Abstract
Microglia, the specialized innate immune cells of the CNS, play crucial roles in neural development and function. Different phenotypes and functions have been ascribed to rodent microglia, but little is known about human microglia (huMG) heterogeneity. Difficulties in procuring huMG and their susceptibility to cryopreservation damage have limited large-scale studies. Here we applied multiplexed mass cytometry for a comprehensive characterization of postmortem huMG (103 – 104 cells). We determined expression levels of 57 markers on huMG isolated from up to five different brain regions of nine donors. We identified the phenotypic signature of huMG, which was distinct from peripheral myeloid cells but was comparable to fresh huMG. We detected microglia regional heterogeneity using a hybrid workflow combining Cytobank and R/Bioconductor for multidimensional data analysis. Together, these methodologies allowed us to perform high-dimensional, large-scale immunophenotyping of huMG at the single-cell level, which facilitates their unambiguous profiling in health and disease.
This is a preview of subscription content, access via your institution
Access options








Similar content being viewed by others
Data availability
Source data associated with Figs. 4–7 can be accessed at https://flowrepository.org/id/FR-FCM-ZYM6.
References
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Sierra, A. et al. Microglia shape adult hippocampal neurogenesis through apoptosis-coupled phagocytosis. Cell Stem Cell 7, 483–495 (2010).
Parkhurst, C. N. et al. Microglia promote learning-dependent synapse formation through brain-derived neurotrophic factor. Cell 155, 1596–1609 (2013).
Prinz, M. & Priller, J. The role of peripheral immune cells in the CNS in steady state and disease. Nat. Neurosci. 20, 136–144 (2017).
Perry, V. H. & Holmes, C. Microglial priming in neurodegenerative disease. Nat. Rev. Neurol. 10, 217–224 (2014).
Colonna, M. & Butovsky, O. Microglia function in the central nervous system during health and neurodegeneration. Annu. Rev. Immunol. 35, 441–468 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Elmore, M. R. et al. Colony-stimulating factor 1 receptor signaling is necessary for microglia viability, unmasking a microglia progenitor cell in the adult brain. Neuron 82, 380–397 (2014).
Bruttger, J. et al. Genetic cell ablation reveals clusters of local self-renewing microglia in the mammalian central nervous system. Immunity 43, 92–106 (2015).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Orre, M. et al. Acute isolation and transcriptome characterization of cortical astrocytes and microglia from young and aged mice. Neurobiol. Aging 35, 1–14 (2014).
Grabert, K. et al. Microglial brain region-dependent diversity and selective regional sensitivities to aging. Nat. Neurosci. 19, 504–516 (2016).
Orre, M. et al. Isolation of glia from Alzheimer’s mice reveals inflammation and dysfunction. Neurobiol. Aging 35, 2746–2760 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Galatro, T. F. et al. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat. Neurosci. 20, 1162–1171 (2017).
Melief, J. et al. Characterizing primary human microglia: a comparative study with myeloid subsets and culture models. Glia 64, 1857–1868 (2016).
Mizee, M. R. et al. Isolation of primary microglia from the human post-mortem brain: effects of ante- and post-mortem variables. Acta Neuropathol. Commun. 5, 16 (2017).
Mildner, A., Huang, H., Radke, J., Stenzel, W. & Priller, J. P2Y12 receptor is expressed on human microglia under physiological conditions throughout development and is sensitive to neuroinflammatory diseases. Glia 65, 375–387 (2017).
Moore, C. S. et al. P2Y12 expression and function in alternatively activated human microglia. Neurol. Neuroimmunol. Neuroinflamm. 2, e80 (2015).
Gaudilliere, B. et al. Clinical recovery from surgery correlates with single-cell immune signatures. Sci. Transl. Med. 6, 255ra131 (2014).
Mei, H. E., Leipold, M. D., Schulz, A. R., Chester, C. & Maecker, H. T. Barcoding of live human peripheral blood mononuclear cells for multiplexed mass cytometry. J. Immunol. 194, 2022–2031 (2015).
