Abstract
Developmental programs that generate the astonishing neuronal diversity of the nervous system are not completely understood and thus present a major challenge for clinical applications of guided cell differentiation strategies. Using direct neuronal programming of embryonic stem cells, we found that two main vertebrate proneural factors, Ascl1 and neurogenin 2 (Neurog2), induce different neuronal fates by binding to largely different sets of genomic sites. Their divergent binding patterns are not determined by the previous chromatin state, but are distinguished by enrichment of specific E-box sequences that reflect the binding preferences of the DNA-binding domains. The divergent Ascl1 and Neurog2 binding patterns result in distinct chromatin accessibility and enhancer activity profiles that differentially shape the binding of downstream transcription factors during neuronal differentiation. This study provides a mechanistic understanding of how transcription factors constrain terminal cell fates, and it delineates the importance of choosing the right proneural factor in neuronal reprogramming strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
Analysis scripts are available at https://github.com/seqcode/Aydin_2019_iAscl1-vs-iNeurog2.
References
Bertrand, N., Castro, D. S. & Guillemot, F. Proneural genes and the specification of neural cell types. Nat. Rev. Neurosci. 3, 517–530 (2002).
Guillemot, F. & Hassan, B. A. Beyond proneural: emerging functions and regulations of proneural proteins. Curr. Opin. Neurobiol. 42, 93–101 (2017).
Urbán, N. & Guillemot, F. Neurogenesis in the embryonic and adult brain: same regulators, different roles. Front. Cell. Neurosci. 8, 396 (2014).
Schuurmans, C. & Guillemot, F. Molecular mechanisms underlying cell fate specification in the developing telencephalon. Curr. Opin. Neurobiol. 12, 26–34 (2002).
Parras, C. M. et al. Divergent functions of the proneural genes Mash1 and Ngn2 in the specification of neuronal subtype identity. Genes Dev. 16, 324–338 (2002).
Osório, J., Mueller, T., Rétaux, S., Vernier, P. & Wullimann, M. F. Phylotypic expression of the bHLH genes Neurogenin2, Neurod, and Mash1 in the mouse embryonic forebrain. J. Comp. Neurol. 518, 851–871 (2010).
Simionato, E. et al. atonal- and achaete-scute-related genes in the annelid Platynereis dumerilii: insights into the evolution of neural basic-helix-loop-helix genes. B Evo.l Biol. 8, 1–13 (2008).
Jarman, A. P. & Ahmed, I. The specificity of proneural genes in determining Drosophila sense organ identity. Mech. Dev. 76, 117–125 (1998).
Fode, C. et al. A role for neural determination genes in specifying the dorsoventral identity of telencephalic neurons. Genes Dev. 14, 67–80 (2000).
Jarman, A. P., Grau, Y., Jan, L. Y. & Jan, Y. N. atonal is a proneural gene that directs chordotonal organ formation in the Drosophila peripheral nervous system. Cell 73, 1307–1321 (1993).
Hirsch, M. R., Tiveron, M. C., Guillemot, F., Brunet, J. F. & Goridis, C. Control of noradrenergic differentiation and Phox2a expression by MASH1 in the central and peripheral nervous system. Development 125, 599–608 (1998).
Lo, L., Dormand, E., Greenwood, A. & Anderson, D. J. Comparison of the generic neuronal differentiation and neuron subtype specification functions of mammalian achaete-scute and atonal homologs in cultured neural progenitor cells. Development 129, 1553–1567 (2002).
Ma, Q., Fode, C., Guillemot, F. & Anderson, D. J. Neurogenin1 and neurogenin2 control two distinct waves of neurogenesis in developing dorsal root ganglia. Genes Dev. 13, 1717–1728 (1999).
Schuurmans, C. et al. Sequential phases of cortical specification involve neurogenin-dependent and -independent pathways. EMBO J. 23, 2892–2902 (2004).
Baker, N. E. & Brown, N. L. All in the family: proneural bHLH genes and neuronal diversity. Development 145, 1–9 (2018).
Flames, N. & Hobert, O. Transcriptional control of the terminal fate of monoaminergic neurons. Annu. Rev. Neurosci. 34, 153–184 (2011).
Tsunemoto, R. et al. Diverse reprogramming codes for neuronal identity. Nature 557, 380 (2018).
Wichterle, H., Gifford, D. & Mazzoni, E. Mapping neuronal diversity one cell at a time. Science 341, 726–727 (2013).
