Möller, T., Verkhratsky, A. & Kettenmann, H. Subcellular heterogeneity of voltage-gated Ca2+ channels in cells of the oligodendrocyte lineage. Glia 13, 1–12 (1995).
Pende, M., Holtzclaw, L. A., Curtis, J. L., Russell, J. T. & Gallo, V. Glutamate regulates intracellular calcium and gene expression in oligodendrocyte progenitors through the activation of DL-alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptors. Proc. Natl Acad. Sci. USA 91, 3215–3219 (1994).
Krasnow, A. M., Ford, M. C., Valdivia, L. E., Wilson, S. W. & Attwell, D. Regulation of developing myelin sheath elongation by oligodendrocyte calcium transients in vivo. Nat. Neurosci. 93, 1–28 (2017).
Yu, X. et al. Reducing astrocyte calcium signaling in vivo alters striatal microcircuits and causes repetitive behavior. Neuron 99, 1170–1187 (2018).
Dimou, L. & Simons, M. Diversity of oligodendrocytes and their progenitors. Curr. Opin. Neurobiol. 47, 73–79 (2017).
Simons, M. & Lyons, D. A. Axonal selection and myelin sheath generation in the central nervous system. Curr. Opin. Cell Biol. 25, 512–519 (2013).
Tsai, E. & Casaccia, P. Mechano-modulation of nuclear events regulating oligodendrocyte progenitor gene expression. Glia 67, 1229–1239 (2019).
Zhu, X. et al. Age-dependent fate and lineage restriction of single NG2 cells. Development 138, 745–753 (2011).
Hill, R. A., Patel, K. D., Goncalves, C. M., Grutzendler, J. & Nishiyama, A. Modulation of oligodendrocyte generation during a critical temporal window after NG2 cell division. Nat. Neurosci. 17, 1518–1527 (2014).
Bergles, D. E., Jabs, R. & Steinhäuser, C. Neuron-glia synapses in the brain. Brain Res. Rev. 63, 130–137 (2010).
Kukley, M., Nishiyama, A. & Dietrich, D. The fate of synaptic input to NG2 glial cells: neurons specifically downregulate transmitter release onto differentiating oligodendroglial cells. J. Neurosci. 30, 8320–8331 (2010).
Wake, H. et al. Nonsynaptic junctions on myelinating glia promote preferential myelination of electrically active axons. Nat. Commun. 6, 7844 (2015).
Karttunen, M. J., Czopka, T., Goedhart, M., Early, J. J. & Lyons, D. A. Regeneration of myelin sheaths of normal length and thickness in the zebrafish CNS correlates with growth of axons in caliber. PLoS ONE 12, e0178058 (2017).
Almeida, R. G., Czopka, T., ffrench-Constant, C. & Lyons, D. A. Individual axons regulate the myelinating potential of single oligodendrocytes in vivo. Development 138, 4443–4450 (2011).
Freeman, J. et al. Mapping brain activity at scale with cluster computing. Nat. Methods 11, 941–950 (2014).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Meyer, M. P. & Smith, S. J. Evidence from in vivo imaging that synaptogenesis guides the growth and branching of axonal arbors by two distinct mechanisms. J. Neurosci. 26, 3604–3614 (2006).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53–56 (2017).
Almeida, R. G. & Lyons, D. A. Intersectional gene expression in zebrafish using the split KalTA4 system. Zebrafish 12, 377–386 (2015).
Kwan, K. M. et al. The Tol2kit: a multisite gateway-based construction kit for Tol2 transposon transgenesis constructs. Dev. Dyn. 236, 3088–3099 (2007).
Walton, E. M., Cronan, M. R., Beerman, R. W. & Tobin, D. M. The macrophage-specific promoter mfap4 allows live, long-term analysis of macrophage behavior during mycobacterial infection in zebrafish. PLoS ONE 10, e0138949 (2015).
Mensch, S. et al. Synaptic vesicle release regulates myelin sheath number of individual oligodendrocytes in vivo. Nat. Neurosci. 18, 628–630 (2015).
