Abstract
Though motor neurons selectively degenerate in amyotrophic lateral sclerosis, other cell types are likely involved in this disease. We recently generated rNLS8 mice in which human TDP-43 (hTDP-43) pathology could be reversibly induced in neurons and expected that microglia would contribute to neurodegeneration. However, only subtle microglial changes were detected during disease in the spinal cord, despite progressive motor neuron loss; microglia still reacted to inflammatory triggers in these mice. Notably, after hTDP-43 expression was suppressed, microglia dramatically proliferated and changed their morphology and gene expression profiles. These abundant, reactive microglia selectively cleared neuronal hTDP-43. Finally, when microgliosis was blocked during the early recovery phase using PLX3397, a CSF1R and c-kit inhibitor, rNLS8 mice failed to regain full motor function, revealing an important neuroprotective role for microglia. Therefore, reactive microglia exert neuroprotective functions in this amyotrophic lateral sclerosis model, and definition of the underlying mechanism could point toward novel therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Block, M. L., Zecca, L. & Hong, J. S. Microglia-mediated neurotoxicity: uncovering the molecular mechanisms. Nat. Rev. Neurosci. 8, 57–69 (2007).
Waisman, A., Ginhoux, F., Greter, M. & Bruttger, J. Homeostasis of microglia in the adult brain: review of novel microglia depletion systems. Trends Immunol. 36, 625–636 (2015).
Hong, S., Dissing-Olesen, L. & Stevens, B. New insights on the role of microglia in synaptic pruning in health and disease. Curr. Opin. Neurobiol. 36, 128–134 (2016).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Dougherty, K. D., Dreyfus, C. F. & Black, I. B. Brain-derived neurotrophic factor in astrocytes, oligodendrocytes, and microglia/macrophages after spinal cord injury. Neurobiol. Dis. 7 6 Pt B, 574–585 (2000).
Henkel, J. S. et al. Presence of dendritic cells, MCP-1, and activated microglia/macrophages in amyotrophic lateral sclerosis spinal cord tissue. Ann. Neurol. 55, 221–235 (2004).
Turner, M. R. et al. Evidence of widespread cerebral microglial activation in amyotrophic lateral sclerosis: an [11C](R)-PK11195 positron emission tomography study. Neurobiol. Dis. 15, 601–609 (2004).
Alshikho, M. J. et al. Glial activation colocalizes with structural abnormalities in amyotrophic lateral sclerosis. Neurology 87, 2554–2561 (2016).
Zürcher, N. R. et al. Increased in vivo glial activation in patients with amyotrophic lateral sclerosis: assessed with [(11)C]-PBR28. Neuroimage Clin. 7, 409–414 (2015).
Tateishi, T. et al. CSF chemokine alterations related to the clinical course of amyotrophic lateral sclerosis. J. Neuroimmunol. 222, 76–81 (2010).
Martínez, H. R. et al. Altered CSF cytokine network in amyotrophic lateral sclerosis patients: A pathway-based statistical analysis. Cytokine 90, 1–5 (2017).
Rabin, S. J. et al. Sporadic ALS has compartment-specific aberrant exon splicing and altered cell-matrix adhesion biology. Hum. Mol. Genet. 19, 313–328 (2010).
Offen, D. et al. Spinal cord mRNA profile in patients with ALS: comparison with transgenic mice expressing the human SOD-1 mutant. J. Mol. Neurosci. 38, 85–93 (2009).
Clement, A. M. et al. Wild-type nonneuronal cells extend survival of SOD1 mutant motor neurons in ALS mice. Science 302, 113–117 (2003).
Boillée, S. et al. Onset and progression in inherited ALS determined by motor neurons and microglia. Science 312, 1389–1392 (2006).
Beers, D. R. et al. Wild-type microglia extend survival in PU.1 knockout mice with familial amyotrophic lateral sclerosis. Proc. Natl Acad. Sci. USA 103, 16021–16026 (2006).
Martínez-Muriana, A. et al. CSF1R blockade slows the progression of amyotrophic lateral sclerosis by reducing microgliosis and invasion of macrophages into peripheral nerves. Sci. Rep. 6, 25663 (2016).
Frakes, A. E. et al. Microglia induce motor neuron death via the classical NF-κB pathway in amyotrophic lateral sclerosis. Neuron 81, 1009–1023 (2014).