Hobert, O. Regulation of terminal differentiation programs in the nervous system. Annu. Rev. Cell Dev. Biol. 27, 681–696 (2011).
Stefanakis, N., Carrera, I. & Hobert, O. Regulatory logic of pan-neuronal gene expression in C. elegans. Neuron 87, 733–750 (2015).
Heinrich, C. et al. Generation of subtype-specific neurons from postnatal astroglia of the mouse cerebral cortex. Nat. Protoc. 6, 214–228 (2011).
Chanda, S. et al. Generation of induced neuronal cells by the single reprogramming factor ASCL1. Stem Cell Rep. 3, 282–296 (2014).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Mall, M. et al. Myt1l safeguards neuronal identity by actively repressing many non-neuronal fates. Nature 544, 245–249 (2017).
Wapinski, O. L. et al. Hierarchical mechanisms for direct reprogramming of fibroblasts to neurons. Cell 155, 621–635 (2013).
Vadodaria, K. C. et al. Generation of functional human serotonergic neurons from fibroblasts. Mol. Psychiatry 21, 49–61 (2016).
Masserdotti, G. et al. Transcriptional mechanisms of proneural factors and REST in regulating neuronal reprogramming of astrocytes. Cell Stem Cell 17, 74–88 (2015).
Smith, D. K., Yang, J., Liu, M.-L. L. & Zhang, C.-L. L. Small molecules modulate chromatin accessibility to promote NEUROG2-mediated fibroblast-to-neuron reprogramming. Stem Cell Rep. 7, 955–969 (2016).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Raposo, A. A. et al. Ascl1 coordinately regulates gene expression and the chromatin landscape during neurogenesis. Cell Rep. 10, 1544–1556 (2015).
Casey, B. H., Kollipara, R. K., Pozo, K. & Johnson, J. E. Intrinsic DNA binding properties demonstrated for lineage-specifying basic helix-loop-helix transcription factors. Genome Biol. 28, 484–496 (2018).
Slattery, M. et al. Absence of a simple code: how transcription factors read the genome. Trends Biochem. Sci. 39, 381–399 (2014).
Powell, L. M., Zur Lage, P. I., Prentice, D. R., Senthinathan, B. & Jarman, A. P. The proneural proteins Atonal and Scute regulate neural target genes through different E-box binding sites. Mol. Cell. Biol. 24, 9517–9526 (2004).
Castro, D. S. et al. Proneural bHLH and Brn proteins coregulate a neurogenic program through cooperative binding to a conserved DNA motif. Dev. Cell 11, 831–844 (2006).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Gordân, R. et al. Genomic regions flanking E-box binding sites influence DNA binding specificity of bHLH transcription factors through DNA shape. Cell Rep. 3, 1093–1104 (2013).
Rohs, R. et al. Origins of specificity in protein–DNA recognition. Annu. Rev. Biochem. 79, 233–269 (2010).
Massari, M. E. & Murre, C. Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Mol. Cell. Biol. 20, 429–440 (2000).
Ma, P. C., Rould, M. A., Weintraub, H. & Pabo, C. O. Crystal structure of MyoD bHLH domain–DNA complex: perspectives on DNA recognition and implications for transcriptional activation. Cell 77, 451–459 (1994).
Ali, F. R. et al. The phosphorylation status of Ascl1 is a key determinant of neuronal differentiation and maturation in vivo and in vitro. Development 141, 2216–2224 (2014).
Hindley, C. et al. Post-translational modification of Ngn2 differentially affects transcription of distinct targets to regulate the balance between progenitor maintenance and differentiation. Development 139, 1718–1723 (2012).
Quan, X.-J. et al. Post-translational control of the temporal dynamics of transcription factor activity regulates neurogenesis. Cell 164, 460–475 (2016).
Li, S. et al. RAS/ERK signaling controls proneural genetic programs in cortical development and gliomagenesis. J. Neurosci. 34, 2169–2190 (2014).
Li, S. et al. GSK3 temporally regulates neurogenin 2 proneural activity in the neocortex. J. Neurosci. 32, 7791–7805 (2012).
Wapinski, O. L. et al. Rapid chromatin switch in the direct reprogramming of fibroblasts to neurons. Cell Rep. 20, 3236–3247 (2017).
Fong, A. P. et al. Conversion of MyoD to a neurogenic factor: binding site specificity determines lineage. Cell Rep. 10, 1937–1946 (2015).
