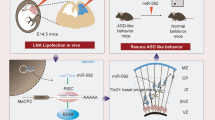
Abstract
Genetic analyses have linked microRNA-137 (MIR137) to neuropsychiatric disorders, including schizophrenia and autism spectrum disorder. miR-137 plays important roles in neurogenesis and neuronal maturation, but the impact of miR-137 loss-of-function in vivo remains unclear. Here we show the complete loss of miR-137 in the mouse germline knockout or nervous system knockout (cKO) leads to postnatal lethality, while heterozygous germline knockout and cKO mice remain viable. Partial loss of miR-137 in heterozygous cKO mice results in dysregulated synaptic plasticity, repetitive behavior, and impaired learning and social behavior. Transcriptomic and proteomic analyses revealed that the miR-137 mRNA target, phosphodiesterase 10a (Pde10a), is elevated in heterozygous knockout mice. Treatment with the Pde10a inhibitor papaverine or knockdown of Pde10a ameliorates the deficits observed in the heterozygous cKO mice. Collectively, our results suggest that MIR137 plays essential roles in postnatal neurodevelopment and that dysregulation of miR-137 potentially contributes to neuropsychiatric disorders in humans.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Genome-wide datasets are deposited at GEO under accession number GSE79661 (RNA-seq) and at the ProteomeXchange database under accession number PXD003874 (proteomics).
References
Pasquinelli, A. E. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat. Rev. Genet. 13, 271–282 (2012).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Li, X. & Jin, P. Roles of small regulatory RNAs in determining neuronal identity. Nat. Rev. Neurosci. 11, 329–338 (2010).
Issler, O. & Chen, A. Determining the role of microRNAs in psychiatric disorders. Nat. Rev. Neurosci. 16, 201–212 (2015).
Schratt, G. microRNAs at the synapse. Nat. Rev. Neurosci. 10, 842–849 (2009).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet. 94, 677–694 (2014).
Duan, J. et al. A rare functional noncoding variant at the GWAS-implicated MIR137/MIR2682 locus might confer risk to schizophrenia and bipolar disorder. Am. J. Hum. Genet. 95, 744–753 (2014).
Szulwach, K. E. et al. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell. Biol. 189, 127–141 (2010).
Smrt, R. D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 28, 1060–1070 (2010).
Siegert, S. et al. The schizophrenia risk gene product miR-137 alters presynaptic plasticity. Nat. Neurosci. 18, 1008–1016 (2015).
Guan, F. et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr. Res. 152, 97–104 (2014).
Carter, M. T. et al. Hemizygous deletions on chromosome 1p21.3 involving the DPYD gene in individuals with autism spectrum disorder. Clin. Genet. 80, 435–443 (2011).
Willemsen, M. H. et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J. Med. Genet. 48, 810–818 (2011).
D’Angelo, C. S., Moller Dos Santos, M. F., Alonso, L. G. & Koiffmann, C. P. Two new cases of 1p21.3 deletions and an unbalanced translocation t(8;12) among individuals with syndromic obesity. Mol. Syndromol. 6, 63–70 (2015).
Tucci, A., Ciaccio, C., Scuvera, G., Esposito, S. & Milani, D. MIR137 is the key gene mediator of the syndromic obesity phenotype of patients with 1p21.3 microdeletions. Mol. Cytogenet. 9, 80 (2016).
Pinkham, A. E., Hopfinger, J. B., Pelphrey, K. A., Piven, J. & Penn, D. L. Neural bases for impaired social cognition in schizophrenia and autism spectrum disorders. Schizophr. Res. 99, 164–175 (2008).
Dykens, E. M. Psychopathology in children with intellectual disability. J. Child Psychol. Psychiatry 41, 407–417 (2000).
Macbeth, A. H., Edds, J. S. & Young, W. S. III. Housing conditions and stimulus females: a robust social discrimination task for studying male rodent social recognition. Nat. Protoc. 4, 1574–1581 (2009).
Kleschevnikov, A. M. et al. Deficits in cognition and synaptic plasticity in a mouse model of Down syndrome ameliorated by GABAB receptor antagonists. J. Neurosci. 32, 9217–9227 (2012).
Marín, O. Interneuron dysfunction in psychiatric disorders. Nat. Rev. Neurosci. 13, 107–120 (2012).
Yizhar, O. et al. Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature 477, 171–178 (2011).
Eichler, S. A. & Meier, J. C. E-I balance and human diseases - from molecules to networking. Front. Mol. Neurosci. 1, 2 (2008).
van Dongen, S., Abreu-Goodger, C. & Enright, A. J. Detecting microRNA binding and siRNA off-target effects from expression data. Nat. Methods 5, 1023–1025 (2008).