Amir, A. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013).
Van der Maaten, L. & Hinton, G. Visualizing high-dimensional data using tSNE. J. Mach. Learn Res. 9, 2579–2605 (2008).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Becher, B. et al. High-dimensional analysis of the murine myeloid cell system. Nat. Immunol. 15, 1181–1189 (2014).
Holder, G. E. et al. Expression of the mannose receptor CD206 in HIV and SIV encephalitis: a phenotypic switch of brain perivascular macrophages with virus infection. J. Neuroimmune Pharmacol. 9, 716–726 (2014).
Cohen, M. et al. Newly formed endothelial cells regulate myeloid cell activity following spinal cord injury via expression of CD200 ligand. J. Neurosci. 37, 972–985 (2017).
Roederer, M., Treister, A., Moore, W. & Herzenberg, L. A. Probability binning comparison: a metric for quantitating univariate distribution differences. Cytometry 45, 37–46 (2001).
Orlova, D. Y. et al. Earth mover’s distance (EMD): a true metric for comparing biomarker expression levels in cell populations. PLoS One 11, e0151859 (2016).
Nichols, T. & Hayasaka, S. Controlling the familywise error rate in functional neuroimaging: a comparative review. Stat. Methods Med. Res. 12, 419–446 (2003).
Chatfield, M. & Mander, A. The Skillings-Mack test (Friedman test when there are missing data). Stata J. 9, 299–305 (2009).
Lun, A. T. L., Richard, A. C. & Marioni, J. C. Testing for differential abundance in mass cytometry data. Nat. Methods 14, 707–709 (2017).
O’Neill, K., Jalali, A., Aghaeepour, N., Hoos, H. & Brinkman, R. R. Enhanced flowType/RchyOptimyx: a BioConductor pipeline for discovery in high-dimensional cytometry data. Bioinformatics 30, 1329–1330 (2014).
Smith, A. M. & Dragunow, M. The human side of microglia. Trends Neurosci. 37, 125–135 (2014).
Schughart, K., Libert, C. & Kas, M. J. Controlling complexity: the clinical relevance of mouse complex genetics. Eur. J. Hum. Genet. 21, 1191–1196 (2013).
Durafourt, B. A., Moore, C. S., Blain, M. & Antel, J. P. Isolating, culturing, and polarizing primary human adult and fetal microglia. Methods Mol. Biol. 1041, 199–211 (2013).
Rustenhoven, J. et al. Isolation of highly enriched primary human microglia for functional studies. Sci. Rep. 6, 19371 (2016).
Melief, J. et al. Microglia in normal appearing white matter of multiple sclerosis are alerted but immunosuppressed. Glia 61, 1848–1861 (2013).
Olah, M. et al. An optimized protocol for the acute isolation of human microglia from autopsy brain samples. Glia 60, 96–111 (2012).
Lambert, C., Ase, A. R., Séguéla, P. & Antel, J. P. Distinct migratory and cytokine responses of human microglia and macrophages to ATP. Brain Behav. Immun. 24, 1241–1248 (2010).
Klegeris, A., Bissonnette, C. J. & McGeer, P. L. Modulation of human microglia and THP-1 cell toxicity by cytokines endogenous to the nervous system. Neurobiol. Aging 26, 673–682 (2005).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl. Acad. Sci. USA 113, E1738–E1746 (2016).
Bianchin, M. M. et al. Nasu-Hakola disease (polycystic lipomembranous osteodysplasia with sclerosing leukoencephalopathy–PLOSL): a dementia associated with bone cystic lesions. From clinical to genetic and molecular aspects. Cell. Mol. Neurobiol. 24, 1–24 (2004).
Korin, B. et al. High-dimensional, single-cell characterization of the brain’s immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Szulzewsky, F. et al. Human glioblastoma-associated microglia/monocytes express a distinct RNA profile compared to human control and murine samples. Glia 64, 1416–1436 (2016).
Hamann, J. et al. EMR1, the human homolog of F4/80, is an eosinophil-specific receptor. Eur. J. Immunol. 37, 2797–2802 (2007).
