Abstract
Schizophrenia is a severely debilitating neurodevelopmental disorder. Establishing a causal link between circuit dysfunction and particular behavioral traits that are relevant to schizophrenia is crucial to shed new light on the mechanisms underlying the pathology. We studied an animal model of the human 22q11 deletion syndrome, the mutation that represents the highest genetic risk of developing schizophrenia. We observed a desynchronization of hippocampal neuronal assemblies that resulted from parvalbumin interneuron hypoexcitability. Rescuing parvalbumin interneuron excitability with pharmacological or chemogenetic approaches was sufficient to restore wild-type-like CA1 network dynamics and hippocampal-dependent behavior during adulthood. In conclusion, our data provide insights into the network dysfunction underlying schizophrenia and highlight the use of reverse engineering to restore physiological and behavioral phenotypes in an animal model of neurodevelopmental disorder.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request
References
Uhlhaas, P. J. & Singer, W. Abnormal neural oscillations and synchrony in schizophrenia. Nat. Rev. Neurosci. 11, 100–113 (2010).
Stephan, K. E., Friston, K. J. & Frith, C. D. Dysconnection in schizophrenia: from abnormal synaptic plasticity to failures of self-monitoring. Schizophr. Bull. 35, 509–527 (2009).
Sigurdsson, T., Stark, K. L., Karayiorgou, M., Gogos, J. A. & Gordon, J. A. Impaired hippocampal-prefrontal synchrony in a genetic mouse model of schizophrenia. Nature 464, 763–767 (2010).
Cardin, J. A. et al. Driving fast-spiking cells induces gamma rhythm and controls sensory responses. Nature 459, 663–667 (2009).
Amilhon, B. et al. Parvalbumin interneurons of hippocampus tune population activity at theta frequency. Neuron 86, 1277–1289 (2015).
Gulyás, A. I. et al. Parvalbumin-containing fast-spiking basket cells generate the field potential oscillations induced by cholinergic receptor activation in the hippocampus. J. Neurosci. 30, 15134–15145 (2010).
Cho, K. K. et al. Gamma rhythms link prefrontal interneuron dysfunction with cognitive inflexibility in Dlx5/6(+/-) mice. Neuron 85, 1332–1343 (2015).
Gonzalez-Burgos, G., Cho, R. Y. & Lewis, D. A. Alterations in cortical network oscillations and parvalbumin neurons in schizophrenia. Biol. Psychiatry 77, 1031–1040 (2015).
Lewis, D. A., Curley, A. A., Glausier, J. R. & Volk, D. W. Cortical parvalbumin interneurons and cognitive dysfunction in schizophrenia. Trends Neurosci. 35, 57–67 (2012).
Steullet, P. et al. Oxidative-stress-driven parvalbumin interneuron impairment as a common mechanism in models of schizophrenia. Mol. Psychiatry 22, 936–943 (2017).
Sauer, J. F., Strüber, M. & Bartos, M. Impaired fast-spiking interneuron function in a genetic mouse model of depression. eLife 4, e04979 (2015).
Nguyen, R. et al. Parvalbumin and GAD65 interneuron inhibition in the ventral hippocampus induces distinct behavioral deficits relevant to schizophrenia. J. Neurosci. 34, 14948–14960 (2014).
Del Pino, I. et al. Erbb4 deletion from fast-spiking interneurons causes schizophrenia-like phenotypes. Neuron 79, 1152–1168 (2013).
Belforte, J. E. et al. Postnatal NMDA receptor ablation in corticolimbic interneurons confers schizophrenia-like phenotypes. Nat. Neurosci. 13, 76–83 (2010).
Korotkova, T., Fuchs, E. C., Ponomarenko, A., von Engelhardt, J. & Monyer, H. NMDA receptor ablation on parvalbumin-positive interneurons impairs hippocampal synchrony, spatial representations, and working memory. Neuron 68, 557–569 (2010).
Carlén, M. et al. A critical role for NMDA receptors in parvalbumin interneurons for gamma rhythm induction and behavior. Mol. Psychiatry 17, 537–548 (2012).
Bhugra, D. The global prevalence of schizophrenia. PLoS Med. 2, e151 (2005).