De Masi, F. et al. Using a structural and logics systems approach to infer bHLH-DNA binding specificity determinants. Nucleic Acids Res. 39, 4553–4563 (2011).
Chien, C. T., Hsiao, C. D., Jan, L. Y. & Jan, Y. N. Neuronal type information encoded in the basic-helix-loop-helix domain of proneural genes. Proc. Natl Acad. Sci. USA 93, 13239–13244 (1996).
Nakada, Y., Hunsaker, T. L., Henke, M. R. & Johnson, J. E. Distinct domains within Mash1 and Math1 are required for function in neuronal differentiation versus neuronal cell-type specification. Development 131, 1319–1330 (2004).
Quan, X.-J. et al. Evolution of neural precursor selection: functional divergence of proneural proteins. Development 131, 1679–1689 (2004).
Iacovino, M. et al. Inducible cassette exchange: a rapid and efficient system enabling conditional gene expression in embryonic stem and primary cells. Stem Cells 29, 1580–1588 (2011).
Mazzoni, E. O. et al. Embryonic stem cell-based mapping of developmental transcriptional programs. Nat. Methods 8, 1056–1058 (2011).
Zappulo, A. et al. RNA localization is a key determinant of neurite-enriched proteome. Nat. Commun. 8, 583 (2017).
Groth, R. D., Lindskog, M., Thiagarajan, T. C., Li, L. & Tsien, R. W. Ca2+/CaM-dependent kinase type II triggers upregulation of GluA1 to coordinate adaptation to synaptic inactivity in hippocampal neurons. Proc. Natl Acad. Sci. USA 108, 828–833 (2011).
Bootman, M. D., Rietdorf, K., Collins, T., Walker, S. & Sanderson, M. Loading fluorescent Ca2+ indicators into living cells. Cold Spring Harb. Protoc. 8, 122–125 (2013).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Kim, D. et al. TopHat2 : accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y. & Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
O’Leary, N. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 2832 (2014).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411 (2018).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, 1–10 (2009).
Mahony, S. et al. An integrated model of multiple-condition ChIP-Seq data reveals predeterminants of Cdx2 binding. PLoS Comput. Biol. 10, e1003501 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Machanick, P. & Bailey, T. L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Kakumanu, A., Velasco, S., Mazzoni, E. O. & Mahony, S. Deconvolving sequence features that discriminate between overlapping regulatory annotations. PLoS Comput. Biol. 13, 1–22 (2017).
Chiu, T. P. et al. DNAshapeR: an R/Bioconductor package for DNA shape prediction and feature encoding. Bioinformatics 32, 1211–1213 (2016).
McLean, C. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, nbt.1630 (2010).
Velasco, S. et al. A multi-step transcriptional and chromatin state cascade underlies motor neuron programming from embryonic stem cells. Cell Stem Cell 20, 205–217.e8 (2017).
Mazzoni, E. O. et al. Synergistic binding of transcription factors to cell-specific enhancers programs motor neuron identity. Nat. Neurosci. 16, 1219–1227 (2013).
Acknowledgements
This work is supported by the NICHD (R01HD079682) and Project ALS (A13-0416) to E.O.M. and by a NYSTEM pre-doctoral training grant (C026880) to B.A. S.M. is supported by the NIGMS (R01GM125722) and the National Science Foundation ABI Innovation grant no. DBI1564466. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation. M.R. is supported by NYU MSTP (T32GM007308) and Developmental Genetics T32 (T32HD007520) grants. N.F. and M.M.-E. are supported by an ERC Starting Grant (ERC STG 2011–281920). The authors would like to thank L. Tejavibulya and A. Ashokkumar for their help with molecular biology, M. Khalfan for his help with scRNA-seq analysis, M. Cammer from the NYU Medical Center Microscopy Core for the ImageJ script used in calcium imaging analysis, and the NYU Genomics Core facility. Finally, the authors would like to thank S. Small, N. Konstantinidis, P. Onal, O. Wapinski, S. Ercan, C. Rushlow, C. Desplan, and Mazzoni Lab members for their helpful suggestions on the manuscript.
