Abstract
Genome-wide association studies have identified 108 schizophrenia risk loci, but biological mechanisms for individual loci are largely unknown. Using developmental, genetic and illness-based RNA sequencing expression analysis in human brain, we characterized the human brain transcriptome around these loci and found enrichment for developmentally regulated genes with novel examples of shifting isoform usage across pre- and postnatal life. We found widespread expression quantitative trait loci (eQTLs), including many with transcript specificity and previously unannotated sequence that were independently replicated. We leveraged this general eQTL database to show that 48.1% of risk variants for schizophrenia associate with nearby expression. We lastly found 237 genes significantly differentially expressed between patients and controls, which replicated in an independent dataset, implicated synaptic processes, and were strongly regulated in early development. These findings together offer genetics- and diagnosis-related targets for better modeling of schizophrenia risk. This resource is publicly available at http://eqtl.brainseq.org/phase1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Birnbaum, R. & Weinberger, D. R. Genetic insights into the neurodevelopmental origins of schizophrenia. Nat. Rev. Neurosci. 18, 727–740 (2017).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Jaffe, A. E. et al. qSVA framework for RNA quality correction in differential expression analysis. Proc. Natl Acad. Sci. USA 114, 7130–7135 (2017).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Nellore, A. et al. Human splicing diversity and the extent of unannotated splice junctions across human RNA-seq samples on the Sequence Read Archive. Genome Biol. 17, 266 (2016).
Collado-Torres, L. et al. Flexible expressed region analysis for RNA-seq with derfinder. Nucleic Acids Res. 45, e9 (2017).
Jaffe, A. E. et al. Developmental regulation of human cortex transcription and its clinical relevance at single base resolution. Nat. Neurosci. 18, 154–161 (2015).
Tan, W. et al. Molecular cloning of a brain-specific, developmentally regulated neuregulin 1 (NRG1) isoform and identification of a functional promoter variant associated with schizophrenia. J. Biol. Chem. 282, 24343–24351 (2007).
Kao, W. T. et al. Common genetic variation in Neuregulin 3 (NRG3) influences risk for schizophrenia and impacts NRG3 expression in human brain. Proc. Natl Acad. Sci. USA 107, 15619–15624 (2010).
Tao, R. et al. Expression of ZNF804A in human brain and alterations in schizophrenia, bipolar disorder, and major depressive disorder: a novel transcript fetally regulated by the psychosis risk variantrs1344706. JAMAPsychiatry 71, 1112–1120 (2014).
Hyde, T. M. et al. Expression of GABA signaling molecules KCC2, NKCC1, and GAD1 in cortical development and schizophrenia. J. Neurosci. 31, 11088–11095 (2011).
Birnbaum, R., Jaffe, A. E., Hyde, T. M., Kleinman, J. E. & Weinberger, D. R. Prenatal expression patterns of genes associated with neuropsychiatric disorders. Am. J. Psychiatry 171, 758–767 (2014).
Buchanan, R. W. & Carpenter, W. T. Domains of psychopathology: an approach to the reduction of heterogeneity in schizophrenia. J. Nerv. Ment. Dis. 182, 193–204 (1994).
Winterer, G. & Weinberger, D. R. Genes, dopamine and cortical signal-to-noise ratio in schizophrenia. Trends Neurosci. 27, 683–690 (2004).
Jaffe, A. E. et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat. Neurosci. 19, 40–47 (2016).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Jaffe, A. E. & Irizarry, R. A. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 15, R31 (2014).
Steijger, T. et al. Assessment of transcript reconstruction methods for RNA-seq. Nat. Methods 10, 1177–1184 (2013).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Weinberger, D. R. & Levitt, P. Neurodevelopmental origins of schizophrenia. in Schizophrenia 393–412 (Wiley-Blackwell, Hoboken, NJ, USA, 2011).
Uhlhaas, P. J. & Singer, W. Abnormal neural oscillations and synchrony in schizophrenia. Nat. Rev. Neurosci. 11, 100–113 (2010).
Tyner, C. et al. The UCSC Genome Browser database: 2017 update. Nucleic Acids Res. 45D1, D626–D634 (2017).