Polazzi, E. et al. Copper-zinc superoxide dismutase (SOD1) is released by microglial cells and confers neuroprotection against 6-OHDA neurotoxicity. Neurosignals 21, 112–128 (2013).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Kabashi, E. et al. Gain and loss of function of ALS-related mutations of TARDBP (TDP-43) cause motor deficits in vivo. Hum. Mol. Genet. 19, 671–683 (2010).
Wegorzewska, I., Bell, S., Cairns, N. J., Miller, T. M. & Baloh, R. H. TDP-43 mutant transgenic mice develop features of ALS and frontotemporal lobar degeneration. Proc. Natl. Acad. Sci. USA 106, 18809–18814 (2009).
Herman, A. M., Khandelwal, P. J., Rebeck, G. W. & Moussa, C. E. Wild type TDP-43 induces neuro-inflammation and alters APP metabolism in lentiviral gene transfer models. Exp. Neurol. 235, 297–305 (2012).
Swarup, V. et al. Deregulation of TDP-43 in amyotrophic lateral sclerosis triggers nuclear factor κB-mediated pathogenic pathways. J. Exp. Med. 208, 2429–2447 (2011).
Tong, J. et al. Expression of ALS-linked TDP-43 mutant in astrocytes causes non-cell-autonomous motor neuron death in rats. EMBO J. 32, 1917–1926 (2013).
Walker, A. K. et al. Functional recovery in new mouse models of ALS/FTLD after clearance of pathological cytoplasmic TDP-43. Acta Neuropathol. 130, 643–660 (2015).
Spiller, K. J. et al. Selective motor neuron resistance and recovery in a new inducible mouse model of TDP-43 proteinopathy. J. Neurosci. 36, 7707–7717 (2016).
Spiller, K. J. et al. Progression of motor neuron disease is accelerated and the ability to recover is compromised with advanced age in rNLS8 mice. Acta Neuropathol. Commun. 4, 105 (2016).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Walker, A. K. et al. An insoluble frontotemporal lobar degeneration-associated TDP-43 C-terminal fragment causes neurodegeneration and hippocampus pathology in transgenic mice. Hum. Mol. Genet. 24, 7241–7254 (2015).
van Rooijen, N., Sanders, A. & van den Berg, T. K. Apoptosis of macrophages induced by liposome-mediated intracellular delivery of clodronate and propamidine. J. Immunol. Methods 193, 93–99 (1996).
Korin, B. et al. High-dimensional, single-cell characterization of the brain’s immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Chiu, I. M. et al. A neurodegeneration-specific gene-expression signature of acutely isolated microglia from an amyotrophic lateral sclerosis mouse model. Cell Rep. 4, 385–401 (2013).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e9 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Buratti, E. & Baralle, F. E. Multiple roles of TDP-43 in gene expression, splicing regulation, and human disease. Front. Biosci. 13, 867–878 (2008).
Elmore, M. R. et al. Colony-stimulating factor 1 receptor signaling is necessary for microglia viability, unmasking a microglia progenitor cell in the adult brain. Neuron 82, 380–397 (2014).
Wake, H., Moorhouse, A. J., Miyamoto, A. & Nabekura, J. Microglia: actively surveying and shaping neuronal circuit structure and function. Trends Neurosci. 36, 209–217 (2013).
Graeber, M. B. et al. The microglia/macrophage response in the neonatal rat facial nucleus following axotomy. Brain Res. 813, 241–253 (1998).
Schafer, D. P. & Stevens, B. Phagocytic glial cells: sculpting synaptic circuits in the developing nervous system. Curr. Opin. Neurobiol. 23, 1034–1040 (2013).
Damani, M. R. et al. Age-related alterations in the dynamic behavior of microglia. Aging Cell 10, 263–276 (2011).
Grabert, K. et al. Microglial brain region-dependent diversity and selective regional sensitivities to aging. Nat. Neurosci. 19, 504–516 (2016).
De Biase, L. M. et al. Local cues establish and maintain region-specific phenotypes of basal ganglia microglia. Neuron 95, 341–356.e6 (2017).
Tay, T. L. et al. A new fate mapping system reveals context-dependent random or clonal expansion of microglia. Nat. Neurosci. 20, 793–803 (2017).