Askew, K. et al. Coupled proliferation and apoptosis maintain the rapid turnover of microglia in the adult brain. Cell Rep. 18, 391–405 (2017).
Tay, T.L. et al. A new fate mapping system reveals context-dependent random or clonal expansion of microglia. Nat. Neurosci. 20, 793–803 (2017).
Kotecha, N., Krutzik, P. O. & Irish, J. M. Web-based analysis and publication of flow cytometry experiments. Curr. Protoc. Cytom. 53, 10.17.1–10.17.24 (2010).
R Core Team. R: a language and environment for statistical computing http://www.r-project.org/ (R Foundation for Statistical Computing, 2014).
Finak, G. et al. OpenCyto: an open source infrastructure for scalable, robust, reproducible, and automated, end-to-end flow cytometry data analysis. PLoS Comput. Biol. 10, e1003806 (2014).
Jiang, M. CytoML: gatingML interface for openCyto. R package version 1.0.1 (R Project for Statistical Computing, 2016).
Spidlen, J., Leif, R. C., Moore, W., Roederer, M. & Brinkman, R. R. Gating-ML: XML-based gating descriptions in flow cytometry. Cytometry A 73A, 1151–1157 (2008).
van Dongen, S. & Enright, A.J. Metric distances derived from cosine similarity and Pearson and Spearman correlations. Preprint at arXiv https://arxiv.org/abs/1208.3145 (2012).
Shekhar, K., Brodin, P., Davis, M. M. & Chakraborty, A. K. Automatic classification of cellular expression by nonlinear stochastic embedding (ACCENSE). Proc. Natl Acad. Sci. USA 111, 202–207 (2014).
Chen, H. et al. Cytofkit: a Bioconductor package for an integrated mass cytometry data analysis pipeline. PLoS Comput. Biol. 12, e1005112 (2016).
Samusik, N., Good, Z., Spitzer, M. H., Davis, K. L. & Nolan, G. P. Automated mapping of phenotype space with single-cell data. Nat. Methods 13, 493–496 (2016).
Rogers, W. T. & Holyst, H. A. flowFP: a Bioconductor package for fingerprinting flow cytometric data. Adv. Bioinforma. https://doi.org/10.1155/2009/193947 (2009).
Japp, A. S. et al. Wild immunology assessed by multidimensional mass cytometry. Cytometry A 91, 85–95 (2017).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Oksanen, J. et al. vegan: community ecology package. (R Project for Statistical Computing, 2008).
Nichols, T. E. & Holmes, A. P. Nonparametric permutation tests for functional neuroimaging: a primer with examples. Hum. Brain Mapp. 15, 1–25 (2002).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Naumann, U., Luta, G. & Wand, M. P. The curvHDR method for gating flow cytometry samples. BMC Bioinformatics 11, 44 (2010).
Hahne, F. et al. flowCore: a Bioconductor package for high throughput flow cytometry. BMC Bioinformatics 10, 106 (2009).
Duong, T. ks: kernel density estimation and kernel discriminant analysis for multivariate data in R. J. Stat. Softw. 21, 1–16 (2007).
Duong, T., Goud, B. & Schauer, K. Closed-form density-based framework for automatic detection of cellular morphology changes. Proc. Natl Acad. Sci. USA 109, 8382–8387 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Diggins, K. E., Greenplate, A. R., Leelatian, N., Wogsland, C. E. & Irish, J. M. Characterizing cell subsets using marker enrichment modeling. Nat. Methods 14, 275–278 (2017).
Hodges, J. L. & Lehmann, E. L. Estimates of location based on rank tests. Ann. Math. Stat. 34, 598–611 (1963).
Rousseeuw, P. & Croux, C. Explicit scale estimators with high breakdown point. In L1-Statistical Analysis and Related Methods (ed. Dodge, Y.) 77–92 (North-Holland, 1992).
Aghaeepour, N. et al. Early immunologic correlates of HIV protection can be identified from computational analysis of complex multivariate T-cell flow cytometry assays. Bioinformatics 28, 1009–1016 (2012).
