Abstract
The thalamus is the central communication hub of the forebrain and provides the cerebral cortex with inputs from sensory organs, subcortical systems and the cortex itself. Multiple thalamic regions send convergent information to each cortical region, but the organizational logic of thalamic projections has remained elusive. Through comprehensive transcriptional analyses of retrogradely labeled thalamic neurons in adult mice, we identify three major profiles of thalamic pathways. These profiles exist along a continuum that is repeated across all major projection systems, such as those for vision, motor control and cognition. The largest component of gene expression variation in the mouse thalamus is topographically organized, with features conserved in humans. Transcriptional differences between these thalamic neuronal identities are tied to cellular features that are critical for function, such as axonal morphology and membrane properties. Molecular profiling therefore reveals covariation in the properties of thalamic pathways serving all major input modalities and output targets, thus establishing a molecular framework for understanding the thalamus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The single-neuron reconstruction data are available at http://ml-neuronbrowser.janelia.org. Raw and processed RNA-seq data have been deposited into the Gene Expression Omnibus repository (GSE133911 for pooled-cell thalamic nuclei-level RNA-seq data and GSE133912 for the single-cell RNA-seq data).
Code availability
The gene expression data can be browsed and plotted and differential gene expression tests can be performed interactively at http://thalamoseq.janelia.org. Scripts for data analysis are available on github (https://github.com/aschulmann/ThalamoSeq).
Change history
01 October 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Sherman, S. M. & Guillery, R. W. The role of the thalamus in the flow of information to the cortex. Philos. Trans. R. Soc. B Biol. Sci. 357, 1695–1708 (2002).
Jones, E. G. The Thalamus (Cambridge Univ. Press, 2007).
Nakajima, M. & Halassa, M. M. Thalamic control of functional cortical connectivity. Curr. Opin. Neurobiol. 44, 127–131 (2017).
Clascá, F., Rubio-Garrido, P. & Jabaudon, D. Unveiling the diversity of thalamocortical neuron subtypes. Eur. J. Neurosci. 35, 1524–1532 (2012).
Willis, T. Cerebri anatome, cui accessit nervorum descriptio et usus. (Londini, 1664).
Jasper, H. Diffuse projection systems: the integrative action of the thalamic reticular system. Electroencephalogr. Clin. Neurophysiol. 1, 405–419 (1949); discussion 1, 419–420 (1949).
Smith, Y. et al. The thalamostriatal system in normal and diseased states. Front. Syst. Neurosci. 8, 5 (2014).
Jones, E. G. & Hendry, S. H. C. Differential calcium binding protein immunoreactivity distinguishes classes of relay neurons in monkey thalamic nuclei. Eur. J. Neurosci. 1, 222–246 (1989).
Jones, E. G. Viewpoint: the core and matrix of thalamic organization. Neuroscience 85, 331–345 (1998).
Jones, E. G. The thalamic matrix and thalamocortical synchrony. Trends Neurosci. 24, 595–601 (2001).
Cowan, W. M. & Powell, T. P. A study of thalamo-striate relations in the monkey. Brain 79, 364–390 (1956).
Kato, S. et al. Action selection and flexible switching controlled by the intralaminar thalamic neurons. Cell Rep. 22, 2370–2382 (2018).
Nelson, S. B., Sugino, K. & Hempel, C. M. The problem of neuronal cell types: a physiological genomics approach. Trends Neurosci. 29, 339–345 (2006).
Kepecs, A. & Fishell, G. Interneuron cell types are fit to function. Nature 505, 318–326 (2014).
Murray, K. D., Choudary, P. V. & Jones, E. G. Nucleus- and cell-specific gene expression in monkey thalamus. Proc. Natl Acad. Sci. USA 104, 1989–1994 (2007).
Frangeul, L. et al. A cross-modal genetic framework for the development and plasticity of sensory pathways. Nature 538, 96–98 (2016).