Barbeito, A. G., Mesci, P. & Boillée, S. Motor neuron-immune interactions: the vicious circle of ALS. J. Neural Transm. (Vienna) 117, 981–1000 (2010).
Hu, J., Ge, H., Newman, M. & Liu, K. OSA: a fast and accurate alignment tool for RNA-Seq. Bioinformatics 28, 1933–1934 (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Hulsen, T., de Vlieg, J. & Alkema, W. BioVenn - a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomics 9, 488 (2008).
Brettschneider, J. et al. TDP-43 pathology and neuronal loss in amyotrophic lateral sclerosis spinal cord. Acta Neuropathol. 128, 423–437 (2014).
Acknowledgements
The authors thank S. Leight for administrative support, K.L. Spiller and K. Brunden for valuable feedback, V. Van Deerlin and E. Suh for the genetic analysis of the ALS patients studied here, E. Lee for help with microscopy, and D. Cotton-Samuel for assistance with tissue sectioning and immunostaining. This work was supported by the Judith and Jean Pape Adams Charitable Foundation and the ALS Association (K.J.S.), AG PO1-017586 (V.M.-Y.L.) as well as gifts from the Koller and Pottruck families. Finally, we thank our ALS patients and their families for making human ALS tissue samples available to us for our studies.
Author information
Authors and Affiliations
Contributions
K.J.S., R.M.R. and V.M.-Y.L. designed research; K.J.S., C.R.R., T.K., K.R.M., M.A.D., T.C.F. and R.G.C. performed research; K.J.S., T.K., M.A.D., T.C.F., R.G.C., C.R. and C.R.R. analyzed data; K.J.S., J.Q.T., R.M.R. and V.M.-Y.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
T.C.F., R.G.C. and C.J.R. are employees and stockholders of Biogen, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 When DOX is removed, there is neuron-specific hTDP-43 expression in rNLS8 mouse SC.
(a) Representative image of microglia (green) and NEFH (blue) in a nTg mouse shows that NEFH is expressed in neurons, but not microglia with orthogonal views to show the lack of IBA-1 and NEFH co-localization. (b) Representative image of microglia (green) and hTDP-43 (blue) from rNLS8 SC after 3 weeks of hTDP43ΔNLS expression, with orthogonal views to show the lack of hTDP-43 localization. Scale bars in (a,b) = 25 μm. Similar staining was observed in 3 independent animals per genotype. (c) Representative images from 1 of 4 rNLS8 mice prior to DOX removal and after 1, 4, and 8 weeks off DOX stained with a marker for TDP-43 (red) that labels both mouse and hTDP-43. Note: nuclear clearance of TDP-43 in many neurons after DOX removal. Scale bar = 100 μm.
Supplementary Figure 2 rNLS8 microglia are capable of responding to inflammatory triggers.
(a-b) Representative cryosections of SC from a nTg mouse (a) and an rNLS8 mouse (b) stained with IBA-1 (red) to label microglia shows reactive gliosis on the ipsilateral (left) side 6 days after sciatic nerve crush similarly for both mice, demonstrating that rNLS8 microglia can appropriately react to inflammatory triggers. (c-f) Zoomed in images of an nTg and rNLS8 SC, left (c,e) and right (d,f) ventral horns show the neurons containing hTDP-43 (blue) and activated or resting microglia interspersed among them. Similar results were obtained from 4 rNLS8 and 4 nTg mice. (g) Dot plot shows the microglia density differences in the crushed side (black dots) versus the uncrushed side (blue dots), which are similar for both nTg and rNLS8 mice; group means indicated with a red line, n = 4 per group, ***, paired, two-tailed t-test with d.f. = 3, t = 13.8, p = 0.0008 and t = 12.8, p = 0.001, between crushed and uncrushed sides of rNLS8 and nTg mice, respectively. (h) Dot plot showing no difference in hTDP-43 expression in MNs on crushed (black) versus control (blue) side of the rNLS8 SC with group means indicated with a red line, n = 4. Scale bar = 100 μm. (i-k) rNLS8 and nTg have similar inflammatory responses to peripheral injection with LPS. Representative cryosections of SC from 1 of 4 nTg and 1 of 4 rNLS8 mice injected with PBS (i) or LPS (4 mg/kg, i.p.) and sacrificed 6 hours (j) or 18 hours (k) later, stained with IBA-1 (red), shows a slight, time-dependent activation of spinal microglia, regardless of hTDP43ΔNLS transgene expression. Scale bar = 100 μm.