Aghaeepour, N. et al. RchyOptimyx: cellular hierarchy optimization for flow cytometry. Cytometry A 81, 1022–1030 (2012).
Acknowledgements
We thank C. Böttcher for excellent technical assistance with FACS analysis. We also acknowledge the assistance of the BCRT Flow Cytometry Lab (Charité – Universitätsmedizin Berlin, Germany). C.B. and J.P. were supported by the German Research Foundation (SFB TRR167, B05 & B07). J.P. received additional funding from the Berlin Institute of Health (CRG2aSP6) and the UK DRI (Momentum Award). S.S. was partially funded by the EU-H2020 project PACE (grant agreement number 733006). A.K. and E.P. were supported by stipends from the NeuroMac School (SFB TRR167, IRTG). A.K. received additional funding from the Cluster of Excellence NeuroCure. H.E.M. and A.R.S. were supported through grant Me3644/5-1. B.S. was supported by the German Research Foundation (SI 749/9-1, 749/10-1, CRC-TRR 241). B.S. and R.G. were supported by the Deutsche Krebhilfe (70112011). M.A.M.S. was supported by a 2014 NARSAD Young Investigator Grant from the Brain & Behavior Research Foundation and L.D.D.W. by the Virgo Consortium, funded by the Dutch government, project number FES0908. The psychiatric donor program of the Netherlands Brain Bank (NBB-Psy) is financially supported by the Netherlands Organization for Scientific Research (NWO). We acknowledge the Leibniz Science Campus for Chronic Inflammation for general support.
Author information
Authors and Affiliations
Consortia
Contributions
C.B. and J.P. conceived and designed the project. C.B., S.S., D.K., B.S. and R.G. designed the antibody panels for mass cytometry. M.A.M.S., G.J.L.S., E.M.H., R.S.K. and L.D.D.W. established and performed the isolation of postmortem huMG. P.F. and L.K. provided biopsy tissues from temporal lobe resections. A.R.S. and H.E.M. set up the fixation approach for cryostorage of human leucocytes and provided guidance in using the system. C.B. established the protocol for cryopreservation of isolated huMG. A.K. and E.P. performed barcoding and antibody staining for CyTOF. A.K. conducted FACS analysis of postmortem huMG. D.K. performed CyTOF measurements. E.J.S. and J.P. provided peripheral blood and cerebrospinal fluid samples. C.B. and S.S. analyzed the data. C.B., S.S., L.D.D.W. and J.P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Immunophenotypic profiling by single-cell mass cytometry.
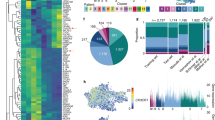
(a) Two-dimensional projections of single-cell data (Panel B) generated by t-SNE of PBMCs (n =4 biologically independent samples), CSF cells (n = 4 biologically independent samples) and brain mononuclear cells (n = 36 biologically independent samples). Each dot represents one cell. The color spectrum represents expression of TMEM119 (red denotes high expression, blue denotes no expression). TMEM119+ cells were gated as huMG (blue) and TMEM119- cells were gated as different circulating immune cells. (b) Representative scatter plots and histograms (two independently repeated experiments with similar results) show expression level of P2Y12 in TMEM119+ cells (upper panel) and vice versa of TMEM119 in P2Y12+ cells (lower panel). (c) The graph shows the quantitative frequencies of P2Y12+ cells in TMEM119+ cell population (P2Y12+TMEM119+, n = 20 biologically independent samples) and vice versa of TMEM119+ cells in P2Y12+cell population (TMEM119+P2Y12+, n = 20 biologically independent samples). Black lines show mean values of the data sets. The values show mean ± SD.
Supplementary Figure 2 Differential marker expressions of PBMCs, CSF cells, and human microglia.