Karayiorgou, M., Simon, T. J. & Gogos, J. A. 22q11.2 microdeletions: linking DNA structural variation to brain dysfunction and schizophrenia. Nat. Rev. Neurosci. 11, 402–416 (2010).
Karayiorgou, M. et al. Schizophrenia susceptibility associated with interstitial deletions of chromosome 22q11. Proc. Natl Acad. Sci. USA 92, 7612–7616 (1995).
Moutin, E. et al. Palmitoylation of cdc42 promotes spine stabilization and rescues spine density deficit in a mouse model of 22q11.2 deletion syndrome. Cereb. Cortex 27, 3618–3629 (2017).
Stark, K. L. et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat. Genet. 40, 751–760 (2008).
Piskorowski, R. A. et al. Age-dependent specific changes in area CA2 of the hippocampus and social memory deficit in a mouse model of the 22q11.2 deletion syndrome. Neuron 89, 163–176 (2016).
Drew, L. J. et al. Evidence for altered hippocampal function in a mouse model of the human 22q11.2 microdeletion. Mol. Cell. Neurosci. 47, 293–305 (2011).
Li, K. X. et al. Neuregulin 1 regulates excitability of fast-spiking neurons through Kv1.1 and acts in epilepsy. Nat. Neurosci. 15, 267–273 (2011).
Harris, K. D., Csicsvari, J., Hirase, H., Dragoi, G. & Buzsáki, G. Organization of cell assemblies in the hippocampus. Nature 424, 552–556 (2003).
Jones, C. A., Watson, D. J. & Fone, K. C. Animal models of schizophrenia. Br. J. Pharmacol. 164, 1162–1194 (2011).
Mena, A. et al. Reduced prepulse inhibition as a biomarker of schizophrenia. Front. Behav. Neurosci. 10, 202 (2016).
Long, J. M. et al. Behavior of mice with mutations in the conserved region deleted in velocardiofacial/DiGeorge syndrome. Neurogenetics 7, 247–257 (2006).
Swerdlow, N. R., Braff, D. L. & Geyer, M. A. Sensorimotor gating of the startle reflex: what we said 25 years ago, what has happened since then, and what comes next. J. Psychopharmacol. 30, 1072–1081 (2016).
Mier, D. et al. Evidence for altered amygdala activation in schizophrenia in an adaptive emotion recognition task. Psychiatry Res. 221, 195–203 (2014).
Paylor, R. et al. Mice deleted for the DiGeorge/velocardiofacial syndrome region show abnormal sensorimotor gating and learning and memory impairments. Hum. Mol. Genet. 10, 2645–2650 (2001).
Cannon, T. D. How schizophrenia develops: cognitive and brain mechanisms underlying onset of psychosis. Trends. Cogn. Sci. 19, 744–756 (2015).
Lieberman, J. A. et al. Hippocampal dysfunction in the pathophysiology of schizophrenia: a selective review and hypothesis for early detection and intervention. Mol. Psychiatry. https://doi.org/10.1038/mp.2017.249 (2018).
Zaremba, J. D. et al. Impaired hippocampal place cell dynamics in a mouse model of the 22q11.2 deletion. Nat. Neurosci. 20, 1612–1623 (2017).
Stefansson, H. et al. Neuregulin 1 and susceptibility to schizophrenia. Am. J. Hum. Genet. 71, 877–892 (2002).
Mukai, J. et al. Molecular substrates of altered axonal growth and brain connectivity in a mouse model of schizophrenia. Neuron 86, 680–695 (2015).
Tamura, M., Mukai, J., Gordon, J. A. & Gogos, J. A. Developmental inhibition of Gsk3 rescues behavioral and neurophysiological deficits in a mouse model of schizophrenia predisposition. Neuron 89, 1100–1109 (2016).
Hu, H., Gan, J. & Jonas, P. Interneurons. Fast-spiking, parvalbumin+ GABAergic interneurons: from cellular design to microcircuit function. Science 345, 1255263 (2014).
Yamada, J. & Jinno, S. Molecular heterogeneity of aggrecan-based perineuronal nets around five subclasses of parvalbumin-expressing neurons in the mouse hippocampus. J. Comp. Neurol. 525, 1234–1249 (2017).