Nagalski, A. et al. Molecular anatomy of the thalamic complex and the underlying transcription factors. Brain Struct. Funct. 221, 2493–2510 (2016).
Franklin, K. B. J. & Paxinos, G. The Mouse Brain in Stereotaxic Coordinates. 4th edn (Elsevier, 2013).
Sugino, K. et al. Mapping the transcriptional diversity of genetically and anatomically defined cell populations in the mouse brain. eLife 8, e38619 (2019).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Freeman, S. A., Desmazières, A., Fricker, D., Lubetzki, C. & Sol-Foulon, N. Mechanisms of sodium channel clustering and its influence on axonal impulse conduction. Cell. Mol. Life Sci. 73, 723–735 (2016).
Heinemann, S. H., Rettig, J., Wunder, F. & Pongs, O. Molecular and functional characterization of a rat brain Kv β3 potassium channel subunit. FEBS Lett. 377, 383–389 (1995).
Rudy, B. & McBain, C. J. Kv3 channels: voltage-gated K+ channels designed for high-frequency repetitive firing. Trends Neurosci. 24, 517–526 (2001).
Okaty, B. W., Miller, M. N., Sugino, K., Hempel, C. M. & Nelson, S. B. Transcriptional and electrophysiological maturation of neocortical fast-spiking GABAergic interneurons. J. Neurosci. 29, 7040–7052 (2009).
Nakamura, K. C., Sharott, A. & Magill, P. J. Temporal coupling with cortex distinguishes spontaneous neuronal activities in identified basal ganglia-recipient and cerebellar-recipient zones of the motor thalamus. Cereb. Cortex 24, 81–97 (2014).
Puil, E., Meiri, H. & Yarom, Y. Resonant behavior and frequency preferences of thalamic neurons. J. Neurophysiol. 71, 575–582 (1994).
Fogerson, P. M. & Huguenard, J. R. Tapping the brakes: cellular and synaptic mechanisms that regulate thalamic oscillations. Neuron 92, 687–704 (2016).
Economo, M. N. et al. A platform for brain-wide imaging and reconstruction of individual neurons. eLife 5, e10566 (2016).
Bickford, M. E., Zhou, N., Krahe, T. E., Govindaiah, G. & Guido, W. Retinal and tectal ‘driver-like’ inputs converge in the shell of the mouse dorsal lateral geniculate nucleus. J. Neurosci. 35, 10523–10534 (2015).
Kuramoto, E. et al. Two types of thalamocortical projections from the motor thalamic nuclei of the rat: a single neuron-tracing study using viral vectors. Cereb. Cortex 19, 2065–2077 (2009).
Groh, A. et al. Convergence of cortical and sensory driver inputs on single thalamocortical cells. Cereb. Cortex 24, 3167–3179 (2014).
Lu, E., Llano, D. A. & Sherman, S. M. Different distributions of calbindin and calretinin immunostaining across the medial and dorsal divisions of the mouse medial geniculate body. Hear. Res. 257, 16–23 (2009).
Mo, C., Petrof, I., Viaene, A. N. & Sherman, S. M. Synaptic properties of the lemniscal and paralemniscal pathways to the mouse somatosensory thalamus. Proc. Natl Acad. Sci. USA 114, E6212–E6221 (2017).
Ramcharan, E. J., Gnadt, J. W. & Sherman, S. M. Higher-order thalamic relays burst more than first-order relays. Proc. Natl Acad. Sci. USA 102, 12236–12241 (2005).
Ferster, D., Chung, S. & Wheat, H. Orientation selectivity of thalamic input to simple cells of cat visual cortex. Nature 380, 249–252 (1996).
Van der Werf, Y. D., Witter, M. P. & Groenewegen, H. J. The intralaminar and midline nuclei of the thalamus. Anatomical and functional evidence for participation in processes of arousal and awareness. Brain Res. Brain Res. Rev. 39, 107–140 (2002).
Saalmann, Y. B. Intralaminar and medial thalamic influence on cortical synchrony, information transmission and cognition. Front. Syst. Neurosci. 8, 83 (2014).