Czopka, T., ffrench-Constant, C. & Lyons, D. A. Individual oligodendrocytes have only a few hours in which to generate new myelin sheaths in vivo. Dev. Cell 25, 599–609 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Meijering, E. et al. Design and validation of a tool for neurite tracing and analysis in fluorescence microscopy images. Cytometry A 58, 167–176 (2004).
Hines, J. H., Ravanelli, A. M., Schwindt, R., Scott, E. K. & Appel, B. Neuronal activity biases axon selection for myelination in vivo. Nat. Neurosci. 18, 683–689 (2015).
Kirby, B. B. et al. In vivo time-lapse imaging shows dynamic oligodendrocyte progenitor behavior during zebrafish development. Nat. Neurosci. 9, 1506–1511 (2006).
Baraban, M., Koudelka, S. & Lyons, D. A. Ca2+ activity signatures of myelin sheath formation and growth in vivo. Nat. Neurosci. 19, 1–23 (2017).
Acknowledgements
We are grateful to R. Almeida, D. Lyons, T. Misgeld, M. Simons and all members of the Czopka laboratory for their comments and suggestions during the assembly of the data and their presentation in this manuscript. We thank B. Khakh (Department of Physiology, University of California, Los Angeles) for providing the CalEx plasmid before publication, K. Kwan (Department of Human Genetics, University of Utah) for providing the pME_nls-Cerulean and pME_nls-mApple plasmids, and D. Kim (Howard Hughes Medical Institute, Janelia Research Campus) for providing the GCaMP6m plasmid. We thank the Single Cell Genomics Facility, the Science for Life Laboratory, the National Genomics Infrastructure and UPPMAX for providing assistance with massive parallel sequencing and computational infrastructure. The bioinformatics computations were performed on resources provided by the Swedish National Infrastructure for Computing at UPPMAX, Uppsala University. E.A. is funded by the European Union (Horizon 2020 Research and Innovation Programme, Marie Skłodowska-Curie actions, grant SOLO, number 794689). Work in G.C.-B.’s research group was supported by the Swedish Research Council (grant 2015-03558), the European Union (Horizon 2020 Research and Innovation Programme, European Research Council Consolidator Grant EPIScOPE, grant 681893), the Swedish Brain Foundation (FO2017-0075), the Ming Wai Lau Centre for Reparative Medicine and the Karolinska Institutet. Work in T.C.’s research group was funded by a Starting Grant from the European Research Council (ERC StG MecMy, grant 714440), the Deutsche Forschungsgemeinschaft through its Emmy Noether Programme for young investigators (ENP CZ226/1-1) and the Deutsche Forschungsgemeinschaft Excellence Strategy within the framework of the Munich Cluster for Systems Neurology (EXC 2145 SyNergy–D 390857198).
Author information
Authors and Affiliations
Contributions
T.H., R.M., L.J.H., W.B., G.C.-B. and T.C. designed the experiments. T.H., R.M., L.J.H., W.B. and F.A. conducted the experiments. T.H., R.M., L.J.H. and T.C. analyzed the imaging data. E.A. and G.C.-B. performed the bioinformatic analysis of RNA sequencing data. T.C. conceived the project and wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of OPCs in the zebrafish spinal cord.