Warren, R. P. et al. Increased frequency of the null allele at the complement C4b locus in autism. Clin. Exp. Immunol. 83, 438–440 (1991).
Odell, D. et al. Confirmation of the association of the C4B null allelle in autism. Hum. Immunol. 66, 140–145 (2005).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Fatemi, S. H., Reutiman, T. J., Folsom, T. D. & Thuras, P. D. GABA(A) receptor downregulation in brains of subjects with autism. J. Autism. Dev. Disord. 39, 223–230 (2009).
Fatemi, S. H., Folsom, T. D., Reutiman, T. J. & Thuras, P. D. Expression of GABA(B) receptors is altered in brains of subjects with autism. Cerebellum 8, 64–69 (2009).
Ramanathan, S. et al. A case of autism with an interstitial deletion on 4q leading to hemizygosity for genes encoding for glutamine and glycine neurotransmitter receptor sub-units (AMPA2, GLRA3, GLRB) and neuropeptide receptors NPY1R, NPY5R. BMC. Med. Genet. 5, 10 (2004).
Fernández, E. et al. Targeted tandem affinity purification of PSD-95 recovers core postsynaptic complexes and schizophrenia susceptibility proteins. Mol. Syst. Biol. 5, 269 (2009).
Beavo, J. A. & Brunton, L. L. Cyclic nucleotide research -- still expanding after half a century. Nat. Rev. Mol. Cell Biol. 3, 710–718 (2002).
Talkowski, M. E. et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149, 525–537 (2012).
Grauer, S. M. et al. Phosphodiesterase 10A inhibitor activity in preclinical models of the positive, cognitive, and negative symptoms of schizophrenia. J. Pharmacol. Exp. Ther. 331, 574–590 (2009).
Wellcome Trust Case Control, C.; Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Ripke, S. et al. Major Depressive Disorder Working Group of the Psychiatric. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Siuciak, J. A. et al. Inhibition of the striatum-enriched phosphodiesterase PDE10A: a novel approach to the treatment of psychosis. Neuropharmacology 51, 386–396 (2006).
Wilson, L. S. & Brandon, N. J. Emerging biology of PDE10A. Curr. Pharm. Des. 21, 378–388 (2015).
Tang, G. B. et al. The histone H3K27 demethylase UTX regulates synaptic plasticity and cognitive behaviors in mice. Front. Mol. Neurosci. 10, 267 (2017).
Silverman, J. L., Yang, M., Lord, C. & Crawley, J. N. Behavioural phenotyping assays for mouse models of autism. Nat. Rev. Neurosci. 11, 490–502 (2010).
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011).
Vicario-Abejón, C. Long-term culture of hippocampal neurons. Curr. Protoc. Neurosci. Chapter 3, 2 (2004).
Terashima, A. et al. An essential role for PICK1 in NMDA receptor-dependent bidirectional synaptic plasticity. Neuron 57, 872–882 (2008).
Xia, S. et al. p21-activated kinase 1 restricts tonic endocannabinoid signaling in the hippocampus. eLife 5, e14653 (2016).
Pagala, V. R. et al. Quantitative protein analysis by mass spectrometry. Methods Mol. Biol. 1278, 281–305 (2015).
Wang, H. et al. Systematic optimization of long gradient chromatography mass spectrometry for deep analysis of brain proteome. J. Proteome. Res. 14, 829–838 (2015).
Wang, X. et al. JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy. Mol. Cell. Proteomics. 13, 3663–3673 (2014).
Peng, J., Elias, J. E., Thoreen, C. C., Licklider, L. J. & Gygi, S. P. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome. Res. 2, 43–50 (2003).
Mertz, J. et al. Sequential elution interactome analysis of the mind bomb 1 ubiquitin ligase reveals a novel role in dendritic spine outgrowth. Mol. Cell. Proteomics. 14, 1898–1910 (2015).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 41, W77–W83 (2013).
Warde-Farley, D. et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214–W220 (2010).
Cerami, E. G. et al. Pathway Commons, a web resource for biological pathway data. Nucleic Acids Res. 39, D685–D690 (2011).
Jourquin, J., Duncan, D., Shi, Z. & Zhang, B. GLAD4U: deriving and prioritizing gene lists from PubMed literature. BMC Genomics 13(Suppl 8), S20 (2012).
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome. Biol. 11, R86 (2010).
Lie, D. C. et al. Wnt signalling regulates adult hippocampal neurogenesis. Nature 437, 1370–1375 (2005).