Rikhye, R. V., Wimmer, R. D. & Halassa, M. M. Toward an integrative theory of thalamic function. Annu. Rev. Neurosci. 41, 163–183 (2018).
Guo, K., Yamawaki, N., Svoboda, K. & Shepherd, G. M. G. Anterolateral motor cortex connects with a medial subdivision of ventromedial thalamus through cell type-specific circuits, forming an excitatory thalamo-cortico-thalamic loop via layer 1 apical tuft dendrites of layer 5B pyramidal tract type neurons. J. Neurosci. 38, 8787–8797 (2018).
Bennett, C. et al. Higher-order thalamic circuits channel parallel streams of visual information in mice. Neuron 102, 477–492.e5 (2019).
Mandelbaum, G. et al. Distinct cortical-thalamic-striatal circuits through the parafascicular nucleus. Neuron https://doi.org/10.1016/j.neuron.2019.02.035 (2019).
Suzuki-Hirano, A. et al. Dynamic spatiotemporal gene expression in embryonic mouse thalamus. J. Comp. Neurol. 519, 528–543 (2011).
Wong, S. Z. H. et al. In vivo clonal analysis reveals spatiotemporal regulation of thalamic nucleogenesis. PLoS Biol. 16, e2005211 (2018).
Altman, J. & Bayer, S. A. Development of the diencephalon in the rat. V. Thymidine-radiographic observations on internuclear and intranuclear gradients in the thalamus. J. Comp. Neurol. 188, 473–499 (1979).
Angevine, J. B. Time of neuron origin in the diencephalon of the mouse. An autoradiographic study. J. Comp. Neurol. 139, 129–187 (1970).
Shi, W. et al. Ontogenetic establishment of order-specific nuclear organization in the mammalian thalamus. Nat. Neurosci. 20, 516–528 (2017).
Grant, E., Hoerder-Suabedissen, A. & Molnár, Z. The regulation of corticofugal fiber targeting by retinal inputs. Cereb. Cortex 26, 1336–1348 (2016).
Antón-Bolaños, N., Espinosa, A. & López-Bendito, G. Developmental interactions between thalamus and cortex: a true love reciprocal story. Curr. Opin. Neurobiol. 52, 33–41 (2018).
Cembrowski, M. S. & Menon, V. Continuous variation within cell types of the nervous system. Trends Neurosci. 41, 337–348 (2018).
Kuramoto, E. et al. Complementary distribution of glutamatergic cerebellar and GABAergic basal ganglia afferents to the rat motor thalamic nuclei. Eur. J. Neurosci. 33, 95–109 (2011).
Tervo, D. G. R. et al. A Designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Kita, T., Shigematsu, N. & Kita, H. Intralaminar and tectal projections to the subthalamus in the rat. Eur. J. Neurosci. 44, 2899–2908 (2016).
Mandelbaum, G. et al. Distinct cortical–thalamic–striatal circuits through the parafascicular nucleus. Preprint at https://doi.org/10.1101/370734 (2018).
Hempel, C. M., Sugino, K. & Nelson, S. B. A manual method for the purification of fluorescently labeled neurons from the mammalian brain. Nat. Protoc. 2, 2924–2929 (2007).
Cembrowski, M. S. et al. Spatial gene-expression gradients underlie prominent heterogeneity of CA1 pyramidal neurons. Neuron 89, 351–368 (2016).
Cembrowski, M. S. et al. Dissociable structural and functional hippocampal outputs via distinct subiculum cell classes. Cell 173, 1280–1292.e18 (2018).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Soumillon, M., Cacchiarelli, D., Semrau, S., Oudenaarden, A. van & Mikkelsen, T. S. Characterization of directed differentiation by high-throughput single-cell RNA-seq. Preprint at biorXiv https://doi.org/10.1101/003236 (2014).
Murphy, S. D. et al. The Janelia workstation for neuroscience. Keystone Big Data Biol. San Fr. CA (2014).