(a) Histogram plots show expression levels of 55 markers analysed in HLA-DR+CD11c+ PBMCs (blue), CSF cells (orange) and huMG from SVZ (green), THA (red), CER (purple), GTS (brown) and GFM (pink). (b) Histogram plots compare expression levels of CD11b and CD115 of huMG and CD3+ T cells (negative for both markers). Expression levels of CD11b and CD115 of individual donors are also shown as histogram plots.
Supplementary Figure 3 Differential marker expressions of PBMCs, CSF cells, and human microglia.
Mean expression levels of selected markers in PBMCs (n = 4), CSF cells (n = 4) and huMG from SVZ (n = 8), THA (n = 8), Cer (n = 5), GTS (n = 8) and GFM (n = 7). Black lines show mean values of the data sets. Each dot represents one individual.
Supplementary Figure 4 CD11c and HLA-DR expression on human microglia.
(a) An overlaid dot plot shows the expression of HLA-DR and CD11c of PBMCs (blue), CSF cells (orange) and P2Y12+ huMG (green). (b) Individual contour plots show the expression of HLA-DR and CD11c of PBMCs, CSF cells, P2Y12+ and TMEM119+ huMG (two independently repeated experiments with similar results).
Supplementary Figure 5 Differential immunophenotypes of postmortem and fresh human microglia.
(a) Heat map and cluster dendrogram demonstrates the mean expression of all analyzed markers and relationships between postmortem GTS-huMG (green), postmortem GFM-huMG (orange) and huMG from fresh biopsies (blue). Heat colours have been scaled per marker (red denotes high and blue denotes low expression). (b) Heat map and cluster dendrogram demonstrates the mean expression of analyzed markers (excluded IRF-8 and P2Y12) and relationships between postmortem GTS-, GFM-huMG and huMG from fresh biopsies, respectively. (c) An overlaid t-SNE plot (left image) of all cells from all samples (green = postmortem GTS-huMG (n = 10), orange = postmortem GFM-huMG (n = 9) and blue = huMG from fresh biopsies (n = 3)). Two main clusters, G1 (blue) and G2 (orange), are detected. Overlaid histograms and single dot plots show mean expression levels of selected markers showing differential marker expressions between the two gates (G1 & G2 in a) in huMG from GTS (green), GFM (orange) and fresh biopsies (blue). Black lines show the mean of data sets.
Supplementary Figure 6
CD206 expression on human microglia and perivascular macrophages. (a) A dot plot graph shows mean expression levels of P2Y12 in CD206-negative huMG (filled dots) from GTS (blue, n = 8) and GFM (orange, n = 7) compared with CD206hi perivascular macrophage (pmΦ, circles) from GTS (blue, n = 8) and GFM (orange, n = 7). Black lines show the mean of data acquired by mass cytometry (CyTOF). (b) An overlaid histogram plot displays low expression of CD206 in P2Y12+ huMG from GTS (blue) and GFM (orange) showing the high autofluorescent background of postmortem samples analyzed by flow cytometry (FACS). Two experiments were independently repeated with similar results.
Supplementary Figure 7 CD206 expression by a subpopulation of human microglia as assessed by antibody Panel B.
(a) High-dimensional tSNE plots of concatenated FCS file (n = 36 biologically independent samples, Panel B). Each dot represents one cell. The color spectrum represents an expression level of CD206 (upper left), TMEM119 (upper middle), HLA-DR (upper right), CD86 (lower left) and CD36 (lower right). Red color denotes high expression, blue color denotes no expression. CD206low cell population is gated as “G1” (red circle) and “G2” (green circle). (b) Frequencies of each CD206-positive population in different brain regions (SVZ, n = 8; THA, n = 8; CER, n = 5; GTS, n = 8 & GFM, n = 7). The values of an individual donor were plotted in the same color. The black line represents the mean value.
Supplementary Figure 8 Donor-specific phenotypic differences in human microglia.