Liu, C. et al. Epigenetic regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell 6, 433–444 (2010).
Liu, P. P. et al. MiR-203 interplays with polycomb repressive complexes to regulate the proliferation of neural stem/progenitor cells. Stem Cell Rep. 9, 190–202 (2017).
Acknowledgements
We thank S. Warren and D. Cook for critical reading of the manuscript. This work was supported by the National Key R&D Program of China (2018YFA0108001 to Z.T.), Natural Science Foundation of China (31590831 and 91640204 to D.C. and 81571212 to Z.T.), Strategic Priority Research Program of Chinese Academy of Science (Grant XDB 19000000), National Institutes of Health (NS051630, NS079625, and MH102690 to P.J. and AG047928 to J.P.), Simons Foundation Autism Research Initiative (239320 to P.J.), and Hundred Talents Program of Chinese Academy of Science to Z.T. P.J. recieves support from the NARSAD Independent Investigator Award-Suzanne and John Golden Investigator sponsorship.
Author information
Authors and Affiliations
Contributions
Y.C., D.C., Z.-Q.T., and P.J. conceived the project. Y.C., Z.-M.W., W.T., and Xiaona Wang performed the experiments. Y.C., B.B., and Yuxin Li performed the bioinformatics analyses. W.T., X.W., Q.S., Z.C., and Z.-J.D. helped to generate and maintain the Mir137-knockout mouse strains. H.-L.Y., Z.-L.C., C.L., Y.-Y.W., and T.-W.M. helped to perform the IHC, ICC, and behavioral experiments. S.X., S.-F.Z., T.-W.M., A.L., Z.Z. and W.X. helped to perform the electrophysiological experiments. G.-B.T. and C.-M.L. performed Golgi staining analysis. J.P. advised on the proteomics experiment and interpreted the data. B.B., H.W., and Xusheng Wang helped to perform proteomics experiment. Yujing Li and L.L. provided technical support and constructed RNA-seq libraries. Y.K. and N.X. helped with the cell experiments. Y.C., D.C., Z.T., and P.J. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Generation and characterization of Mir137-knockout mice.
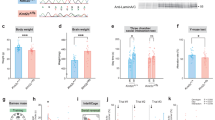
a, PCR genotyping confirmation of offspring of miR-137 global knockout (gKO) mice from heterozygous × heterozygous (Mir137+/– × Mir137+/–). The results are routinely repeated. b, Verification of decreased expression of mature miR-137 in Mir137+/– and Mir137–/– mice in both hippocampus (F2,6 = 308.1, P < 0.0001) and cortex (F2,6 = 1357, P < 0.0001) using independent real-time PCR (n = 3 mice per group). Data represent means ± s.e.m; One-way ANOVA with Tukey’s post hoc test; ****, P < 0.0001. c, Endogenous Dypd mRNA levels in hippocampus (t = 0.7159, P = 0.5136) and cortex (t = 0.3181, P = 0.7663) shows no difference in between Mir137+/+ and Mir137–/– mice (n = 3 mice per group). Data shown are means ± s.e.m. Unpaired Two-Tailed t test; n.s., nonsignificant. d, Kaplan-Meier graph shows survival curves of the Mir137+/+ (n = 8 mice), Mir137+/– (n = 8 mice) and Mir137–/– (n = 9 mice) mice groups. Mice in the Mir137–/– group exhibited significantly decreased survival (dot) relative to the other groups (P < 0.0001, Two-Tailed log-rank tests). All 9 tested Mir137–/– mice died by postnatal day 21.
Supplementary Figure 2 Impact of miR-137 on embryonic and postnatal development.
a, Morphology of miR-137 gKO embryos. Timed mating occurred using heterozygous × heterozygous. Embryos at embryonic day 15.5 (E15.5) were collected, and no apparent morphological difference among Mir137+/+, Mir137+/– and Mir137–/– mice were found. b, At the postnatal day 0 (P0), Mir137+/+ and Mir137–/– mice were similar regarding size/weight. c, The major organs (heart, liver, lung, and kidney) of Mir137–/– mice at P14 are smaller than those of Mir137+/+ and Mir137+/– littermates. d, Histopathological examinations of heart, liver, lung, and kidney tissues (H&E staining, 20X) showed no apparent differences in organs’ histological sections among each genotype. All the experiments were repeated independently at least three times with similar results.