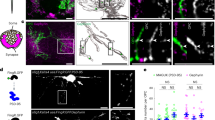
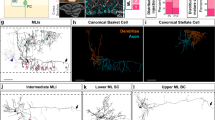
a, Confocal image of a Tg(olig1:memEYFP),Tg(olig1:nls-mApple) zebrafish at the level of the spinal cord at 21 d.p.f. (example of three animals from one experiment). Scale bar, 50 µm. b, Cross-sectional view of the spinal cord showing the distribution of myelin in Tg(mbp:EGFP-CAAX) at 7 d.p.f. (example of 12 animals from four experiments). Scale bar, 10 µm. c, Cross-sectional view of the spinal cord showing the distribution of pre- and postsynapses (Tg(elavl3:synaptophysin-RFP), anti-mCherry, anti-gephyrin) at 7 d.p.f. (example of 12 animals from four experiments). Scale bar, 10 µm. d, Confocal images of Tg(mbp:nls-EGFP),Tg(olig1:nls-mApple) transgenic animals between 4 and 28 d.p.f. (n values as in e). Scale bar, 20 µm. e, Cell numbers of OPCs (olig1: nls-mApple-positive, mbp:nls-EGFP-negative) and myelinating oligodendrocytes (mbp:nls-EGFP-positive) in the spinal cord. Data are expressed as mean cells per field ± s.d. at 3 (n = 17 animals in two experiments), 5 (n = 15 animals in three experiments), 7 (n = 15 in three experiments), 10 (n = 16 in three experiments), 13 (n = 17 in two experiments), 16 (n = 17 in two experiments), 20 (n = 20 in three experiments), 24 (n = 12 in three experiments) and 28 (n = 13 in three experiments) d.p.f. f, Example images of individual OPCs showing a range of morphologies. The soma can be localized within axo-dendritic (top) or neuron-rich areas (middle, bottom). The process network of an individual cell can be restricted to one side of the spinal cord (top and middle cells), but it can also reach to both sides of the spinal cord (bottom cell) (n values as in g). Scale bar, 10 µm. g, OPC morphometry using three-dimensional process tracing and creation of a volume hull around the reconstructed filaments (n = 228 cells from 56 animals between 3 and 16 d.p.f. in 24 experiments). Scale bar, 10 µm.
Extended Data Fig. 2 Analysis of single-cell RNA sequencing clusters.
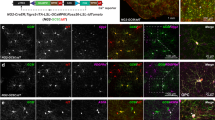
a, Schematic overview of cell isolation, sorting and sequencing. b, Flow cytometry plots of olig1:memEYFP-sorted cells and wild-type control cells. Dotted lines indicate the gating used (example from two independent experiments). c, t-SNE plot showing expression of sox10 (total sample size n = 310 cells). Immunohistochemistry for sox10 on transverse spinal cord sections of 7 d.p.f. Tg(olig1:nls-mApple),Tg(mbp:nls-EGFP) animals and quantification of sox10-expressing OPCs (olig1:nls-mApple-positive, mbp:nls-EGFP-negative) in neuron-rich and axo-dendritic areas (100% (68/68) versus 100% (49/49) positive cells, n = 16 animals in four experiments). Dotted lines indicate the outlines of the spinal cord. Scale bar,10 µm. d, t-SNE plot showing expression of olig2 and nkx2.2a (sample size as in c). e, t-SNE plot showing expression of ppp1r14bb, mbpa and plp1a (sample size as in c). f, Confocal images with in situ hybridizations for cspg4, gpr17, myrf, and labeling of EDU incorporated cells on transverse spinal cord sections of 7 d.p.f. Tg(olig1:nls-mApple),Tg(mbp:nls-EGFP) animals (see Fig. 2e,i,k,l for respective n values). Scale bar, 10 µm.
Extended Data Fig. 3 Quantification of OPC morphology and position before differentiation.
Quantification of OPC complexity and soma position from imaging timelines between 3 and 15 d.p.f. Measured is the last timepoint as OPC before differentiation, as assessed by myelin sheath formation (imaging intervals of 1 d between 3 and 7 d.p.f., and 2 d between 7 and 15 d.p.f.). n = 10, n = 6, n = 3, n = 3, n = 3, n = 2, n = 1, n = 2, n = 2, n = 2, n = 1, n = 5 and n = 1 cells at 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 and 15 d.p.f. Data from 23 animals in six experiments.
Extended Data Fig. 4 Time-lapse imaging of OPC population dynamics.
a, Overview images of transgenic zebrafish labeling nuclei of OPCs (olig1:nls-mApple) and myelinating oligodendrocytes (mbp:nls-EGFP) at the beginning and end of a timelapse between 3 and 5 d.p.f. Dashed boxes indicate the areas shown in panel c (n = 3 animals in two experiments). Scale bar, 10 µm. b, Quantification of the fates of OPCs found in neuron-rich areas. A detailed breakdown of the data shown in Fig. 4e. c, Zoom-ins and false coloring of the time-lapse in a, showing potential behaviors of OPCs in neuron-rich areas: remaining quiescent (red cell), generating new OPCs in neuron-rich areas (magenta cells) or generating new OPCs in axo-dendritic areas (green cells). The insets at the first and last timepoints show the absence of myelin markers (mbp:nls-EGFP) in the cells studied (n = 3 animals in two experiments). Scale bar, 10 µm.