Johnson, H., Harris, G. & Williams, K. BRAINSFit: mutual information rigid registrations of whole-brain 3D images, using the insight toolkit. Insight J. http://hdl.handle.net/1926/1291 (2007).
Ng, L. et al. An anatomic gene expression atlas of the adult mouse brain. Nat. Neurosci. 12, 356–362 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Cembrowski, M. S., Wang, L., Sugino, K., Shields, B. C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
Miller, M. N., Okaty, B. W. & Nelson, S. B. Region-specific spike-frequency acceleration in layer 5 pyramidal neurons mediated by Kv1 subunits. J. Neurosci. 28, 13716–13726 (2008).
Miller, M. N., Okaty, B. W., Kato, S. & Nelson, S. B. Activity-dependent changes in the firing properties of neocortical fast-spiking interneurons in the absence of large changes in gene expression. Dev. Neurobiol. 71, 62–70 (2011).
Dani, V. S. et al. Reduced cortical activity due to a shift in the balance between excitation and inhibition in a mouse model of Rett syndrome. Proc. Natl Acad. Sci. USA 102, 12560–12565 (2005).
Acknowledgements
The authors thank Karel Svoboda, Albert Lee and Amy Chuong for critical input throughout the project. They also thank Matthew Phillips, Mark Cembrowski, Andre Marques-Smith, Virginia Rutten and Yves Weissenberger for comments on the manuscript. Thanks are also given to the following individuals: Kshama Aswath and Jingqun Ma for technical assistance with library preparation and RNA-seq; Monique Copeland and Amy Hu for help with FISH and imaging; Vilas Menon, Damian Kao and Mark Cembrowski for help with single-cell RNA-seq analysis; the MouseLight annotators for single neuron reconstructions; Kim Ritola and the Janelia Viral Tools and the Anatomy and Histology facilities for production of viruses and histology; Jody Clements for website engineering; and Daniel Morozoff, Yajie Liang, Justin Little, Ondrej Zelenka, Amy Chuong and Na Ji for surgical protocols and assistance in identifying nuclei for dissection. Finally, they also thank the Janelia Vivarium for animal care and surgeries. This project was funded as a small project team (ThalamoSeq) by the HHMI at the Janelia Research Campus, following a pilot project in the Dudman/Hantman labs. S.B.N., V.V. and C.L. were also supported by grants from the NINDS (NS109916) and the NIMH (MH105949). A.S. is funded via the Janelia Visiting Scientists Program.
Author information
Authors and Affiliations
Contributions
J.W.P. contributed to all aspects of this project. A.S. analyzed and collected data, planned the project and wrote the paper. E.H. planned the project and collected data. J.W. collected and analyzed the single-cell reconstruction data. C.L. and V.V. collected and analyzed the electrophysiology data. L.W. and B.C.S. collected data. W.K. supervised the project. J.C. supervised the single-cell reconstruction project. A.L.L. collected data and developed methods. B.M. edited the paper. J.T.D. supervised the project and edited the paper. S.B.N. supervised the project and edited the paper. A.W.H. initiated and supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Pooled-cell RNAseq quality control and additional analyses.
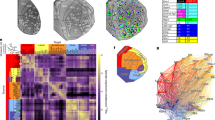
a. Markers of non-neuronal sample contamination are low across our dataset. Expression (TPM) in pooled-cell samples shown for 8 genes marking astrocytes, microglia, oligodendrocytes and erythrocytes. Only a small number of samples showed expression of contamination markers. b. ERCC spike-in level correlation with their mean TPM (left), and spike-in detection (TPM>0) across samples (right) indicate high accuracy and sensitivity of the pooled-cell RNAseq measurements (n=120). Blue lines show linear and logistic function fits, respectively. Shaded areas are 95% confidence intervals. c. Heatmap of the top 500 differentially expressed genes. Mean expression per nucleus is shown (22 nuclei; n=120 samples). Rows and columns are ordered by hierarchical clustering with Euclidean distance metric. Colors represent gene-wise Z-scores. d. Samples of the same nucleus obtained via different labelling methods cluster similarly. Principal component analysis of those samples, for which multiple collection methods were used (that is GENSAT lines in addition to retrograde labeling; n=46 for projection labeling and n=31 for genetic labeling), using the top 500 genes with highest variance. Samples are colored by collection approach or transgenic line used.
