Abstract
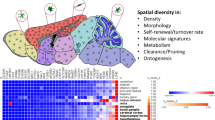
Microglia are tissue-resident macrophages of the CNS that orchestrate local immune responses and contribute to several neurological and psychiatric diseases. Little is known about human microglia and how they orchestrate their highly plastic, context-specific adaptive responses during pathology. Here we combined two high-dimensional technologies, single-cell RNA-sequencing and time-of-flight mass cytometry, to identify microglia states in the human brain during homeostasis and disease. This approach enabled us to identify and characterize a previously unappreciated spectrum of transcriptional states in human microglia. These transcriptional states are determined by their spatial distribution, and they further change with aging and brain tumor pathology. This description of multiple microglia phenotypes in the human CNS may open promising new avenues for subset-specific therapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data for this project are available at the Gene Expression Omnibus under accession code GSE135437. The data can be explored in a browser widget at https://single-cell.shinyapps.io/human-microglia/.
Code availability
Computer code for this project can be provided upon request and found at https://github.com/rsankowski/sankowski-et-al-microglia.
Change history
30 May 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41593-022-01089-3
References
Ransohoff, R. M. & Cardona, A. E. The myeloid cells of the central nervous system parenchyma. Nature 468, 253–262 (2010).
Prinz, M., Priller, J., Sisodia, S. S. & Ransohoff, R. M. Heterogeneity of CNS myeloid cells and their roles in neurodegeneration. Nat. Neurosci. 14, 1227–1235 (2011).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Prinz, M., Erny, D. & Hagemeyer, N. Ontogeny and homeostasis of CNS myeloid cells. Nat. Immunol. 18, 385–392 (2017).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Takahashi, K., Rochford, C. D. & Neumann, H. Clearance of apoptotic neurons without inflammation by microglial triggering receptor expressed on myeloid cells-2. J. Exp. Med. 201, 647–657 (2005).
Jordao, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Hickman, S. E. et al. The microglial sensome revealed by direct RNA sequencing. Nat. Neurosci. 16, 1896–1905 (2013).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Galatro, T. F. et al. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat. Neurosci. 20, 1162–1171 (2017).
Ajami, B. et al. Single-cell mass cytometry reveals distinct populations of brain myeloid cells in mouse neuroinflammation and neurodegeneration models. Nat. Neurosci. 21, 541–551 (2018).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395.e6 (2018).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Li, Q. et al. Developmental heterogeneity of microglia and brain myeloid cells revealed by deep single-cell rna sequencing. Neuron 101, 207–223.e10 (2019).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Mathys, H. et al. Temporal tracking of microglia activation in neurodegeneration at single-cell resolution. Cell Rep. 21, 366–380 (2017).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Böttcher, C. et al. Human microglia regional heterogeneity and phenotypes determined by multiplexed single-cell mass cytometry. Nat. Neurosci. 22, 78–90 (2019).
Darmanis, S. et al. Single-cell RNA-seq analysis of infiltrating neoplastic cells at the migrating front of human glioblastoma. Cell Rep. 21, 1399–1410 (2017).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Venteicher, A. S. et al. Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355, eaai8478 (2017).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Jakel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Herman, J. S., Sagar & Grun, D. FateID infers cell fate bias in multipotent progenitors from single-cell RNA-seq data. Nat. Methods 15, 379–386 (2018).
Grun, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Grun, D. et al. De novo prediction of stem cell identity using single-cell transcriptome data. Cell Stem Cell 19, 266–277 (2016).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Butovsky, O. et al. Identification of a unique TGF-beta-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Grabert, K. et al. Microglial brain region-dependent diversity and selective regional sensitivities to aging. Nat. Neurosci. 19, 504–516 (2016).
Hagemeyer, N. et al. Microglia contribute to normal myelinogenesis and to oligodendrocyte progenitor maintenance during adulthood. Acta Neuropathol. 134, 441–458 (2017).
Wlodarczyk, A. et al. A novel microglial subset plays a key role in myelinogenesis in developing brain. EMBO J. 36, 3292–3308 (2017).
Chabas, D. et al. The influence of the proinflammatory cytokine, osteopontin, on autoimmune demyelinating disease. Science 294, 1731–1735 (2001).
Hambardzumyan, D., Gutmann, D. H. & Kettenmann, H. The role of microglia and macrophages in glioma maintenance and progression. Nat. Neurosci. 19, 20–27 (2016).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Safaiyan, S. et al. Age-related myelin degradation burdens the clearance function of microglia during aging. Nat. Neurosci. 19, 995–998 (2016).