Extended Data Fig. 5 Cell fate analysis of OPCs with their soma in neuron-rich areas.
a, Time series of an individual OPC with its soma in neuron-rich areas that gives rise to myelinating oligodendrocytes by proliferation-mediated generation of daughter OPCs in axo-dendritic areas. Left panel, confocal images. Middle panel, reconstructions of the starting cell and the individual daughter cells. Cells that will differentiate are shown in blue. Right panels, y-axis rotations showing olig1:nls-mApple cell body positions within the hemi-spinal cord. Dashed lines depict the outline of the spinal cord. One of eight examples from seven animals in six experiments. Scale bar, 10 µm. b, Graphical summary of cell fates from the data analyzed in Fig. 5a–d. See also Supplementary Fig. 2.
Extended Data Fig. 6 Characterization of OPC GCaMP reporter lines.
a, Example images of individual olig1:GCaMP-CAAX-labeled OPCs in axo-dendritic areas of the zebrafish spinal cord at 4 d.p.f. The absence of nascent ensheathments indicates that these cells are not early differentiating oligodendrocytes (n = 9 independent experiments). Scale bar, 10 µm. b, Dorsal views of Tg(olig1:GCaMP6m),Tg(mbp:KillerRed) transgenic zebrafish at 4 d.p.f. to label OPCs and differentiated oligodendrocytes. Dotted box indicates position of zoom-ins in bottom row (n = 3 animals in one experiment). Scale bars, 50 µm (top) and 20 µm (bottom). c, Quantification of single- and double-positive cells from images as shown in b. d, ∆F/F0 GCaMP transients of individual cells in two Tg(olig1:GCaMP6m) zebrafish. Green traces depict cells in axo-dendritic areas, and gray traces depict cells in neuron-rich areas (total of eight animals in eight experiments).
Extended Data Fig. 7 Effects of chronic 4-AP incubation on zebrafish.
a, Minimum intensity projections of a 2 min time-lapse of fish freely swimming in a 3 cm petri dish in different treatment conditions (n = 6, n = 7, n = 3 and n = 3 animals in control, 4-AP, TTX and 4-AP+TTX conditions, three independent experiments). b, Traces of GCaMP transients from Tg(elavl3:h2b-GCaMP6s) zebrafish at 4 d.p.f. and after overnight incubation in 0.1 mM 4-AP and before and after 10 µM TTX (seven animals per condition in two experiments). c, Confocal images of Tg(mfap4:memCerulean),Tg(olig1:nls-mApple) zebrafish at 4 d.p.f. after treatment with 0.1 mM 4-AP, 0.5 mM 4-AP, or Danieau’s solution as a control. Transmitted light images show spinal cord morphology and tissue integrity after drug treatment. Scale bars, 100 µm. The graph shows the number of macrophages that accumulate in a 400 µM length of spinal cord of Tg(mfap4:memCerulean) zebrafish after 1 d of control (2 ± 0.25/2 cells), 0.1 mM (2 ± 1/2 cells) and 0.5 mM (3 ± 0.25/2 cells) 4-AP treatment (median ± 25%/75% percentiles). P = 0.43 (control versus 0.1 mM 4-AP), P = 0.03 (control versus 0.5 mM 4-AP), Kruskal–Wallis test, test statistic=3.003, n = 16, n = 19 and n = 8 animals in three experiments. d, Representative images of Tg(mbp:nls-EGFP),Tg(olig1:nls-mApple) zebrafish in control treatment and after 2 d of 0.1 mM 4-AP treatment (see Fig. 7e for n values). Scale bar, 20 µm.
Supplementary information
Supplementary Information
Supplementary Figures 1 and 2, Supplementary Tables 1–3.
Supplementary Video 1
Pan and zoom of OPC reporter lines. Transgenic olig1:memEYFP, olig1:nls-Cerulean zebrafish at 5 d.p.f. showing the distribution of OPCs and their process network within the CNS. Representative images from four animals in two independent experiments.