Supplementary Figure 2 Modality and core/matrix dichotomy do not account for the major split of thalamic gene expression profiles.
a. Hierarchical clustering tree from Fig. 1, colored by modality (left), core/matrix (middle) and this paper’s nomenclature (right). b. Expression of and Pvalb, Calb1, and Calb2 (mean ± standard error of the mean; each dot represents a replicate; n=120 samples), with nuclei on the x-axis colored by their profile from Fig. 1. The core/matrix organizational theory proposes that thalamus is divided into two discrete groups, expressing Calb1 or Pvalb. In three unsupervised clustering approaches (hclust: hierarchical clustering; kmeans: k-means clustering with k=2; mclust: two-component Gaussian mixture model) the main split consistently separated tertiary and secondary profiles, both of which are marked by Calb1 and would thus both be considered ‘matrix’ nuclei in this theory. Thus, core/matrix differences do not reflect the main split in thalamic organizational structure. Calb2 exhibits heterogeneity even within a profile.
Supplementary Figure 3 Dimensionality reduction of thalamic gene expression data.
a. Scree plot of the PCA from Fig. 2 (22 nuclei, n=120 samples) shows the variance explained by the top 10 PCs compared to a shuffled matrix with the order of samples permuted for each gene (mean and standard deviation from 1000 permutations). The variance explained by the first 6 PCs is substantially larger than PCs of shuffled data. b. Dot plot illustrating separation of the three major profiles of thalamic projection neuron (primary, secondary, tertiary) along the first principal component (PC1). Thalamic nuclei are ordered by their mean position on PC1. Dots represent samples. c. Multidimensional scaling using an alternative distance metric also identifies a similar leading axis of variance with classical relay nuclei on one end and midline/intralaminar nuclei on the other end. Distance was defined as the quadratic mean of the log2 fold changes of the top 500 differentially expressed genes between any two samples (meaning that the gene set used for the distance comparison varies between each sample pair).
Supplementary Figure 4 Separation of thalamic nuclei by profiles and projection in the first six principal components.
Clustering of profiles (a, as defined by hierarchical clustering in Fig. 1c) and projections (b, as defined by cortical target area, in which the retrograde tracer was injected) on PC1-6 from Fig. 2a. Dots represent single samples (total n=90). Boxes show median and quartiles, and whiskers extend up to the highest and lowest value within 1.5x interquartile range of the upper and lower quartiles. Individual samples are shown as black dots. Asterisks denote significance of each group in a two-sided Wilcoxon rank-sum test compared to all samples taken together (*: p<0.05, **: p<0.01, ***: p<0.001, ****: p<0.0001; for exact p-values see Source Data 2). Specific striatal projection types were not examined here because of the coarseness of our striatal viral injections.
Supplementary Figure 5 Additional analysis of profile- and projection-specific gene expression differences.
a. Classification accuracy for distinguishing primary (n=28), secondary (n=50), and tertiary (n=30) type nuclei, as well as for distinguishing motor (VL,VA,VM; n=20) vs. sensory (LGd,LP,VB,PO; n=25) and visual (LGd,LP; n=14) vs. somatosensory (VB,PO; n=11) nuclei samples. Classifiers were obtained using elastic-net logistic regression on 20 random genes over 100 iterations. To prevent bias due to sample size difference, larger groups were subsampled to the size of the smallest group (n=11) at each iteration. Accuracy was assessed using 5-fold cross-validation. Boxes show median and quartiles, and whiskers extend up to the highest and lowest value within 1.5x interquartile range of the upper and lower quartiles. b. Genes that best distinguish motor from sensory nuclei samples (LGd,VB,LP,PO vs. VL,VA,VM; n=25 and n=20, respectively). Plotted are the top 20 genes with false discovery rate < 10-3 (likelihood ratio test), fold change > 2, and ordered by highest signal-to-noise ratio (mean log fold change between vs. within group).