Supplementary Figure 3 The loss of miR-137 in the nervous system leads to the synaptic overgrowth but has a negligible effect on cell viability.
a, A cartoon schematic indicating the position of brain section for IHC staining. PSD-95 and synaptophysin IHC staining were carried out on 40-μm thick floating sections containing cortical tissue from four pairs of littermate cKO mice, that is, Mir137loxP/+, Mir137loxP/+;Nestin-Cre and Mir137loxP/loxP;Nestin-Cre mice, at the age of postnatal day 18 (P18). b, Relative fluorescence intensity of PSD-95 (F2,9 = 6.733, P = 0.0163) and synaptophysin (F2,9 = 7.607, P = 0.0116) elevated upon the loss of miR-137 in the cortex (n = 4 mice per group). Image analyses and quantification were performed using ImageJ software. Data represent means ± s.e.m; One-way ANOVA with Tukey’s post hoc test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01. c, Partial loss of miR-137 resulted in a significant increase in PSD-95 (t = 5.331, P = 0.0060) and Synaptophysin (t = 3.566, P = 0.0235) protein levels (relative to Actin) in the hippocampus (n = 3 mice). Full-length blots are presented in Supplementary Fig. 15. Data are means ± s.e.m; Unpaired Two-Tailed t-test; *, P < 0.05; **, P < 0.01.d, Immunohistochemistry (IHC) staining of Caspase-3 and DAPI in Mir137loxP/+ and Mir137loxP/+;Nestin-Cre mice hippocampal tissues. Caspase-3 and DAPI IHC staining was carried out on 40-μm thick floating sections containing hippocampal tissue from four pairs of littermate Mir137loxP/+ and Mir137loxP/+;Nestin-Cre mice (n = 10 slices from 4 mice per group) at the age of postnatal day 0 (P0), 18 (P18) and 60 (P60). Relative fluorescence intensity of Caspase-3 and DAPI did not change upon the loss of miR-137 in the hippocampus (tP0 = 0.1055, P = 0.9171; tP18 = 0.1561, P = 0.8777; tP60 = 0.5895, P = 0.5629). Image analyses and quantification were performed using ImageJ software. Data are means ± s.e.m; Unpaired Two-Tailed t test; n.s., nonsignificant.
Supplementary Figure 4 The loss of miR-137 in germline leads to synaptic overgrowth.
a–b, IHC staining of PSD-95 and Synaptophysin in miR-137 gKO mice. Significant higher PSD-95 and Synaptophysin fluorescence intensity were observed in the CA1 region of the hippocampus in 8-week-old Mir137+/– mice (n = 4 mice; tPSD-95 = 2.841, P = 0.0295; tSynaptophysin = 5.912, P = 0.0010) (a) and P18-old Mir137–/– mice (n = 6; tPSD-95 = 3.206, P = 0.0094; tSynaptophysin = 2.987, P = 0.0136) (b). Data shown are means ± s.e.m. Unpaired Two-Tailed t test; *, P < 0.05; **, P < 0.01.
Supplementary Figure 5 Partial loss of miR-137 results in impaired spatial learning and memory.
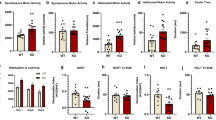
a–d, In the Morris water maze test, Mir137+/– mice (n = 13 mice) exhibited learning and memory deficits in the Morris water maze test. (a) During the training phase, in which four trials occurred per day for four successive days, Mir137+/– mice failed to diminish the latency to the platform, unlike Mir137+/+ mice (n = 12 mice). Data represent means ± s.e.m; tDay1 = 0.0022, P > 0.9999; tDay2 = 0.8859, P > 0.9999; tDay3 = 2.953, P = 0.0160; tDay4 = 2.64, P = 0.0390. Two-way ANOVA with Bonferroni post hoc test; *, P < 0.05). (b) In probe trials on Day 5, the swimming speeds between two groups of mice showed no significant difference (t = 1.202, P = 0.2414). In the subsequent probe test phase, Mir137+/– mice exhibited significantly longer times to find targets (t = 3.79, P = 0.0009) (c) and a fewer number of target crossings (t = 3.39, P = 0.0025) (d). Data shown are means ± s.e.m. Unpaired Two-Tailed t-test; n.s., nonsignificant; **, P < 0.01; ***, P < 0.001. e–h, The miR-137 gKO mice were subject to the Barnes maze test. (e) During the training phase of four trials conducted per day for four successive days, compared to the littermate Mir137+/+ mice (n = 10 mice), Mir137+/– mice (n = 11 mice) failed to diminish the time required of first entrance to the hiding box, indicating that loss of miR-137 results in impaired spatial memory. Data shown are means ± s.e.m; tDay1 = 0.6531, P > 0.9999; tDay2 = 2.658, P = 0.0356; tDay3 = 2.565, P = 0.0491; tDay4 = 3.155, P = 0.0092. Two-way ANOVA with Bonferroni post hoc test; *, P < 0.05; **, P < 0.01. Probe trials on Day 5 demonstrated that Mir137+/– mice also had spatial memory deficits as they spent less time in the target quadrant (t = 2.709, P = 0.0139) (f) and visited the target hole less often (t = 3.052, P = 0.0066) (g) than Mir137+/+ mice. The total distance traveled showed no significant difference between two groups (t = 1.766, P = 0.0935) (h). Data shown are means ± s.e.m. Unpaired Two-Tailed t-test; *, P < 0.05; ***, P < 0.001.