Lipska, B. K. et al. Critical factors in gene expression in postmortem human brain: Focus on studies in schizophrenia. Biol. Psychiatry 60, 650–658 (2006).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Ramu, A., Abbott, T. & Griffith, M. regtools v. 0.1.0. https://github.com/griffithlab/regtools (2016).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Frazee, A. C. et al. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 33, 243–246 (2015).
Scharpf, R. B., Irizarry, R. A., Ritchie, M. E., Carvalho, B. & Ruczinski, I. Using the R Package crlmm for Genotyping and Copy Number Estimation. J. Stat. Softw. 40, 1–32 (2011).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Delaneau, O., Coulonges, C. & Zagury, J. F. Shape-IT: new rapid and accurate algorithm for haplotype inference. BMC Bioinforma. 9, 540 (2008).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Pohl, A. & Beato, M. bwtool: a tool for bigWig files. Bioinformatics 30, 1618–1619 (2014).
BrainSpan. Atlas of the Developing Human Brain http://developinghumanbrain.org (2011).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 13, 86 (2012).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Buja, A. & Eyuboglu, N. Remarks on Parallel Analysis. Multivar. Behav. Res. 27, 509–540 (1992).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Stouffer, S.A.S. E.A., DeVinney, L.C., Star, S.A. & Williams, R.M. Jr. The American Soldier, Vol.1: Adjustment during ArmyLife (Princeton Univ. Press, Princeton, NJ, USA, 1949).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, e3 (2004).
Acknowledgements
We thank R. Zielke, R. D. Vigorito and R. M. Johnson of the National Institute of Child Health and Human Development Brain and Tissue Bank for Developmental Disorders at the University of Maryland for providing fetal, child and adolescent brain specimens. This work was supported by the funding from Lieber Institute for Brain Development and the Maltz Research Laboratories and partially supported by NIH R21MH109956 (A.E.J.) and Consejo Nacional de Ciencia y Tecnología México 351535 (L.C.-T.). The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health. Additional funds were provided by the NCI, NHGRI, NHLBI, NIDA, NIMH and NINDS. Donors were enrolled at Biospecimen Source Sites funded by NCI/SAIC-Frederick, Inc. (SAIC-F) subcontracts to the National Disease Research Interchange (10XS170), Roswell Park Cancer Institute (10XS171) and Science Care, Inc. (X10S172). The Laboratory, Data Analysis, and Coordinating Center (LDACC) was funded through a contract (HHSN268201000029C) to The Broad Institute, Inc. Biorepository operations were funded through an SAIC-F subcontract to the Van Andel Institute (10ST1035). Additional data repository and project management were provided by SAIC-F (HHSN261200800001E). The Brain Bank was supported by supplements to University of Miami grants DA006227 and DA033684 and to contract N01MH000028. Statistical methods development grants were made to the University of Geneva (MH090941 and MH101814), the University of Chicago (MH090951, MH090937, MH101820, MH101825), the University of North Carolina - Chapel Hill (MH090936 and MH101819), Harvard University (MH090948), Stanford University (MH101782), Washington University St Louis (MH101810) and the University of Pennsylvania (MH101822). The data used for the analyses described in this manuscript were obtained from dbGaP accession number phs000424.v6.p1 on October 6, 2015. Data were generated as part of the CommonMind Consortium supported by funding from Takeda Pharmaceuticals Company Limited, F. Hoffman-La Roche Ltd and NIH grants R01MH085542, R01MH093725, P50MH066392, P50MH080405, R01MH097276, R01-MH-075916, P50M096891, P50MH084053S1, R37MH057881, R37MH057881S1, HHSN271201300031C, AG02219, AG05138 and MH06692. Brain tissue for the study was obtained from the following brain bank collections: the Mount Sinai NIH Brain and Tissue Repository, the University of Pennsylvania Alzheimer’s Disease Core Center, the University of Pittsburgh NeuroBioBank and Brain and Tissue Repositories and the NIMH Human Brain Collection Core. CMC leadership: Pamela Sklar and Joseph Buxbaum (Icahn School of Medicine at Mount Sinai), Bernie Devlin and David Lewis (University of Pittsburgh), Raquel Gur and Chang-Gyu Hahn (University of Pennsylvania), Keisuke Hirai and Hiroyoshi Toyoshiba (Takeda Pharmaceuticals Company Limited), Enrico Domenici and Laurent Essioux (F. Hoffman-La Roche Ltd), Lara Mangravite and Mette Peters (Sage Bionetworks), and Thomas Lehner and Barbara Lipska (NIMH).