Supplementary Figure 6 Topographical arrangement of gene expression across thalamic nuclei.
Relationship of PC1-6 with topographical position of nuclei. Rostrocaudal, dorsoventral, and mediolateral positions are the x, y, and z voxel coordinates, respectively, in the Allen Mouse Brain Reference Atlas. 1 voxel = 1 µm. All correlations are determined via linear regression and p-values are calculated via two-sided Student’s t-test.
Supplementary Figure 7 Expression of ion channels and receptors across thalamic nuclear profiles.
Expression of voltage-gated ion channels (left) and neurotransmitter/neuromodulator receptors (right) across thalamic nuclei. The mean expression is shown for each nucleus (22 nuclei, n=120 samples). Genes are plotted separately for each group and ordered by their PC1 loading within each group. Colors are gene-wise Z-scores. 5-HT=serotonin; DA=dopamine; NE=norepinephrine; HA=histamine.
Supplementary Figure 8 Additional electrophysiological properties vary between thalamic nuclear profiles.
a. Additional analysis of action potential waveform features. All statistical tests and experimental details are the same as in Fig. 3c, d. b. Analysis of mEPSCs across motor cortex-projecting nuclei. Statistics as in Fig. 3d. Left: example mEPSC traces. Middle: mean trace for mEPSC from each nucleus. Right: decay time constant, frequency, and amplitude in each nucleus. The number of recorded neurons were 11 for VL, 15 for VA and 12 for CM. Asterisks denote significance level in a Tukey HSD test (*: p<0.05, **: p<0.01; for exact p-values see Source Data 2). c. As in Supplementary Fig. 8b, but for mIPSCs. The number of recorded neurons were 13 for VL, 9 for VA and 11 for CM. d. Cumulative frequency plots of the mEPSC (left) and mIPSC (right) amplitudes.
Supplementary Figure 9 Variation in axonal projections of reconstructed thalamic neurons.
a. 3D visualization of four reconstructed thalamic neurons (total n=106). Colors represent the calculated PC1 score based on each neuron’s gene expression score. Dashed line represents the coronal position shown in b. b. Coronal view of the neurons shown in Supplementary Fig. 9a projected onto the normalized cortical depth map (see Methods). Variations in cortical depth innervation could hereby be reliably measured independently of cortical area. c. Histogram of the distribution of middle layer proportion scores. d. Histogram of measured axonal length within the caudoputamen for all neurons. e. Relationship between a neuron’s gene expression PC1 score and its axon density in cortex. f. Histogram of axonal density in cortex for all reconstructed thalamic neurons. g. The gene expression PC1 score correlated significantly with a neuron’s mediolateral position in the thalamus. Correlations are determined via linear regression and p-values are calculated via two-sided Student’s t-test.
Supplementary Figure 10 Quality control for single-cell RNAseq data.
a. Unique molecular identifier (UMI) count (upper) and gene detection rate (lower) for all collected single cells. Cutoffs for downstream use are indicated by dashed lines. b. PCA on the single-cell RNAseq data revealed that principal component 3 represented contamination with oligodendrocytic transcripts (top 30 genes with the highest absolute loadings for the top 100 cells with the highest scores on PC3 are shown; left). Cells with PC3 position <0.05 were removed (right). c. ERCC spike-in level correlation with their mean UMI count (left), and spike-in detection (UMI>0) across cells (right) indicate high accuracy and sensitivity of the single-cell RNAseq measurements. Blue lines show linear and logistic function fits, respectively. Shaded areas are 95% confidence intervals. d. PC1 loadings for the most differentially expressed genes between nuclei (gene set as in Fig. 2) are highly correlated in pooled-cell and single-cell RNAseq data. Correlations are determined via linear regression and p-values are calculated via two-sided Student’s t-test.