Supplementary Figure 6 Partial loss of miR-137 results in impaired social behaviors.
a, Representative data for social interactions in Mir137loxP/+ (n = 10 mice) and Mir137loxP/+;Nestin-Cre (n = 8 mice) mice. Mir137loxP/+;Nestin-Cre mice exhibited relatively lower levels of following (t = 1.104, P = 0.2859), crawling over/under (t = 1.522, P = 0.1474) and sniffing (Nose-to-nose: t = 2.266, P = 0.0377; Anogenital t = 1.888, P = 0.0773), and higher levels of self-grooming (t = 3.164, P = 0.0060) than Mir137loxP/+ mice, Data shown are means ± s.e.m. Unpaired Two-Tailed t-test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01. b, In 3-chamber test, both Mir137loxP/+ (n = 14 mice; t = 0.1743, P > 0.9999) and Mir137loxP/+;Nestin-Cre (n = 15 mice; t = 0.2245, P > 0.9999) mice had no preference for either left chamber or right chamber during the habituation phase. Data represent means ± s.e.m; Two-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant. c–e, In the 3-chamber test, Mir137+/– mice exhibited impaired social behavior. In miR-137 gKO mice, both Mir137+/+ (n =9 mice) and Mir137+/– (n = 12 mice) mice had no preference for either left chamber (t = 1.177, P > 0.9999) or right chamber (t = 0.598, P > 0.9999) during the habituation phase (c), but Mir137+/– mice lacked a social preference for a mouse over an object (Mir137+/+ mice: tEmpty vs. Stranger1 = 2.857, P = 0.0414; Mir137+/– mice: tEmpty vs. Stranger1 = 0.8122, P > 0.9999; Mir137+/+ vs. Mir137+/–: tStranger1 = 3.052, P = 0.0248) (d). Moreover, unlike Mir137+/+ control mice, Mir137+/– mice displayed impaired social novelty recognition by demonstrating no preference for a novel mouse over a familiar mouse (Mir137+/+ mice: tStranger2 vs. Stranger1 = 3.496, P = 0.0351; Mir137+/– mice: tStranger2 vs. Stranger1 = 0.9823, P > 0.9999; Mir137+/+ vs. Mir137+/–: tStranger2 = 3.496, P = 0.0073) (e). Data represent means ± s.e.m; Two-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01. f–g, In the social discrimination test, the Mir137+/– mice (n = 15 mice) exhibited impaired social discrimination ability. When compared with control mice, heterozygous cKO and gKO mice exhibited significantly fewer entries number (Mir137+/+ mice: tNovel vs. Familiar = 1.157, P > 0.9999; Mir137+/– mice: tNovel vs. Familiar = 0.2959, P > 0.9999; Mir137+/+ vs. Mir137+/–: tNovel = 2.744, P = 0.0489) (f) and lower % time spent (Mir137+/+ mice: tNovel vs. Familiar = 2.742, P = 0.0490; Mir137+/– mice: tNovel vs. Familiar = 1.598, P = 0.6943; Mir137+/+ vs. Mir137+/–: tNovel = 3.64, P = 0.0036) (g) in novel mouse zone. Data represent means ± s.e.m; Two-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01.
Supplementary Figure 7 Quantitative proteomics analysis by LC-MS/MS.
(a) An example of an MS2 raw scan of a peptide from protein Satb2. The inset is the relative intensity of each TMT reporter ion of the peptide from six samples respectively. Matched peptides MS b and y fragment ions are indicated in the peptide shown on the top and labeled in the spectrum. Western blotting validated the protein result below (see the first protein in the list). Xcorr score for identification of this spectrum is 74.83, suggesting high confidence. (b) Validation of some proteins by Western blotting. The table shows the relative abundance of the protein derived from the TMT-based MS quantification. The same protein lysate used for MS proteomic analysis underwent Western Blot analysis.
Supplementary Figure 8 Altered protein expression profile caused by the loss of miR-137.