Klausberger, T. et al. Spike timing of dendrite-targeting bistratified cells during hippocampal network oscillations in vivo. Nat. Neurosci. 7, 41–47 (2004).
Lee, S. H. et al. Parvalbumin-positive basket cells differentiate among hippocampal pyramidal cells. Neuron 82, 1129–1144 (2014).
Donato, F., Chowdhury, A., Lahr, M. & Caroni, P. Early- and late-born parvalbumin basket cell subpopulations exhibiting distinct regulation and roles in learning. Neuron 85, 770–786 (2015).
Viney, T. J. et al. Network state-dependent inhibition of identified hippocampal CA3 axo-axonic cells in vivo. Nat. Neurosci. 16, 1802–1811 (2013).
Howard, A., Tamas, G. & Soltesz, I. Lighting the chandelier: new vistas for axo-axonic cells. Trends Neurosci. 28, 310–316 (2005).
Curley, A. A. & Lewis, D. A. Cortical basket cell dysfunction in schizophrenia. J. Physiol. (Lond.) 590, 715–724 (2012).
Bartos, M. et al. Fast synaptic inhibition promotes synchronized gamma oscillations in hippocampal interneuron networks. Proc. Natl Acad. Sci. USA 99, 13222–13227 (2002).
Katona, I., Acsády, L. & Freund, T. F. Postsynaptic targets of somatostatin-immunoreactive interneurons in the rat hippocampus. Neuroscience 88, 37–55 (1999).
Acsády, L., Görcs, T. J. & Freund, T. F. Different populations of vasoactive intestinal polypeptide-immunoreactive interneurons are specialized to control pyramidal cells or interneurons in the hippocampus. Neuroscience 73, 317–334 (1996).
Harris, K. D. et al. Classes and continua of hippocampal CA1 inhibitory neurons revealed by single-cell transcriptomics. PLoS Biol. 16, e2006387 (2018).
Hamm, J. P., Peterka, D. S., Gogos, J. A. & Yuste, R. Altered cortical ensembles in mouse models of schizophrenia. Neuron 94, 153–167.e8 (2017).
Merscher, S. et al. TBX1 is responsible for cardiovascular defects in velo-cardio-facial/DiGeorge syndrome. Cell 104, 619–629 (2001).
Hippenmeyer, S. et al. A developmental switch in the response of DRG neurons to ETS transcription factor signaling. PLoS Biol. 3, e159 (2005).
Kastin, A. J., Akerstrom, V. & Pan, W. Neuregulin-1-beta1 enters brain and spinal cord by receptor-mediated transport. J. Neurochem. 88, 965–970 (2004).
Rösler, T. W. et al. Biodistribution and brain permeability of the extracellular domain of neuregulin-1-β1. Neuropharmacology 61, 1413–1418 (2011).
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G-protein-coupled receptors potently activated by an inert ligand. Proc. Natl Acad. Sci. USA 104, 5163–5168 (2007).
Stoppini, L., Buchs, P. A. & Muller, D. A simple method for organotypic cultures of nervous tissue. J. Neurosci. Methods 37, 173–182 (1991).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Urban, D. J. & Roth, B. L. DREADDs (designer receptors exclusively activated by designer drugs): chemogenetic tools with therapeutic utility. Annu. Rev. Pharmacol. Toxicol. 55, 399–417 (2015).
Fisahn, A., Pike, F. G., Buhl, E. H. & Paulsen, O. Cholinergic induction of network oscillations at 40 Hz in the hippocampus in vitro. Nature 394, 186–189 (1998).
Pietersen, A. N. et al. Transition between fast and slow gamma modes in rat hippocampus area CA1 in vitro is modulated by slow CA3 gamma oscillations. J. Physiol. (Lond.) 592, 605–620 (2014).
Marissal, T. et al. Pioneer glutamatergic cells develop into a morpho-functionally distinct population in the juvenile CA3 hippocampus. Nat. Commun. 3, 1316 (2012).
Gschwend, O. et al. Neuronal pattern separation in the olfactory bulb improves odor discrimination learning. Nat. Neurosci. 18, 1474–1482 (2015).
Abraham, N. M., Guerin, D., Bhaukaurally, K. & Carleton, A. Similar odor discrimination behavior in head-restrained and freely moving mice. PLoS One 7, e51789 (2012).