Members of the BrainSeq consortium include: Christian R. Schubert, Patricio O’Donnell, Jie Quan, Jens R. Wendland, Hualin S. Xi, Ashley R. Winslow, Enrico Domenici, Dheeraj Malhotra, Laurent Essioux, Tony Kam-Thong, David C. Airey, John N. Calley, David A. Collier, Hong Wang, Brian Eastwood, Philip Ebert, Yushi Liu, Laura Nisenbaum, Cara Ruble, James Scherschel, Ryan Matthew Smith, Hui-Rong Qian, Kalpana Merchant, Michael Didriksen, Mitsuyuki Matsumoto, Takeshi Saito, Nicholas J. Brandon, Alan J. Cross, Qi Wang, Husseini Manji, Hartmuth Kolb, Maura Furey, Wayne C. Drevets, Joo Heon Shin, Andrew E. Jaffe, Rujuta Narurkar, Yankai Jia, Richard E. Straub, Amy Deep-Soboslay, Thomas M. Hyde, Joel E. Kleinman and Daniel R. Weinberger.
Author information
Authors and Affiliations
Consortia
Contributions
A.E.J. performed primary data processing and analyses, and led the writing of the manuscript. R.E.S. contributed to data analysis and writing of the manuscript. J.H.S., R.T. and Y.G. performed RNA sequencing data generation (RNA extraction, library preparation and sequencing) and QC analyses. L.C.-T. and J.T.L. performed region-level data generation and assisted in data analysis and interpretation. Q.C. performed risk profile score analysis. T.K.-T., H.S.X., J.Q., C.C., B.J.M., A.J.C., N.J.B. and the BrainSeq Consortium provided feedback on the manuscript and contributed to data analyses and interpretations on eQTL analyses. W.S.U. created the user-friendly database of eQTLs. A.D.-S. obtained consent from and clinically characterized human brain donors. T.M.H. and J.E.K. obtained consent from donors, collected, characterized and dissected human brain tissue and contributed to the design of the study. D.R.W. designed and oversaw the research project and analysis of the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The following named authors have competing interests: T.K.-T. is employed by F. Hoffmann-La Roche; H.S.X. and J.Q. were employees of Pfizer Inc. at the time these studies were conducted; A.J.C. and N.J.B. were full time employees and shareholders in AstraZeneca at the time these studies were conducted. The following BrainSeq Consortium members have competing interests: C.R.S., P.O., J.Q., J.R.W., H.S.X. and A.R.W. are employees and stockholders of Pfizer. E.D., L.E. and T.K.-T. are employees and stockholders of F Hoffmann-La Roche, AG. D.C.A., J.N.C., D.A.C., H.W., B.E., P.E., Y.L., L.N., C.R., J.E.S., R.M.S. and H.-R.Q. are employees and stockholders of Eli Lilly and Company. K.M. is an employee of TransThera Consulting and an ex-employee of Eli Lilly and Company. M.D. is an employee and stockholder of H Lundbeck A/S. M.M. and T.S. are employees of Astellas Pharma. N.J.B., A.J.C. and Q.W. are employees and stockholders of AstraZeneca LP. H.M., H.K., M.F. and W.C.D. are employees of Janssen Research and Development, LLC, and of Johnson and Johnson, and stockholders of Johnson and Johnson. J.H.S., A.E.J., Y.J., R.E.S., A.D.-S., R.N., T.M.H., J.E.K. and D.R.W. are employees of the Lieber Institute for Brain Development, a nonprofit organization.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
ackn
Supplementary Figure 1
Study overview and cartoon describing the five different expression summarizations.
Supplementary Figure 2 Cartoon describing the four different splice junction annotation classes, relative to annotated exons.
(A) Annotated splice junctions map between two exons (dark blue rectangles) in a known transcript. (B) Exon-skipping splice junctions map to two annotated exons in different transcripts. (C) Alternative start/exon junctions map to only one annotated exon on either the 5’ or 3’ end. (D) Completely novel junction do not map to any known exon.