(a) Sylamer enrichment landscape plots for 7-nt word. The x-axes represent the sorted protein list (n = 7,810 identified proteins were ranked from upregulated to downregulated in Mir137–/– versus Mir137+/+) from most upregulated (left) to most downregulated (right). The y-axes show the hypergeometric significance for each word at each leading bin. Positive values indicate enrichment (–log10(P-value)) and negative values, depletion (log10(P-value)). The horizontal dotted line represents an E-value threshold (Bonferroni-corrected) of 0.01 (orange) and 0.05 (green). (b) Gene Ontology (GO)- ‘Biological process’ analysis on the differentially expressed (DE) protein (n = 150 for upregulated DE genes; n = 227 for downregulated DE genes) identified in proteomics analysis, by comparing ‘Mir137–/– versus Mir137+/+.
Supplementary Figure 9 miR-137-mediated regulation of the genes implicated in neuropsychiatric disorders.
Venn diagrams show overlaps among the disease candidate genes with the differentially expressed (DE) proteins (that is, 377 DE genes) that identified by proteomics analysis (Mir137–/–versus Mir137+/+). The upregulated DE (DEup) and downregulated DE (DEdown) genes overlapped with ASD-, schizophrenia (SCZ)-, Intellectual disability (ID)-candidate genes, then overlapped with miR-137 predicted targets. Pearson’s chi-squared test calculated the P-value.
Supplementary Figure 10 Significant overlap exists between downregulated proteins and PSD–PSD95 complex proteins.
(a) Proteomic analysis identified 417 differentially expressed (DE) proteins (377 genes) in Mir137–/– mice compared to Mir137+/+ mice. By overlapping with previously reported PSD/PSD95 core complex proteins (Fernandez et al., 2009), we found a significant fraction of PSD/PSD95 core complex proteins were downregulated in Mir137–/– mice. (b) Many of these downregulated PSD/PSD95 core complex proteins are associated with ASD (in red) and schizophrenia (underlined). (c) A significant fraction of PSD/PSD95 core complex proteins has miR-137 predicted targets. Pearson’s chi-squared test calculated the P-value.
Supplementary Figure 11 Identification of the key mRNA targets of miR-137.
(a) To identify the direct mRNA targets of miR-137, we overlapped the predicted miR-137 targets with those DE proteins and identified 41 upregulated miR-137 predicted targets. The targets ranked from high to low in average log2 fold change of two replicates. (b) qPCR measured luciferase mRNA levels in two of our reporter gene assays (Ptpn2 and Pde10a). The decreased luciferase level was regulated by the overexpressed miR-137 rather than altered luciferase mRNA level, demonstrating by the little-changed expression level of luciferase. All data shown are means ± s.e.m. n = 9 independent samples. tPtpn2 = 0.954, P = 0.3543; tPde10a = 0.3656, P = 0.7195. Unpaired Two-Tailed t- test; n.s., nonsignificant. (c) Western blot was performed to validate the result of the proteomics analysis. In 12- day-old mice cortex tissue, protein expression of the most upregulated miR-137 target was examined (n = 3 mice per group). GAPDH (AM4300; Thermo Fisher) was used for loading controls. ImageJ software quantified the band intensities. All data shown are means ± s.e.m. tPtpn2 = 2.496, P = 0.1300; tSatb2 = 0.985, P = 0.3804; tPde10a = 4.217, P = 0.0135. Unpaired Two-Tailed t- test; n.s., nonsignificant; *, P < 0.05.
Supplementary Figure 12 Endogenous miR-137 regulates the expression of Pde10a through the predicted binding site in the 3′ UTR of Pde10a.
(a) The relative expression level of mature miR-137 in mouse hippocampus and Neuron-2a cells. n = 3 mice or Neuron-2a cell samples. All data shown are means ± s.e.m. t = 32.09, P < 0.0001. Unpaired Two-Tailed t- test. (b) The co-transfection experiment was performed for the two plasmids, including “Pde10a-3′ UTR” and “Pde10a-3′ UTRΔMir137” constructs in Neuron-2a cells. mirVana miR-137 inhibitor (ID: MH10513, 10uM) or Negative Control #1 was transfected to Neuron-2a cells 1 day before the plasmid transfection. All the other three samples were normalized to “Negative Control + Pde10a-3′ UTR”. There is a significant difference in gene expression when directly comparing Pde10a-3′ UTR and Pde10a-3′ UTRΔMir137 constructs. MiR-137 inhibition increased Pde10a-3′ UTR-dependent expression of a luciferase reporter gene. Renilla luciferase-Pde10a-3′ UTR expression was normalized to firefly luciferase. n = 9 independent experiments. Data represent means ± s.e.m. miR-137NC+ Pde10a-3' UTR vs. miR-137KD+ Pde10a-3' UTR: t = 4.182, P = 0.0020; miR-137NC+ Pde10a-3' UTR vs. miR-137 NC+ Pde10a-3' UTRΔMir137: t = 4.39, P = 0.0012; miR-137NC+ Pde10a-3' UTRΔMir137 vs. miR-137KD+ Pde10a-3' UTRΔMir137: t = 0.9731, P > 0.9999; miR-137 NC+ Pde10a-3' UTRΔMir137 vs. miR-137OE+ Pde10a-3' UTRΔMir137: t = 2.642, P = 0.0856; Two-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; **, P < 0.01.