Gschwend, O., Beroud, J. & Carleton, A. Encoding odorant identity by spiking packets of rate-invariant neurons in awake mice. PLoS One 7, e30155 (2012).
Gschwend, O., Beroud, J., Vincis, R., Rodriguez, I. & Carleton, A. Dense encoding of natural odorants by ensembles of sparsely activated neurons in the olfactory bulb. Sci. Rep. 6, 36514 (2016).
Patterson, M. A., Lagier, S. & Carleton, A. Odor representations in the olfactory bulb evolve after the firstbreath and persist as an odor afterimage. Proc. Natl Acad. Sci. USA 110, E3340–E3349 (2013).
Rossant, C. et al. Spike sorting for large, dense electrode arrays. Nat. Neurosci. 19, 634–641 (2016).
Csicsvari, J., Hirase, H., Czurko, A. & Buzsáki, G. Reliability and state dependence of pyramidal cell-interneuron synapses in the hippocampus: an ensemble approach in the behaving rat. Neuron 21, 179–189 (1998).
Barthó, P. et al. Characterization of neocortical principal cells and interneurons by network interactions and extracellular features. J. Neurophysiol. 92, 600–608 (2004).
Engel, M. et al. Neuregulin 1 prevents phencyclidine-induced behavioral impairments and disruptions to GABAergic signaling in mice. Int. J. Neuropsychopharmacol. 18, pyu114 (2015).
Stefanelli, T., Bertollini, C., Lüscher, C., Muller, D. & Mendez, P. Hippocampal somatostatin interneurons control the size of neuronal memory ensembles. Neuron 89, 1074–1085 (2016).
Alexander, G. M. et al. Remote control of neuronal activity in transgenic mice expressing evolved G-protein-coupled receptors. Neuron 63, 27–39 (2009).
Acknowledgements
This paper is dedicated to the memory of D. Müller, whose ideas had inspired this study. He will continue to inspire all of us. We thank R. Kucherlapati (Harvard University) for generously providing the Lgdel/+ mice. We thank L. Jourdain and M.-P. Hervé for their technical support. We thank Y. Bernardelli, P. Mendez Garcia, T. Stefanelli, S. Eliez, P. Caroni, and other members of the NCCR SYNAPSY for helpful discussions and/or comments on the manuscript. This research was supported by the University of Geneva, the Swiss National Science Foundation (grant numbers: 31003A_172878 to A.C., 310030B_144080 to D.M. and 31003A_170114 to I.R.), the National Center of Competence in Research (NCCR) “SYNAPSY - The Synaptic Bases of Mental Diseases” financed by the Swiss National Science Foundation (grant 51NF40-158776, D.M. and A.C.), the Novartis Foundation for medical-biological research (grant 17A057 to A.C.) and the Lejeune Foundation (T.M.).
Author information
Authors and Affiliations
Contributions
T.M., D.M., and A.C. conceived the study. T.M., R.S., C.B., S.M., M.D.R., I.R., D.M., and A.C. contributed to the experimental design. TM performed the calcium imaging experiments and analyzed the data. C.B. carried out the in vitro electrophysiological experiments. R.S. performed the in vivo electrophysiological recordings and developed most of the MATLAB-based programs used for the analysis of the calcium imaging and the electrophysiological recordings. M.D.R. developed some MATLAB-based scripts used to analyze calcium-imaging data. T.M. and C.B. performed stereotaxic viral infections. T.M. performed behavioral experiments with the help of S.M. A.C., T.M., R.S., and I.R. wrote and edited the manuscript with comments from all of the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 A subpopulation of neurons displays lower calcium transient frequency in Lgdel/+ mice when compared to WT mice.
(a,b) Cumulative distributions of the frequency of calcium events recorded in Lgdel/+ and WT mice. The distributions are plotted either for every slice (a) or for the mean (± SEM, b). Statistical comparisons were done using two-way repeated measures ANOVA with two-sided Bonferroni post-hoc test (*P < 0.05: see Supplementary Table 1 for detailed statistics).
Supplementary Figure 2 Representative example to illustrate the co-activation analysis of calcium transient onsets.