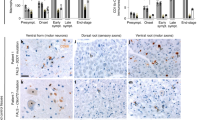
Supplementary Figure 3 A slight increase in IBA-1 immunoreactivity is found in postmortem brains of ALS patients with SOD1 mutations, compared to samples from control or sALS patients.
Representative images from cerebellum (a-c), hippocampus (d-f), and thalamus (g-i) of a control patient (a, d, g), a sporadic ALS patient (b, e, h), and a fALS patient with a SOD1 mutation (c,f,i) immunostained with IBA-1 (red) show slightly higher levels of reactive microgliosis in samples from patients with mutations in SOD1, but not in sporadic ALS or controls. Staining was done on brain regions from 4 control patients, 3 sporadic patients, and 3 patients with a SOD1 mutation with similar results. Scale bars = 100 μm.
Supplementary Figure 4 Microgliosis following transgene suppression occurs in cortex and hippocampus in addition to SC, though without a concurrent change in astrocytosis.
(a-c) Representative images of cortex, hippocampus, and lumbar SC of control rNLS8 mice (a), disease rNLS8 mice at 6 weeks off DOX (b), and early recovery rNLS8 mice (c) show a slight increase in the number of activated microglia (IBA-1, red) during recovery relative to controls in all regions, with changes in the SC being most pronounced. Similar results were observed in n = 3 animals per treatment group. (d) GFAP staining (green) on SC cryosections at multiple time-points during disease and recovery shows no obvious increase in astrocytosis at any time-point, relative to control rNLS8 mice maintained on DOX. Tissue was processed from n = 4 animals for disease time-points, and n = 5 animals for recovery time-points. All scale bars = 100 μm. (e-g) Consistent with this, we did not find significant changes in expression of genes associated with the A1 (Serping1 and Fkbp5) or A2 (Emp1) astrocyte phenotype by qPCR in RNA isolated from whole lysate of rNLS8 SC before (0 weeks off DOX, n = 4), during (4 weeks off DOX, n = 5), or after disease (6 weeks off DOX + 1 week on DOX, n = 6). Whiskers in box plots show minima and maxima, and box shows Q1-Q3 with median indicated, with individual points represented by black circles overlaid. Analysis by one way ANOVA, F2,14 = 0.63, p = 0.55 (e), F2,14 = 1.73, p = 0.22 (f), and F2,14 = 0.95, p = 0.42 (g).
Supplementary Figure 5 Expression of the C-terminal fragment of TDP-43 in NEFH-208 mice does not result in nuclear clearance or MN loss.
(a) Representative images from lumbar SCs of NEFH-208 mice immunostained with panTDP43 (purple) and hTDP-43 (green) show that the diffuse, punctate expression of hTDP-43 fragments in the cytoplasm of MNs does not cause nuclear clearance of mTDP-43. Similar staining results were seen in n = 6 animals. (b-c) No MN loss is observed in the ventral horns of NEFH-208 mice that had been off DOX for minimum 6 weeks. Bars represent mean ± S.D., n = 6 per group; unpaired, two-tailed t-test, t = 1.8, d.f. = 10, p = 0.10. All scale bars = 100 μm.
Supplementary Figure 6 rNLS8 animals have an intact blood—brain barrier at 4 weeks off DOX and clodronate liposomes do not cross into the CNS and alter microglia.
(a-d) Images of the CNS of 1 of 3 rNLS8 mice that were given tail vein injections with dye and sacrificed to detect brain penetrance of dye into the CNS from blood. Note that there is no evidence of blue dye grossly (a), in 2 mm coronal sections (b), or in 20 µm sections of hippocampus (c) or SC (d). (e-f) Representative cryosections of lumbar SC (e) and spleen (f) from 1 of 2 nTg mouse that was injected with clodronate liposomes immunostained with IBA-1 (red) alone or F4/80 (green), IBA-1 (red), and DAPI (blue) show intact microglia in the SC with a non-reactive morphology, but dramatically depleted splenic macrophage populations after clodronate-liposome treatment. Similar results were seen in n = 3 rNLS8 mice. Scale bar = 100 µm.