Author information
Authors and Affiliations
Contributions
B.A. performed cell differentiations, RNA-seq, ChIP-seq, ATAC-seq, scRNA-seq, immunohistochemistry experiments, and generated the inducible chimera line. Two replicates of iAscl1 48 h RNA-seq experiments were performed by M.M.-E. with guidance from N.F. M.R. performed calcium imaging of neurons with guidance from N.R. G.G provided the Tubb3::GFP line. A.K., B.A., and S.M. performed analysis of all sequencing data. B.A., S.M., and E.O.M. conceived the experiments and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Neuroscience thanks Diogo Castro, Carol Schuurmans, and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Ascl1 and Neurog2 induction programs neuronal fate.
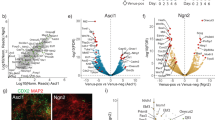
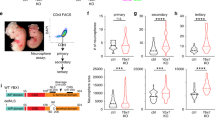
a, Immunocytochemistry of iASCL1 (iA - left panel) and iNEUROG2 (iN - right panel) embryoid bodies (EBs) showing the expression of neuronal Tubb3 and V5-tagged Ascl1 or FLAG-tagged Neurog2 transgene expression over time, respectively (scale bar: 1 µm). 0h time point (before induction of TFs) is used as a control. Similar results were observed in at least n = 2 cell differentiations. b, Schematic representation of time-series (bulk) RNA-seq experiments (not to scale) showing the overlap of the genes up or downregulated early (12 h) or late (48 h) between iA and iN neurons. c, Gene Ontology (GO-terms) biological process terms enriched at the early and late shared upregulated genes between iA and iN neurons (PANTHER- Fisher’s Exact test: FDR-corrected p-value < 0.05) (iA 12 h n = 2; iN 12 h n = 2; iA 48 h n = 5; iN 48 h n = 2 independent cell differentiations). d, Bulk RNA-seq heatmaps showing the expression of subtype-specific markers in iA and iN neurons 48h after induction (EB t = 0 n = 5; iA 48 h n = 5; iN 48 h n = 2 independent cell differentiations). log2(fold change) values are plotted with respect to before induction (EB t = 0) in magenta-orange heatmap. The ratio of iN (48 h) over iA (48 h) is plotted on the blue-green heatmap.
Supplementary Figure 2 Divergent binding of Ascl1 and Neurog2 is not an artifact.
ChIP-seq heatmap with all sites identified in 12 h Ascl1 and Neurog2 datasets shows the divergent binding pattern when 10 k (Fig. 2) or all sites are analyzed (n = 3). b, Ascl1 and Neurog2 bind to largely non-overlapping sites even at 48 h after induction. ChIP-seq heatmap with top 10k sites identified in 48 h Ascl1 and Neurog2 datasets shows that the late binding of Ascl1 and Neurog2 is also divergent (n = 2). c, ChIP-seq heatmap with all sites identified in 48 h Ascl1 and Neurog2 datasets showing that the late divergent binding is still retained when all sites are analyzed (n = 2). d, e, Comparison of Ascl1 ChIP-seq experiments across published datasets in mouse embryonic fibroblasts (MEFs) (d), and ESCs (e). Heatmaps display shared binding sites (that is peaks called in both experiments) and significantly differentially bound sites across experiments. f, ChIP-seq and RNA-seq genome browser snapshots displaying gene expression and differential binding of Ascl1 and Neurog2 at the Dll1 locus. g, Comparison of Ascl1 and Neurog2 binding across 12 h and 48 h time-points. Table displays counts of shared binding sites (that is peaks called in both experiments) and significantly differentially bound sites across experiments.
Supplementary Figure 3 Nucleotides flanking the core E-box motif contribute to the binding specificity of Ascl1 and Neurog2.
a, Nucleotides flanking the core “CAGNTG” k-mer also contribute to Ascl1 or Neurog2 differential binding (A > N or N > A) and shared binding (A = N) in vivo. b, DNA shape prediction at the Ascl1- and Neurog2-preferred sites around the core E-box motifs. Larger predicted propeller twist and minor groove width were noted at alternate sides of the core E-box motif in Ascl1-preferred sites (A > N).
Supplementary Figure 4 The k-mer preference and the expression profile of the A[N]bHLH chimera.
a, The A[N]bHLH chimera binding sites are enriched in Neurog2-preferred core and flanking k-mers. b, Volcano plot comparing the gene expression between iA and iA[N]bHLH at 48 h after induction by RNA-seq. Colored dots represent differentially expressed genes in iA (blue) or iA[N]bHLH(pink) (q-value cut-off < 0.01, Wald test) (iA 48 h n = 5; iA[N]bHLH 48 h n = 2 independent cell differentiations). c, Volcano plot comparing the gene expression between iN and iA[N]bHLH at 48 h after induction by RNA-seq. Colored dots represent differentially expressed genes in iN (green) or iA[N]bHLH(pink) (q-value cut-off < 0.01, Wald test) (iA[N]bHLH 48 h n = 2; iN 48 h n = 2 independent cell differentiations).