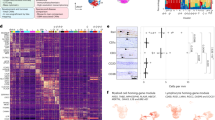
(a) The tSNE plots of concatenated FCS files from 36 huMG samples analysed by antibody Panel A with colours indicating distribution of each donor’s cells (same as in Fig. 5a). (b) Assessment of subject-to-subject variability by probability binning and bin-wise testing for donor-specific differences between 8 donors on n = 35 biologically independent samples – SVZ (n = 8); THA (n = 8); CER (n = 5); GTS (n = 8) and GFM (n =6) – (using negative binomial generalized linear model and quasi-likelihood test of edgeR). The colour spectrum indicates FDR-adjusted p-values thresholded to 0.05 (blue). Top significant bins were automatically gated (G1) to identify donor #6-specific huMG phenotype. (c) Median bin-expression levels as shown in boxplot representation for bins inside (red boxes, n = 24 bins) and outside (green boxes, n = 488 bins) of G1 and all markers indicate a specifically higher expression of CD64 and EMR1 in huMG samples of donor #6. Box center line and limits represent median, 16th and 84th percentiles; whiskers define the data minimum and maximum. (d) Heatmap representing the pairwise earth mover’s distances (EMD) between cellular density distributions among 31 huMG samples after t-SNE embedding without the 5 samples from donor #6 or after t-SNE-embedding of 36 huMG samples excluding the outlier markers CD64 and EMR1 (e). (f) The tSNE plots of concatenated FCS files (n = 31 biologically independent samples). The colouring indicates five brain regions (left image) and eight donors (middle image, without donor #6). The right image shows the smoothed representation of statistical tSNE map after thresholding to 0.05 FDR-adjusted p-values (blue). Three highly differentially abundant subsets with distinct phenotypes (indicated by arrows) were detected when donor #6 was excluded before embedding. The phenotypes of these subsets are equivalent to subset 1, 2 & 3 seen in Figs. 6 & 7, as shown with expression levels of the key markers (lower boxplot graph representing median bin-expression levels across 31 concatenated samples with subset 1 (n = 73 bins), 2 (n = 23 bins), 3 (n = 8 bins), and all cells – subsets (n = 408 bins). (g) The t-SNE plots of concatenated FCS files (n = 36 biologically independent samples). The colouring indicates five brain regions (left image) and nine donors (middle image). The tSNE embedding was performed without CD64 and EMR1. The right image shows the smoothed representation of statistical tSNE map after thresholding to 0.05 FDR-adjusted p-values (blue). Boxplots represent median bin-expression levels across 36 concatenated samples with subset 1 (n = 56 bins), 2 (n = 53 bins) and all cells – subsets (n = 403 bins). Two highly significant subsets (white lines) were detected when CD64 and EMR1 were excluded from embedding. Subset 1 is phenotypically identical to subset 1 shown in Figs. 6 & 7 with regard to co-expression of CD11c, CD45, CD68, CX3CR1, and HLA-DR, whereas the CD206+ subset 2 is equivalent to subsets 2 & 3 in Figs. 6 & 7, as shown in the boxplot graph below. For the boxplot graph shown in both Supplementary Fig. 8f, g, box center line and limits represent median, 16th and 84th percentiles; whiskers define the data minimum and maximum. (h) Comparable results (as shown in Fig. 6c and Supplementary Fig. 8f, g) are obtained when the analysis was performed using the cydar/edgeR framework for detection of differentially abundant (DA) subsets as hyperspheres in original multiparameter space (on same set of all markers or excluding CD64 and EMR1 in n = 35 biologically independent samples or in n= 30 independent samples excluding samples from donor #6). The colour spectrum indicates FDR-adjusted P-values (edgeR quasi-likelihood test) thresholded to 0.05, overlaid onto the tSNE map (non-significant hyperspheres in blue). (i) Mean signal intensity levels of Ki-67, Cyclin A and Cyclin B staining across four different defined subsets (subset 1, n = 36; subset 2, n = 34; subset 3, n = 36 and subset 4, n = 35). The black line represents the mean value. *P < 0.05, **P < 0.01, ***P < 0.001, ****P <0.0001, one-way ANOVA with Bonferroni correction.
Supplementary Figure 9 Assessment of regional differences in human microglia phenotypes by probability binning using the antibody Panel B.