Supplementary Figure 12 Intermediate cell identities in the single-cell RNAseq data.
a. Transitions between single-cell clusters are relatively continuous. tSNE plots of single cells for each projection system as in Fig. 6a (total n=1,952). Color reflects cluster identity as in Fig. 6a. The alpha value (transparency of the color fill) of each cell indicates the class probability for its respective cluster using a random forest classifier (see Methods). b. Expression in counts per million (CPM) of three single-cell marker genes (Tnnt1, Necab1, Calb2) across all 1,952 single thalamic neurons shows intermediate cells expressing more than one marker gene. Cells are colored by their position on pooled-cell PC1.
Supplementary Figure 13 Multi-FISH show cells with mixed expression of cluster marker genes.
Expanded views of example regions from Fig. 7 showing intermediate cells expressing combinations of Tnnt1, Necab1, and Calb2, which are preferentially expressed in primary, secondary and tertiary nuclear profiles respectively. The inset images for mediodorsal, reuniens/rhomboid/SMT (submedial thalamic nucleus) and visual/somatosensory thalamus are the same as in Fig. 7b. Multi-FISH experiments were repeated twice to ensure reproducibility.
Supplementary Figure 14 Quantification of multi-FISH images shows intermediate cells.
Quantification of multi-FISH gene expression images. Regions of interest (ROIs) were drawn in ImageJ. Intensity was normalized first to the ROI size, then divided by the maximum for that channel. Only cells that express at least one of the marker genes were included.
Supplementary Figure 15 Summary: a conserved molecular architecture across thalamic projection systems.
a. Thalamic neurons projecting to a given cortical area are distributed across multiple nuclei (left), in this example, motor thalamus. We applied pooled-cell RNAseq to each nucleus and found three major classes of thalamic nuclei (middle). Single-cell RNAseq and in situ hybridizations revealed that these profiles lie along a continuum with cells having intermediate identities (right). b. The continuum of cell types that existed across nuclei of a projection system could also be found within nuclei boundaries. The prefrontal-projecting mediodorsal nucleus (left) and VL provide examples (right). c. In summary, three major thalamic gene expression profiles are repeated across each projection system and there is continuous cell-type variation between them. Ion channel encoding genes are differentially expressed along the axis, in a manner predictive of action potential waveform (tested in motor thalamus). These gene expression profiles correlate with differing morphological projections, as shown for the motor, visual and somatosensory systems, but morphological features also likely show projection-system specific differences. IL = intralaminar nuclei. * by Reuniens indicates that tertiary status is inferred by expression of tertiary markers, though it contains additional transcriptional differences that make it cluster separately in hierarchical clustering.
Supplementary information
Supplementary Data 1
Pooled-cell RNA-seq metadata and differential gene expression.
Supplementary Data 2
Pooled-cell RNA-seq principal component analysis, Panther protein class enrichment and summary of electrophysiology results.
Supplementary Data 3
Single-cell RNA-seq metadata and cluster marker genes.
Rights and permissions
About this article
Cite this article
Phillips, J.W., Schulmann, A., Hara, E. et al. A repeated molecular architecture across thalamic pathways. Nat Neurosci 22, 1925–1935 (2019). https://doi.org/10.1038/s41593-019-0483-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0483-3
This article is cited by
-
Skeletal interoception in osteoarthritis
Bone Research (2024)
-
Prognostic analysis and risk assessment based on RNA editing in hepatocellular carcinoma
Journal of Applied Genetics (2024)
-
A phylogenetically-conserved axis of thalamocortical connectivity in the human brain
Nature Communications (2023)
-
The impact of the human thalamus on brain-wide information processing
Nature Reviews Neuroscience (2023)
-
Spatial transcriptomics for profiling the tropism of viral vectors in tissues
Nature Biotechnology (2023)