Supplementary Figure 5 Global chromatin accessibility dynamics mimic the divergent-to-convergent gene expression dynamics in Ascl1- and Neurog2-induced neurogenesis.
a, Global comparison of sites that differentially gained or lost accessibility at 12 h (top) and 48 h (bottom) iA and iN neurons with respect to EB 0 h. Note the increased shared changes by 48 h (p-val cut-off < 0.05) (n = 2 independent cell differentiations). b, Comparison of differentially accessible regions in iAscl1 EBs (this paper) to iAscl1 in MEFs (Wapinski et al45). c, Time-series ATAC-seq heatmaps showing the dynamics of the accessibility gain at the differentially bound and shared sites of Ascl1 and Neurog2 along with the accessibility dynamics in the iA[N]bHLH chimeric TF line. Note that the A[N]bHLH chimera has increased ATAC-seq read counts at the Neurog2-preferred sites (N > A) both at 12 h and 48 h.
Supplementary Figure 6 Examples of shared targets that have differential Ascl1 or Neurog2 binding.
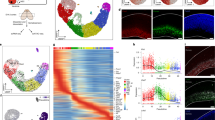
a, Comparison of endogenous Brn2 ChIP-seq in mESC (this paper) with exogenous Brn2 in MEFs programming when expressed exogenously with Ascl1 and Myt1l (iBAM – Wapinski et al25). Heatmaps display shared binding sites (that is peaks called in both experiments) and significantly differentially bound sites across experiments. b, Maximum motif log-likelihood scores of downstream TFs shows weaker enrichment of their respective motifs in iA > iN (left) or iN > iA (right) sites that have E-box motifs as opposed to their top 10 K binding sites (150 bp window around sites) (n = 2 independent cell differentiations) c, tSNE plots showing the cells that express shared targets in iA (top cluster) and iN (bottom cluster) neuron clusters (top panel / n = 1 cell differentiation) that have differential Ascl1 or Neurog2 binding (genome browser shots – bottom panel / n = 3 independent cell differentiations).
Supplementary Figure 7 Working model.
Ascl1 and Neurog2 bind to their preferred sites in the genome mediated intrinsically by their bHLH DNA-binding domain. This divergent binding, in return, results in distinct chromatin landscapes that affect the activity of downstream TFs in establishing generic and neuron-specific phenotypes (TF, transcription factor).
Supplementary information
Supplementary Table 1
Summary of genes vs. peaks associations from GREAT analysis. The peaks are the same sets represented in Fig 2a,b. Gene sets evaluated in Fig. 7 represent genes that are significantly upregulated in both iA and iN compared with EBs (iA = iN), and genes that are significantly differentially expressed between iA and iN (iA > iN and iN > iA) for each relevant timepoint. Overrepresentation table shows the ratio between genes overlapped by real peaks versus genes overlapped by random peaks. P-value tables are calculated using GREAT’s binomial testing tool.
Rights and permissions
About this article
Cite this article
Aydin, B., Kakumanu, A., Rossillo, M. et al. Proneural factors Ascl1 and Neurog2 contribute to neuronal subtype identities by establishing distinct chromatin landscapes. Nat Neurosci 22, 897–908 (2019). https://doi.org/10.1038/s41593-019-0399-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0399-y
This article is cited by
-
RFX4 is an intrinsic factor for neuronal differentiation through induction of proneural genes POU3F2 and NEUROD1
Cellular and Molecular Life Sciences (2024)
-
Promoter hypermethylation of neural-related genes is compatible with stemness in solid cancers
Epigenetics & Chromatin (2023)
-
Efficient generation of functional neurons from mouse embryonic stem cells via neurogenin-2 expression
Nature Protocols (2023)
-
Ascl1 and Ngn2 convert mouse embryonic stem cells to neurons via functionally distinct paths
Nature Communications (2023)
-
Longitudinal scRNA-seq analysis in mouse and human informs optimization of rapid mouse astrocyte differentiation protocols
Nature Neuroscience (2023)