The top panel shows the same tSNE map of concatenated FCS files (n = 36 biologically independent samples). The coloring in (a) indicates nine donors and five brain regions (left and right plot, respectively, legends shown in (e)), (b) overall cellular density of the concatenated files (left plot, spectrum from blue, low density, to red, high density) with a superimposed binning grid (512 bins) and unadjusted P-values (right plot) of bin-wise Skillings-Mack test for frequency differences between five brain regions, and (c) unadjusted p-values in differentially abundant hyperspheres (using cydar/edgeR framework), overlaid onto the tSNE map. The testing was performed on n = 35 independent huMG samples (SVZ (n = 8); THA (n = 8); CER (n = 5); GTS (n = 8) and GFM (n =6)) from 8 biologically independent donors. (d) tSNE plots of concatenated FCS files of each brain region. (e) Heatmap representing the pairwise earth mover’s distances (EMD) between cellular density distributions over the tSNE-space among all huMG samples. Hierarchical clustering highlights samples that have a similar phenotype, as indicated by low EMD values (top left colour bar).
Supplementary Figure 10 Region-dependent human microglia subpopulations analysed by antibody Panel B.
(a) Smoothed representation of statistical tSNE map (show in Supplementary Fig. 9b) after thresholding to 0.05 FDR-adjusted p-values (blue) shows selection of four differentially abundant subsets by automated gating (black contour lines). Skilling-Mack testing was performed on n = 35 biologically independent huMG samples (SVZ (n = 8); THA (n = 8); CER (n = 5); GTS (n = 8) and GFM (n =6)) from 8 biologically independent donors. (b) Snail plot shows mean marker expression levels of each huMG subset (subset 1 = red; 2 = orange; 3 = green and 4 = purple). The snail shell represents transversal (perpendicular) axis mapping marker expression levels on exponential scale. Each line denotes each sample (total of 36 samples). (c) Gates of four detected subsets overlaid on cell density tSNE plots of concatenated FCS files for each brain region (biologically independent samples of SVZ = 8; THA = 8; CER = 5; GTS = 8 and GFM = 7). (d) Top panel shows median bin-expression levels in boxplot representation of markers that discriminate between the four subsets (subset 1 = red box (n = 6 bins); 2 = orange box plot (n = 95 bins); 3 = green box (n = 4 bins); 4 = purple box (n = 12 bins); remaining cells (cells – subsets) = blue box (n = 395 bins) according to robust separation score. Box center line and limits represent median, 16th and 84th percentiles; whiskers define the data minimum and maximum.
Supplementary Figure 11 Aged-related marker expression on human microglia.
Scatter plots showing correlation between mean marker expression and age of all nine biologically independent donors analyzed (SVZ = 8; THA = 8; CER = 5; GTS = 8 and GFM = 7). *P < 0.05, **P < 0.01, non-parametric Spearman correlation test (r), two-sided.
Supplementary information
Supplementary Figures 1–11
Supplementary Figs. 1–11 and Supplementary Tables 1–5
Supplementary Software
Supplementary Software.
Rights and permissions
About this article
Cite this article
Böttcher, C., Schlickeiser, S., Sneeboer, M.A.M. et al. Human microglia regional heterogeneity and phenotypes determined by multiplexed single-cell mass cytometry. Nat Neurosci 22, 78–90 (2019). https://doi.org/10.1038/s41593-018-0290-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0290-2
This article is cited by
-
Disease and brain region specific immune response profiles in neurodegenerative diseases with pure and mixed protein pathologies
Acta Neuropathologica Communications (2024)
-
iPSC-derived PSEN2 (N141I) astrocytes and microglia exhibit a primed inflammatory phenotype
Journal of Neuroinflammation (2024)
-
The roles of tissue resident macrophages in health and cancer
Experimental Hematology & Oncology (2024)
-
Multiomic spatial landscape of innate immune cells at human central nervous system borders
Nature Medicine (2024)
-
Monocyte-derived cells invade brain parenchyma and amyloid plaques in human Alzheimer’s disease hippocampus
Acta Neuropathologica Communications (2023)