Supplementary Figure 3 RNA-based cellular profiles across age for fetal replicating and quiescent neurons, and adult neurons, oligodendrocytes oligodendrocyte precursor cells, astrocytes, endothelial and microglia cells.
RNA-based cellular profiles across age for fetal (A) replicating and (B) quiescent neurons, and adult (C) neurons, (D) oligodendrocytes (E) oligodendrocyte precursor cells (OPCs), (F) astrocytes, (G) endothelial and (H) microglia cells.
Supplementary Figure 4 Associating global components of gene expression with estimates of fetal replicating, fetal quiescent and combined fetal neuronal cell types.
Associating global components of gene expression with estimates of (A) fetal replicating, (B) fetal quiescent and (C) combined fetal neuronal cell types. N=320 independent samples/subjects.
Supplementary Figure 5 Venn diagram of developmentally regulated features mapped back to Ensembl Gene IDs by the five feature summarization methods.
DER: differentially expressed region.
Supplementary Figure 6 Comparing F-statistics from the full models of the age spline versus the eight cell composition proportion estimates using the same null model for sex and ethnicity.
N=320 independent samples/subjects.
Supplementary Figure 7 Example of CRTC2 containing a developmental isoform shift.
Example of CRTC2 (A) containing a developmental isoform shift. (B) Gene-level analysis shows no developmental regulation but at the junction-level (C) one splice junction significantly decreases in expression and (D) another splice junction significantly increases in expression over the lifespan. Exons in panels (E), (F), and (H) show some marginal increases in expression across the lifespan, but only the exon in (G) is unique to a single isoform and shows significant decreases in expression. N=320 independent samples/subjects; P-values (two-sided) are based on an F-test for an age spline (see Methods).
Supplementary Figure 8 Venn diagram of Ensembl Gene IDs that contain significant isoform shifts by the four feature summarization methods that allow for multiple features per gene.
DER: differentially expressed region.
Supplementary Figure 9
Discovery (LIBD) and replication (CMC) t-statistics for eQTLs identified in the DLPFC for the best SNP–feature pair for each feature across five feature summarization types.
Supplementary Figure 10 Assessing regional specificity of eQTLs in GTEx for the best SNP–feature pair for each feature across five feature summarization types.
(A) Extensive concordance of many eQTLs within discovery (LIBD) and GTEx Frontal Cortex samples – 59.9% of Bonferroni-significant eQTLs in the LIBD dataset were directionally consistent and marginally significant at p < 0.01 in the replication data. (B) These DLPFC-identified eQTLs showed significant meta-analysis T-statistics across the 13 brain regions in GTEx (N=62-115 subjects per brain region) – 81.1% of Bonferroni-significant eQTLs from LIBD were significant at meta-analysis p-value < 1e-5 in GTEx. (C) These DLPFC-identified eQTLs showed lack of regional specificity even within GTEx.
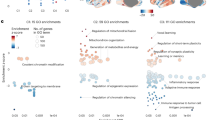
Supplementary Figure 11 Enrichment among chromatin states of 127 human cell types and tissues of unannotated sequences with expression levels associated with clinical risk variants in schizophrenia and the more general NHGRI GWAS catalog.
Enrichment among chromatin states of 127 human cell types and tissues of unannotated sequences with expression levels associated with clinical risk variants in A) schizophrenia and B) the more general NHGRI GWAS catalog. Points in red correspond to human brain tissues from different regions (N=13).
Supplementary Figure 12 Scatter plot of effect sizes (fold changes) in discovery and replication datasets for those features significant and replicated.
Colors have the same legend as Figure 3A.
Supplementary Figure 13 GWAS loci set-level analysis for all features together and then stratified by only genes, exons, junctions, transcripts and expressed regions.
GWAS loci set-level analysis for (A) all features together (N=1,011,825) and then stratified by only (B) genes (N=24,122), (C) exons (N=420,022), (D) junctions (N=229,846), (E) transcripts (N=61,950) and (F) expressed regions (N=275,885). P-values (two-sided) were based on linear regression Wald statistics, adjusting for mean expression levels.