Supplementary Figure 7 Microglia proliferate in the rNLS8 SC after hTDP43ΔNLS transgene suppression.
(a-c) Double-labeling of BrdU (white) and microglia (IBA-1+, red) in the SC of rNLS8 mice during disease (a) or early recovery (b-c) following injection of BrdU at 50 mg/kg, i.p. daily for 7 days from 6 weeks off DOX shows an increase in the number of newborn microglia in recovery. Though most of the BrdU-labeled cells were microglia, some were other cell types and these are indicated with yellow stars. Similar staining results were observed in n = 4 mice. Scale bar = 50 μm. (d) Quantitative analysis of BrdU-positive cells showing there was a trend towards increased BrdU+ cells in the ventral horn during early recovery when compared to disease rNLS8 mice, with most of these being co-positive for IBA-1. Data are mean ± S.D., n = 4 per group; unpaired t-test, t = 2.3, d.f. = 6, p = 0.07.
Supplementary Figure 8 Transcriptional profiling of purified SC microglia.
(a) SC homogenate from rNLS8 and nTg littermates were subjected to FACS analysis and DRAQ5/CD45/CD11b+ microglia purification for RNA-seq. Shown here is the gating strategy and purity of sorted microglia. (b) Total RNA extracted from the following sorted samples: controls (green, pooled rNLS8 on DOX and nTg, n = 14); early disease (purple, rNLS8 2 weeks off DOX, n = 8); late disease (red, rNLS8 6 weeks off DOX, n = 7), and recovery (blue, rNLS8 6 weeks off + 1 week on DOX, n = 9). Bars represent mean ± S.D. (c) PCA of SC samples (n = 38) described in (b) shows some clustering of disease groups, though early and late disease groups are less discrete. (d) Cell lineage specific gene expression shows that all groups have high expression of microglial genes (Cx3cr1, Irf8), but low/absent markers for astrocytes (Gfap) and neurons (Nefh). Whiskers in box plots show minima and maxima, and box shows Q1-Q3 with median indicated, with individual points represented by squares overlaid. Group sizes are same as in (b). (e) Hierarchical clustering of all samples from SC based on logarithm of expression > 10 (7771 genes), shows some overlap between disease samples at different time-points. Yellow and blue indicate higher and lower expression levels, respectively.
Supplementary Figure 9 rNLS8 microglia change their gene expression profiles during late disease.
(a) Hierarchical clustering of SC samples based on FDR and Fold Change (FDR <0.05|FC| 1.5) shows gene expression changes in late disease (red) compared to control (green) samples. Yellow and blue indicate higher and lower expression levels, respectively. (b) Volcano plot of gene expression changes during after 6 weeks of hTDP43ΔNLS expression shows that 375 genes are upregulated (yellow) and 90 are downregulated (blue) between control and 6 weeks off DOX in rNLS8 SCs. (c) Table indicating the top activated canonical pathways identified by Ingenuity pathway analysis of the genes changed between late disease and control, with associated statistics reported by Ingenuity software using a right handed Fisher exact test (unadjusted p-value), their propriety “knowledge base” to calculate a z-score, and the genes included in these pathways indicated.
Supplementary Figure 10 rNLS8 microglia have a unique gene expression signature during recovery, which has not been previously identified in other models of neurodegeneration.
(a) Heatmap showing the levels of microglia-specific genes in rNLS8 samples that were identified as being up- or down-regulated in neurodegeneration previously by Chiu et al., 2013, Keren-Shaul et al., 2017, and Krasemann et al., 2017. Many of these genes were similarly upregulated (yellow) in both late disease (red) and early recovery (cyan) samples. (b) Heatmap showing the levels of microglia-specific genes in rNLS8 samples that were uniquely unregulated in recovery (FC |1.5|) and included in Fig. 6b. Hierarchical clustering with trees to show numerically accurate gene grouping are shown in both (a) and (b).
Supplementary Figure 11 Reactive microglia express CD68 and contain intracellular hTDP-43.