Supplementary Figure 13 Papaverine could ameliorate the abnormalities in Mir137loxP/+;Nestin-Cre mice.
a, In the Barnes maze test, papaverine ameliorated spatial learning memory by significantly increasing the latency of first entrance to the hiding box during the training phase (Day4: Mir137loxP/+ + vehicle vs. Mir137loxP/+;Nestin-Cre + vehicle: t = 4.203, P = 0.0057; Mir137loxP/+ + papaverine vs. Mir137loxP/+;Nestin-Cre + papaverine: t = 1.398, P > 0.9999; Two-way ANOVA with Bonferroni post hoc test), without significantly affecting the total moving distance (Data represent means ± s.e.m; F3,34 = 0.6295, P = 0.6010; One-way ANOVA with Tukey’s post hoc test). Data represent means ± s.e.m; Mir137loxP/++vehicle: n = 9 mice; Mir137loxP/+ + papaverine: n = 9 mice; Mir137loxP/+;Nestin-Cre + vehicle: n = 10 mice; Mir137loxP/+;Nestin-Cre + papaverine: n = 10 mice. n.s., nonsignificant; **, P < 0.01. b, In the 3-chamber test, papaverine had an insignificant effect on the entry numbers in the left or right chamber of Mir137loxP/+ and Mir137loxP/+;Nestin-Cre mice during the habituation phase. Data represent means ± s.e.m; Mir137loxP/+ + vehicle: n = 9 mice, t = 0.3032, P > 0.9999; Mir137loxP/+ + papaverine: n = 9 mice, t = 0. 2421, P > 0.9999; Mir137loxP/+;Nestin-Cre + vehicle: n = 10 mice, t = 0.2584, P > 0.9999; Mir137loxP/+;Nestin-Cre + papaverine: n = 10 mice, t = 0.9475, P > 0.9999.Two-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant. c, In the open field test, papaverine did not change the locomotor activity, demonstrating by the little-changed moving distance (F3,63 = 2.243, P = 0.0920), but significantly ameliorated the anxiety-like behavior (F3,63 = 5.651, P = 0.0017). Data represent means ± s.e.m; Mir137loxP/+ + vehicle: n = 17 mice; Mir137loxP/+ + papaverine: n = 17 mice; Mir137loxP/+;Nestin-Cre + vehicle: n = 18 mice; Mir137loxP/+;Nestin-Cre + papaverine: n = 15 mice. One-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01. d–g, Papaverine could rescue the impaired neuronal phenotype in vitro. Primary hippocampal neurons were isolated from P0 littermate of from Mir137loxP/+ and Mir137loxP/+;Nestin-Cre mice. (d) After adding papaverine on the day in vitro 4 (DIV4), the primary hippocampal neurons usually died in between day DIV9 to DIV11. Thus we decided to investigate the dendritic growth at day DIV7. (e) Papaverine reduced dendritic complexity in Mir137loxP/+;Nestin-Cre neurons compared with controls, as determined by Sholl analysis (Data represent means ± s.e.m; Interaction: F102,1015 = 4.797, P < 0.0001; Distance: F34,1015 = 62.46, P < 0.0001; Treatment: F3,1015 = 233.3, P < 0.0001. Two-way ANOVA with Bonferroni post hoc test). With papaverine treatment, the dendritic length (F3,29 = 41.12, P < 0.0001) (f) and dendritic nodes (F3,29 = 18.7, P < 0.0001) (g) were significantly reduced in both Mir137loxP/+ and Mir137loxP/+;Nestin-Cre neurons. Data represent means ± s.e.m; One-way ANOVA with Bonferroni post hoc test; *, P < 0.05; **, P < 0.01; ***, P < 0.001. Mir137loxP/+ + vehicle: n = 7 samples; Mir137loxP/+ + papaverine: n = 8 samples; Mir137loxP/+;Nestin-Cre + vehicle: n = 9 samples; Mir137loxP/+;Nestin-Cre + papaverine: n = 9 samples.