(a,b) Raster plots of the calcium transient onsets for the actual data imaged in CA1 neurons of a representative WT slice. (c,d) Raster plots of the calcium transient onsets for the same dataset after one iteration of temporal shuffling (see methods, coactive neurons are indicated in cyan). (e,f) Ten thousands shuffling iterations were made and the iteration with the maximum number of co-activated cells at each frame was chosen as the threshold (magenta). All frames were considered as significant when the number of neurons in the actual data exceeded this threshold and were used to calculate the duration of the co-activations over the whole recordings. The numbers of active cells during these frames were considered as significant co-active neurons above chance (magenta).
Supplementary Figure 3 Representative example to illustrate the cross-correlation analysis of paired cells.
(a) Calcium traces of two neurons (blue and red) and identification of their calcium transient onsets. (b) The cross-correlation is presented as a histogram of the number of coincidences (shaded area) as a function of frame lags. A temporal shuffling procedure was used to estimate the significance threshold (dashed line, see methods). (c) Each time the number of coincidences exceeded the threshold, the corresponding frame lag (lower panel) was considered as significant. Data is shown for a few pairs of neurons.
Supplementary Figure 4 Synaptic parameters recorded in PCs and PVIs from WT and Lgdel/+ mice.
(a-d) Patch-clamp recordings of CA1 PCs from WT and Lgdel/+ mice. Summary graphs of the mean sEPSC (b) and mean sIPSC (c) amplitude. (d) Summary graph of the mean mEPSC IEI. (e-h) Patch-clamp recordings of CA1 PVIs from PvalbtdTom;+/+ and PvalbtdTom;Lgdel/+ mice. Summary graphs of the mean sEPSC (f) and mean sIPSC (g) amplitude. (h) Summary graph of the mean mEPSC IEI. All data are presented as mean ± SEM ; each circle represents a cell, number of cells indicated in parentheses; two-sided Mann-Whitney test were performed.
Supplementary Figure 5 Effect of NRG1P application in vitro and in vivo on WT mice.
(a-d) Network dynamics in slices from WT mice treated or not with NRG1P (data for not treated slices are the same as in Fig. 1). Percentage of co-active neurons above chance (a) and occurrence of these co-activations (b). (c) Percentage of correlated pairs of neurons above chance. (d) Distributions of significant time lags in correlated pairs (two-way repeated measures ANOVA, interaction genotype x time lags: F20,441 = 0.82, P = 0.69). The grey dashed line represents the equidistribution. Box: 25-75th percentiles, whiskers: 10-90th percentiles, white bar: median. (e,f) Distribution of the LFP PSDs in the theta band for WT mice before (same data as in Fig. 6a) and after NRG1P injection (PSDs averaged between 5 to 8 Hz in f, the number of shank/mouse pairs is indicated in parentheses). (g) Percentage of correlated neurons (median) in the CA1 region of awake mice as a function of time lags for WT animals before (same data as in Fig. 6d) and after NRG1P injection. The numbers of recorded animals are indicated in parentheses. (h) Distribution of central peaks of correlation curves (computed in ± 60 ms time window). (i-k) WT mice in the open field arena and during contextual fear conditioning after NRG1P administration. (i) Locomotor tracks during 10 min in an open field. (j) Distance travelled in the open field. (k) WT mice freezing behaviour in the control conditions and after treatments with NRG1P. All data are presented as mean ± SEM unless otherwise indicated. Each circle represents a mouse (total number of animal indicated in parentheses). (a-c,j,k): Two-sided Mann-Whitney test. (f,h): Two-sided Wilcoxon paired test.
Supplementary Figure 6 Effects of PVI activation on various parameters in Lgdel/+ mice during calcium imaging.
(a) Proportion of neurons displaying Ca2+ transients. (b) Frequency (c), amplitude (c) and duration (d) of Ca2+ transients recorded in CA1 neurons. (e) Percentage of co-active neurons above random co-activation. (f) Occurrences of the co-activations. Statistical comparisons were done with two-sided Mann-Whitney test. Data are presented as mean ± SEM (number of slices in indicated in parentheses).
Supplementary Figure 7 In vivo local field potential recordings using polytrodes and DREADD infection in the hippocampus.