(a) Representative images of microglia (green), CD68 (red), hTDP-43 (orange), and DAPI (blue) in an rNLS8 SC after 6 weeks of hTDP43ΔNLS expression followed by 1 week suppression. The arrow points to a cluster of several microglia that are filled with hTDP-43 and highly express CD68. (b) Many IBA-1+ (red) cells in the lumbar SC also express CD68 (green) at this time. Scale bar = 100 μm. Similar results were seen in tissue from 4 other early recovery rNLS8 SCs. (c) There is a non-significant increase in the % area occupied by CD68 after 1 week of transgene suppression, relative to rNLS8 SC at 6 weeks off DOX (n = 4 rNLS8 mice at 6 weeks off; n = 5 in early recovery group; t = 2.0 d.f. = 7, p = 0.08). Bars represent mean ± S.D.
Supplementary Figure 12 During early recovery in rNLS8 mice, microglia often contain high levels of hTDP-43, but rarely other neuronal cytoplasmic proteins.
(a-c) Adjacent cryosections of rNLS8 lumbar SC double-stained for IBA-1 (red) and either hTDP-43 (a), NeuN (b), or ChAT (c) shows that the majority of these microglia contain hTDP-43 but not the other proteins that are highly expressed by MNs. (d-f) Adjacent sections showing a very activated cluster of microglia immmunostained for IBA-1 (red) and hTDP-43 (d), NeuN (e), and ChAT (f), shows that NeuN, but not ChAT, is occasionally found in these very activated clusters (labeled with #). Scale bar = 100 μm. Similar results were obtained on tissue from additional rNLS8 mice at 6 weeks off DOX + 1 week on (n = 3) and 2 weeks off DOX + 1 week on (n = 4).
Supplementary Figure 13 Neonatal intracerebroventricular injection with AAV9.GFP transduces MNs in the lumbar SC.
Lumbar SC cryosections from adult rNLS8 mice that were injected with AAV9.GFP at P1 immunostained with NeuN (a), IBA-1 (b) and GFAP (c) in blue, shows robust expression of GFP (green) in neurons (a), but not microglia (b) or astrocytes (c). Scale bar = 100 μm. Similar results were obtained on tissue from nTg mice (n = 4) and rNLS8 littermates (n = 3).
Supplementary Figure 14 PLX3397 treatment itself does not affect transgene expression, suppression or overall motor function.
(a) Though the hTDP-43 levels are different at the protein level, qPCR analysis reveals that the control and PLX3397-treated animals both have similar low hTDP-43 mRNA levels in rNLS8 mice maintained continuously on DOX (black bar, n = 3), off DOX for 6 weeks (white bar, n = 5), or 4 weeks off DOX + 2 weeks on and treated with PLX3397 (blue bar, n = 4) or control Nutella (grey bar, n = 4), Bars represent mean ± S.D. (b-e) Representative cryosections of lumbar SC from sham-treated (b,d) and PLX3397-treated (c,e) rNLS8 mice immunostained for IBA-1 (red) and LPL or Trem2 (blue) shows that surviving microglia in the PLX3397-treated lumbar SC still express LPL and Trem2. Staining was performed on tissue from n = 5 per treatment, with similar results. Scale bars = 100 μm (f-i). Treating nTg mice with the same PLX3397 regimen does not result in any obvious motor phenotype (f), change in maximum evoked CMAP from the gastrocnemius muscle (g), loss of MN in the SC (h), or axonal dieback from the TA muscle (i). Bars represent mean ± S.D., n = 4 mice.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14
Rights and permissions
About this article
Cite this article
Spiller, K.J., Restrepo, C.R., Khan, T. et al. Microglia-mediated recovery from ALS-relevant motor neuron degeneration in a mouse model of TDP-43 proteinopathy. Nat Neurosci 21, 329–340 (2018). https://doi.org/10.1038/s41593-018-0083-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0083-7
This article is cited by
-
Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases
Signal Transduction and Targeted Therapy (2024)
-
Dysregulated N6-methyladenosine modification in peripheral immune cells contributes to the pathogenesis of amyotrophic lateral sclerosis
Frontiers of Medicine (2024)
-
Molecular hallmarks of ageing in amyotrophic lateral sclerosis
Cellular and Molecular Life Sciences (2024)
-
Regulation of cortical hyperexcitability in amyotrophic lateral sclerosis: focusing on glial mechanisms
Molecular Neurodegeneration (2023)
-
Microglial CD68 and L-ferritin upregulation in response to phosphorylated-TDP-43 pathology in the amyotrophic lateral sclerosis brain
Acta Neuropathologica Communications (2023)