Supplementary Figure 14 Knockdown of Pde10a ameliorates the abnormal behaviors associated with the partial loss of miR-137.
a, The knockdown efficiencies of three lentivirus lines encoding shRNA targeting Pde10a (sh-Pde10a) were tested in primary neurons. The ICC staining by using PDE10A antibody indicated the #1 lentivirus line had the most significant knockdown efficiency. Thus we used this line in the following experiments. n = 10 slices from 4 independent experiment. Data represent means ± s.e.m; F3,36 = 33.67, P < 0.0001; One-way ANOVA with Bonferroni post hoc test; ****, P < 0.0001. b-d, Knockdown of Pde10a ameliorated the abnormal behaviors in Mir137loxP/+;Nestin-Cre mice. In self-grooming test (F3,33 = 14.62, P < 0.0001), sh-Pde10a resulted in less time spent grooming in Mir137loxP/+;Nestin-Cre, although not statistically significant (Mir137loxP/+;Nestin-Cre + sh-Neg vs. Mir137loxP/+;Nestin-Cre + sh-Pde10a: t = 2.292, P = 0.1705) (b). In the marble-burying test (F3,33 = 4.247, P = 0.0121), sh-Pde10a ameliorated the impaired repetitive behaviors in Mir137loxP/+;Nestin-Cre mice to the same level in Mir137loxP/+ mice (Mir137loxP/+ + sh-Pde10a vs. Mir137loxP/+;Nestin-Cre + sh-Pde10a: t = 0.935, P > 0.9999) (c). In the open field test, papaverine did not change the locomotor activity, demonstrating by the little-changed moving distance (F3,32 = 2.153, P = 0.1129), but significantly ameliorated the anxiety-like behavior (F3,32 = 9.899, P < 0.0001) (d). Data represent means ± s.e.m; Mir137loxP/+ + sh-Neg: n = 9 mice; Mir137loxP/+ + sh-Pde10a: n = 10 mice; Mir137loxP/+;Nestin-Cre + sh-Neg: n = 9 mice; Mir137loxP/+;Nestin-Cre + sh-Pde10a: n = 9 mice. One-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; *, P < 0.05; ***, P < 0.001; ****, P < 0.0001. e, At DIV 14, cultured hippocampal neurons from Mir137loxP/+;Nestin-Cre mice (n = 3 mice) had more PSD-95 punctas (left) and Synaptophysin punctas (right) compared with Mir137loxP/+ mice (n = 3 mice). Knockdown of Pde10a significantly reduced the spine numbers of PSD-95 (n = 41 slices from 3 samples per group. F3,164 = 13.71, P < 0.0001) and synaptophysin (n = 31 slices from 3 samples per group. F3,120 = 24.93, P < 0.0001) in the Mir137loxP/+;Nestin-Cre neurons to similar levels in Mir137loxP/+ neurons. Data represent means ± s.e.m; One-way ANOVA with Bonferroni post hoc test; n.s., nonsignificant; *, P < 0.05; **, P < 0.01; ***, P < 0.001.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15
Supplementary Table 1
Summary of primers used in the experiments.
Supplementary Table 2
Summary of proteomics analyses.
Supplementary Table 3
Summary of RNA-seq analyses.
Supplementary Table 4
Summary of miR-137 predicted targets and candidate genes associated with ASD, schizophrenia, and ID.
Rights and permissions
About this article
Cite this article
Cheng, Y., Wang, ZM., Tan, W. et al. Partial loss of psychiatric risk gene Mir137 in mice causes repetitive behavior and impairs sociability and learning via increased Pde10a. Nat Neurosci 21, 1689–1703 (2018). https://doi.org/10.1038/s41593-018-0261-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0261-7
This article is cited by
-
Beyond traditional pharmacology: evaluating phosphodiesterase inhibitors in autism spectrum disorder
Neuropsychopharmacology (2024)
-
Divergent projections of the prelimbic cortex mediate autism- and anxiety-like behaviors
Molecular Psychiatry (2023)
-
Remember the poke: microRNAs are required for long-term memory formation following operant conditioning in Lymnaea
Journal of Comparative Physiology A (2023)
-
Epigenome-wide DNA methylation analysis of whole blood cells derived from patients with GAD and OCD in the Chinese Han population
Translational Psychiatry (2022)
-
Signalling pathways in autism spectrum disorder: mechanisms and therapeutic implications
Signal Transduction and Targeted Therapy (2022)