Paloneva, J. et al. Mutations in two genes encoding different subunits of a receptor signaling complex result in an identical disease phenotype. Am. J. Hum. Genet. 71, 656–662 (2002).
Rademakers, R. et al. Mutations in the colony stimulating factor 1 receptor (CSF1R) gene cause hereditary diffuse leukoencephalopathy with spheroids. Nat. Genet. 44, 200–205 (2011).
Wu, Y. E., Pan, L., Zuo, Y., Li, X. & Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron 96, 313–329.e6 (2017).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-seq. Genome Biol. 17, 77 (2016).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Baker, S. C. et al. The external RNA controls consortium: a progress report. Nat. Methods 2, 731–734 (2005).
Grun, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Allaire, J. & Chollet, F. keras: R Interface to ‘Keras’. R package version 2.2.4.1.9001 (2017).
Abadi, M. et al. Tensorflow: large-scale machine learning on heterogeneous distributed systems. Preprint at arXiv https://arxiv.org/abs/1603.04467 (2016).
Ma, F. & Pellegrini, M. Automated identification of cell types in single cell RNA sequencing. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/532093v1 (2019).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
The authors thank J. Bodinek-Wersing, T. El Gaz, E. Barleon and A. Frömming for excellent technical support. R.S. is supported by the Berta-Ottenstein Programme for Clinician Scientists. M.P. is supported by the Sobek Foundation, the Ernst-Jung Foundation, the German Research Foundation (SFB 992, SFB1160, Reinhart-Koselleck-Grant) and the Ministry of Science, Research and Arts, Baden-Wuerttemberg (Sonderlinie “Neuroinflammation”) and by the BMBF-funded competence network of multiple sclerosis (KKNMS). This study was supported by the German Research Foundation (DFG) under Germany’s Excellence Strategy (CIBSS, EXC-2189, project ID 390939984). The authors would also like to acknowledge the assistance of the BCRT Flow Cytometry Lab (Charité—Universitätsmedizin Berlin, Germany). C.B., M.P. and J.P. are supported by the German Research Foundation (SFB/TRR167 “NeuroMac”). J.P. received additional funding from the Berlin Institute of Health (CRG2aSP6) and the UK DRI (Momentum Award). D.G. was supported by the Max Planck Society, the German Research Foundation (DFG) (SPP1937 GR4980/1-1, GR4980/3-1 and GRK2344 MeInBio), by the DFG under Germany’s Excellence Strategy (CIBSS, EXC-2189, project ID 390939984), by the ERC (818846, ImmuNiche, ERC-2018-COG) and by the Behrens-Weise-Foundation.
Author information
Authors and Affiliations
Contributions
R.S., C.B., T.M., L.G. and E.S. conducted the experiments and analyzed the data. C.S., M.J.S., D.H.H. and O.S. performed brain surgeries for tissue acquisition. R.S. and S. analyzed the scRNA-seq data (under the supervision of D.G.). R.S., C.B., T.S., A.M., D.G., J.P. and M.P. supervised the project and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.M. and T.S. are shareholders of AC Immune. All the other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised. In the version of this article initially published, Fig.3b,c, Fig. 5a–d, Extended Data Fig. 1 and Supplementary Fig. 8b contained errors. Fig. 3b and c showed incorrect immunohistochemistry quantifications. Fig. 5a contained a duplicate image for the TMEM119 control sample, and panels b and c showed incorrect immunohistochemistry quantifications. Fig. 5d contained an incorrect image for the t-SNE map of P2RY12; it was a duplicate of the map for HLA-DRA. Extended Data Fig. 1 contained incorrect images in panel a for Pat 7 (TMEM119), Pat 12 (TMEM119, CD74) and Pat 15 (CD68) and in panel b for Pat 3 (GFAP), Pat 12 (TMEM119 and P2YR12) and Pat 13 (TMEM119). Supplementary Fig. 8b contained an incorrect image for the CSF2 t-SNE panel. In addition, the legend for Fig. 5b as well as the description of Fig. 1b in Methods incorrectly said that the test used was a paired t-test; it should read ‘unpaired t-test’ in both places. In the Results section, the sentence “Normalization for the number of IBA1+ cells revealed lower expression of the bona fide microglial markers P2RY12 and TMEM119, and higher expression of HLA-DR (Fig. 5c)” should read “no change of expression” instead of “lower expression”.