Supplementary Video 2
Segmentation of individual OPCs. Animation of process tracing and hull construction to reconstruct single OPCs at 4 d.p.f. in the spinal cord (example of n = 76 cells from 23 animals in 11 experiments).
Supplementary Video 3
Time-lapse of process remodeling of an OPC within axo-dendritic areas. 60 min time-lapse of individual OPC within axo-dendritic areas of the spinal cord recorded at 5 min intervals (n = 4 cells from four animals in four experiments). See also Supplementary Video 4.
Supplementary Video 4
Time-projection of process remodeling of an OPC within axo-dendritic areas. Projection of a 60 min time-lapse of an individual OPC within axo-dendritic areas of the spinal cord. Stable processes are green, and remodeled processes appear in magenta (n = 4 cells from four animals in four experiments).
Supplementary Video 5
Time-lapse of process remodeling of an OPC with its soma in neuron-rich areas. 60 min time-lapse of an individual OPC with its soma in neuron-rich areas of the spinal cord recorded at 5 min intervals (n = 4 cells from four animals in four experiments). See also Supplementary Video 6.
Supplementary Video 6
Time-projection of process remodeling of an OPC with its soma in neuron-rich areas. Projection of a 60 min time-lapse of an individual OPC with its soma in neuron-rich areas of the spinal cord. Stable processes are green, and remodeled processes appear in magenta (n = 4 cells from four animals in four experiments).
Supplementary Video 7
Behavior and fate of individual OPC with its soma in neuron-rich areas. 24 h time-lapse showing a cell division of an individual OPC with high process complexity typically found when the OPC soma resides in neuron-rich areas (n = 13 cells from four animals in three experiments).
Supplementary Video 8
Behavior and fate of an individual OPC within axo-dendritic areas. 13 h time-lapse showing differentiation of an individual OPC with low process complexity typically found residing in axo-dendritic areas (n = 21 cells from nine animals in seven experiments).
Supplementary Video 9
Transition of OPC soma between neuron-rich and axo-dendritic areas is linked to cell divisions. 42 h timelapse of olig1:nls-mApple-labeled OPCs. The rounded nucleus shaded in green represents an OPC residing in neuron-rich areas. It divides to give rise to a daughter cell in axo-dendritic areas (elliptic nucleus), and the daughter cell continues to divide. Representative results from three animals in two independent experiments.
Supplementary Video 10
OPC calcium transients can be restricted to process subdomains. Time-lapse of two olig1-GCaMP6m-CAAX-labeled OPCs showing transients in process subdomains. Example of 27 animals in 23 independent experiments.
Supplementary Video 11
OPC calcium transients can spread throughout the cell. Time-lapse of two olig1-GCaMP6m-CAAX-labeled OPCs where one cell shows transients throughout the cell. Example of 27 animals in 23 independent experiments.
Supplementary Video 12
Population analysis of OPC calcium transients. Time-lapse of a Tg(olig1:GCaMP6m) at the level of the spinal cord (dorsal view) showing a GCaMP transient in the soma of a single OPC within the population. Example of eight animals in eight independent experiments.
Rights and permissions
About this article
Cite this article
Marisca, R., Hoche, T., Agirre, E. et al. Functionally distinct subgroups of oligodendrocyte precursor cells integrate neural activity and execute myelin formation. Nat Neurosci 23, 363–374 (2020). https://doi.org/10.1038/s41593-019-0581-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0581-2
This article is cited by
-
Microglia regulation of central nervous system myelin health and regeneration
Nature Reviews Immunology (2024)
-
Synaptic input and Ca2+ activity in zebrafish oligodendrocyte precursor cells contribute to myelin sheath formation
Nature Neuroscience (2024)
-
A preliminary study of the effects of an antimuscarinic agent on anxious behaviors and white matter microarchitecture in nonhuman primates
Neuropsychopharmacology (2024)
-
Reconstruction of macroglia and adult neurogenesis evolution through cross-species single-cell transcriptomic analyses
Nature Communications (2024)
-
Oligodendrocyte calcium signaling promotes actin-dependent myelin sheath extension
Nature Communications (2024)