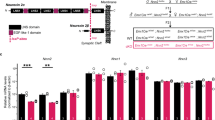
(a) Photograph of the track resulting from the implantation in the dorsal CA1 (dCA1) of the silicon probes coated with DiI. For each shank, the recording sites close the CA1 layer were used for LFP spectral analysis (see methods). (b) Distributions of LFP power spectra for WT and Lgdel/+ (mean ± SEM; number in parentheses indicate the numbers of mouse/shank pairs). (c) Photographs showing the AAV-mediated expression of hM3Gq-tdTomato in dCA1 PVI from two different mice (experiment was repeated independently 12 times with similar results, see Figs. 6,7 for effects on physiology and behavior).
Supplementary Figure 8 Power spectral densities of local field potentials (LFP) in the delta and gamma frequency ranges.
(a,b) Distribution of the LFP PSDs in the delta and gamma frequency ranges for WT (+/+) and Lgdel/+ mice (two-sided Mann-Whitney test; mean ± SEM; number in parentheses indicate the number of shank/mouse pairs). Data from Lgdel/+ and Pvalbcre/+;Lgdel/+ during the baseline were grouped and referred as Lgdel/+ control mice (all ctrl Lgdel/+). (c) PSDs in the gamma frequency range in WT and Lgdel/+ mice in the control conditions or after treatments with NRG1P or CNO (two-sided Wilcoxon paired test; mean ± SEM; number in parentheses indicate the number of shank/mouse pairs).
Supplementary Figure 9 Effect of increasing PVI excitability on mouse behaviour.
(a) Averaged startle response to a 120 dB tone in two representative mice (a.u. arbitrary unit). (b) Summary graph of the startle response. Statistical comparisons were done with two-sided Mann-Whitney test except for the chemogenetic manipulation where we used a two-sided Wilcoxon paired test. (c) Summary graph of the pre-pulse inhibition of the acoustic startle reflex. Statistical comparisons were done with two-way repeated measures ANOVA (number of animals indicated in parentheses). Data presented as box plots (box: 25-75th percentiles, whiskers: 10-90th percentiles, white bar: median). (d) Mice freezing behaviour in the different conditions during the fear conditioning (two-sided Mann-Whitney or two-sided Wilcoxon paired test for genotype comparison or drug treatments, respectively). All data are presented as mean ± SEM. Each circle represents a mouse (total number of animals indicated in parentheses).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Supplementary Table 1
Statistics summary table. Details about all statistical tests presented in the study.
Supplementary Video 1
Population dynamics in a WT slice. Supplementary video showing the neuronal co-activations during calcium imaging of a hippocampal slice culture (treated with Carbachol) originating from a WT mouse. (experiment was repeated 14 times independently with similar results, see Fig. 1e-h for quantifications).
Supplementary Video 2
Population dynamics in a Lgdel/+ slice. Supplementary video showing the neuronal co-activations during calcium imaging of a hippocampal slice culture (treated with Carbachol) originating from a Lgdel/+ mouse (experiment was repeated 22 times independently with similar results, see Fig. 1e-h for quantifications).
Rights and permissions
About this article
Cite this article
Marissal, T., Salazar, R.F., Bertollini, C. et al. Restoring wild-type-like CA1 network dynamics and behavior during adulthood in a mouse model of schizophrenia. Nat Neurosci 21, 1412–1420 (2018). https://doi.org/10.1038/s41593-018-0225-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0225-y
This article is cited by
-
Control of sustained attention and impulsivity by Gq-protein signalling in parvalbumin interneurons of the anterior cingulate cortex
Translational Psychiatry (2023)
-
Transcriptional linkage analysis with in vivo AAV-Perturb-seq
Nature (2023)
-
Pharmacogenetic activation of parvalbumin interneurons in the prefrontal cortex rescues cognitive deficits induced by adolescent MK801 administration
Neuropsychopharmacology (2023)
-
Reduced expression of the psychiatric risk gene DLG2 (PSD93) impairs hippocampal synaptic integration and plasticity
Neuropsychopharmacology (2022)
-
Pathological oligodendrocyte precursor cells revealed in human schizophrenic brains and trigger schizophrenia-like behaviors and synaptic defects in genetic animal model
Molecular Psychiatry (2022)