Supplementary Figure 14
Replication assessment of isoform shift features in the BrainSpan project.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Notes 1–5
Supplementary Table 1
Demographic information for subjects in the present study, stratified by age and diagnosis group. Dx: diagnosis, N: sample size, F: Female, Cauc: Caucasian, SD: standard deviation, PCW: post-conception weeks. Antipsychotic use was measured using toxicology at time of death. P-values for diagnosis differences in continuous variables are based on linear regression and P-values for categorical variables are based on chi-squared tests.
Supplementary Table 2
Splice junction annotation and characterization in GTEx and GEUVADIS for any junction or highly expressed junctions (mean reads per 80 M mapped reads, RP80M > 0, > 1 and > 5). Each column represents a 2 × 2 table for presence of identified junctions in 495 DLPFC samples in two independent poly(A)+ datasets.
Supplementary Table 3
Summary statistics for those features significantly developmentally regulated in the control-only analyses across the lifespan.
Supplementary Table 4
Significant developmentally regulated features collapsed to Ensembl Gene ID, used to make Supplementary Figure 5.
Supplementary Table 5
Isoform shifts by Ensembl Gene ID and feature summarization type.
Supplementary Table 6
Gene set analyses for those genes with significant isoform shift, stratified by feature summarization type. Q-values, which control the false discovery rate, FDR, are shown. BP, biological processes; MF, molecular function.
Supplementary Table 7
Genes within the PGC schizophrenia GWAS risk regions that contain isoform shifts by feature summarization type. 21.8% of PGC2 genes had developmental isoform shifts using exon counts (N = 96/440) and 31.9% showed this isoform shift association based on junction counts (N = 137/430).
Supplementary Table 8
Significant eQTLs to schizophrenia GWAS index variants, including replication statistics and additional annotation metrics for variants and expressed features. “condIndep” column refers to those associations that were conditionally independent. Based on N = 412 independent samples/subjects and linear regression (two-sided P-values); see Methods.
Supplementary Table 9
Differential expression statistics for those features that were significant and replicated in case-control comparisons. Based on N = 351 independent samples/subjects and linear regression (two-sided P-values); see Methods.
Supplementary Table 10
Genes consistently differentially expressed by case-control analysis for the different feature summarizations.
Supplementary Table 11
Gene set analysis for genes with features differentially expressed by case-control status, stratified by directionality and feature summarization type. Based on N = 351 independent samples/subjects and linear regression (two-sided P-values) and 24,122 expressed genes; see Methods.
Supplementary Table 12
GWAS region set-level analyses for diagnosis-associated differentially expressed features, testing whether features in the PGC risk loci were more or less expressed as a set in cases compared to controls. Qual: qSVA adjusted analysis, Adj: observed covariate adjusted analysis. P-values (two-sided) were based on linear regression Wald statistics, adjusting for mean expression levels.
Supplementary Table 13
Associations between diagnosis, RPS and expression at gene and exon levels using linear regression. First two columns for each feature: P-values (two-sided) for gene set tests for the significant case-control features among statistics capturing the effect of RPS on expression. Second two columns for each feature: directionality between RPS on expression associations and diagnosis. N = 209 Caucasian samples; see Methods.
Supplementary Table 14
Reference-based deconvolution profiles from single-cell RNA-seq data.
Rights and permissions
About this article
Cite this article
Jaffe, A.E., Straub, R.E., Shin, J.H. et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat Neurosci 21, 1117–1125 (2018). https://doi.org/10.1038/s41593-018-0197-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0197-y
This article is cited by
-
Cortical gene expression architecture links healthy neurodevelopment to the imaging, transcriptomics and genetics of autism and schizophrenia
Nature Neuroscience (2024)
-
Risk factors associated with age at onset of Parkinson’s disease in the UK Biobank
npj Parkinson's Disease (2024)
-
Immunophenotypes in psychosis: is it a premature inflamm-aging disorder?
Molecular Psychiatry (2024)
-
Decreased CNNM2 expression in prefrontal cortex affects sensorimotor gating function, cognition, dendritic spine morphogenesis and risk of schizophrenia
Neuropsychopharmacology (2024)
-
Central neurocytoma exhibits radial glial cell signatures with FGFR3 hypomethylation and overexpression
Experimental & Molecular Medicine (2024)