Extended data
Extended Data Fig. 1 Representative immunohistochemical pictures of control samples.
(a) grey and (b) white matter from each subject included in the study. Each image is representative of n=10 independent images taken per specimen.
Extended Data Fig. 2 Pregating Strategy and FACS plots – control samples.
Overview of FACS plots for the control samples included in the study. The boxplot whiskers represent 1.5 times the interquartile range. Each sample represents an independent experiment.
Extended Data Fig. 3 Cell signatures and microglia cell-to-cell-distance heatmap – control samples.
a) Cell signatures of microglial and non-microglial cells. Aggregate transcript counts of genes enriched in the given cell type (indicated in parentheses) are color-coded on t-SNE maps. b) Unsupervised clustering of microglia after exclusion of non-microglial cells divided microglia in 9 major populations, that is clusters. The heatmap displays pairwise cell-to-cell distances with clusters color-coded on the bottom. C4 (grey color) was removed from subsequent analysis due to upregulation of cell-stress genes indicating dissociation-induced artefacts. Minor clusters containing individual cells that comprise less than 1 % of all cells were also removed (white ‘x’ on black background), leading to n = 4,116 microglia that passed quality control. The color scale represents the pairwise distance between cells.
Extended Data Fig. 4 t-SNE maps and line plots of selected genes - control samples.
t-SNE maps color-coded for transcript counts with the corresponding line plots.
Extended Data Fig. 5 Dataset comparison with published data.
a) Workflow for the neural network classifier. b) Heatmap depicting the comparison between the true (y-axis) and the predicted (x-axis) cluster assignments on the test data. Microglia that were assigned correctly are lying on the diagonal. The color-coding indicates the ratio between the number of microglia in a given cell divided by the sum of all cells in this row. Absolute numbers are provided in each cell.
Extended Data Fig. 6 Line plots of selected GO terms - control samples.
Aggregate transcript counts of genes enriched in the respective GO term are visualized using line plots.
Extended Data Fig. 7 SPP1 immunohistochemistry and clusterwise SPP1 expression across age bins - control samples.
a) Immunohistochemistry of SPP1 and Iba1 in control grey and white matter. Each image is representative of n=5 independent images taken per specimen. b) Violin and box plots showing the transcript counts of SPP1 across age bins and. Statistically significant p-values are indicated resulting from two-way analysis of variance followed by a Tukey post-hoc test. Violin plots represent kernel density estimates of expression values from n=4,396 cells from 15 samples. The boxplot spans the 25th to 75th percentiles and the whiskers represent 1.5 times the interquartile range. The bold line indicates the median.
Extended Data Fig. 8 FACS plots of GBM samples. Overview of FACS plots for the GBM samples included in the study.
Each row represents a separate experiment.
Extended Data Fig. 9 Cell signatures and cell-to-cell distance heatmap – GBM samples.
a) Cell signatures of microglial and non-microglial cells. Aggregate transcript counts of genes enriched in the given cell type (indicated in parentheses) are color-coded on the respective t-SNE maps. b) Unsupervised clustering of microglia after exclusion of non-microglial cells resulted in 14 major populations, that is clusters. The heatmap displays pairwise cell-to-cell distances. The color scale represents the pairwise distance between cells.
Extended Data Fig. 10 t-SNE maps and line plots of selected genes - GBM samples.
t-SNE maps color-coded for transcript counts with the corresponding line plots.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Supplementary Tables
Excel sheet with all Supplementary Tables 1–12.
Rights and permissions
About this article
Cite this article
Sankowski, R., Böttcher, C., Masuda, T. et al. Mapping microglia states in the human brain through the integration of high-dimensional techniques. Nat Neurosci 22, 2098–2110 (2019). https://doi.org/10.1038/s41593-019-0532-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0532-y
This article is cited by
-
Regulation of human microglial gene expression and function via RNAase-H active antisense oligonucleotides in vivo in Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
The roles of tissue resident macrophages in health and cancer
Experimental Hematology & Oncology (2024)
-
Transcriptional characterization of iPSC-derived microglia as a model for therapeutic development in neurodegeneration
Scientific Reports (2024)
-
SorLA restricts TNFα release from microglia to shape a glioma-supportive brain microenvironment
EMBO Reports (2024)
-
Harnessing innate immune pathways for therapeutic advancement in cancer
Signal Transduction and Targeted Therapy (2024)