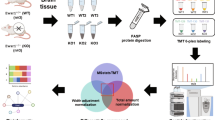
Abstract
Detailed observations of transcriptional, translational and post-translational events in the human brain are essential to improving our understanding of its development, function and vulnerability to disease. Here, we exploited label-free quantitative tandem mass-spectrometry to create an in-depth proteomic survey of regions of the postnatal human brain, ranging in age from early infancy to adulthood. Integration of protein data with existing matched whole-transcriptome sequencing (RNA-seq) from the BrainSpan project revealed varied patterns of protein–RNA relationships, with generally increased magnitudes of protein abundance differences between brain regions compared to RNA. Many of the differences amplified in protein data were reflective of cytoarchitectural and functional variation between brain regions. Comparing structurally similar cortical regions revealed significant differences in the abundances of receptor-associated and resident plasma membrane proteins that were not readily observed in the RNA expression data.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Silbereis, J. C., Pochareddy, S., Zhu, Y., Li, M. & Sestan, N. The cellular and molecular landscapes of the developing human central nervous system. Neuron 89, 248–268 (2016).
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015).
Swanson, L. W. & Lichtman, J. W. From Cajal to connectome and beyond. Annu. Rev. Neurosci. 39, 197–216 (2016).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Ouyang, A. et al. Spatial mapping of structural and connectional imaging data for the developing human brain with diffusion tensor imaging. Methods 73, 27–37 (2015).
Lerner, T. N., Ye, L. & Deisseroth, K. Communication in neural circuits: tools, opportunities, and challenges. Cell Rep. 164, 1136–1150 (2016).
Fertuzinhos, S. et al. Laminar and temporal expression dynamics of coding and noncoding RNAs in the mouse neocortex. Cell Rep. 6, 938–950 (2014).
Pletikos, M. et al. Temporal specification and bilaterality of human neocortical topographic gene expression. Neuron 81, 321–332 (2014).
Bakken, T. E. et al. A comprehensive transcriptional map of primate brain development. Nature 535, 367–375 (2016).
BrainSpan Consortium. Technical white paper: transcriptome profiling by RNA sequencing and exon microarray. http://help.brain-map.org/download/attachments/3506181/Transcriptome_Profiling.pdf?api=v2 (2013).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Krishnaswami, S. R. et al. Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat. Protoc. 11, 499–524 (2016).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Akbarian, S. et al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
Burgess, D. J. Technology: a drop in single-cell challenges. Nat. Rev. Genet. 16, 376–377 (2015).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Li, J. J., Bickel, P. J. & Biggin, M. D. System wide analyses have underestimated protein abundances and the importance of transcription in mammals. PeerJ 2, e270 (2014).
Schwanhäusser, B., Wolf, J., Selbach, M. & Busse, D. Synthesis and degradation jointly determine the responsiveness of the cellular proteome. Bioessays 35, 597–601 (2013).
Kitchen, R. R., Rozowsky, J. S., Gerstein, M. B. & Nairn, A. C. Decoding neuroproteomics: integrating the genome, translatome and functional anatomy. Nat. Neurosci. 17, 1491–1499 (2014).
Mann, M., Kulak, N. A., Nagaraj, N. & Cox, J. The coming age of complete, accurate, and ubiquitous proteomes. Mol. Cell 49, 583–590 (2013).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Beck, M. et al. The quantitative proteome of a human cell line. Mol. Syst. Biol. 7, 549 (2011).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Wang, H., Alvarez, S. & Hicks, L. M. Comprehensive comparison of iTRAQ and label-free LC-based quantitative proteomics approaches using two Chlamydomonas reinhardtii strains of interest for biofuels engineering. J. Proteome Res. 11, 487–501 (2012).
Latosinska, A. et al. Comparative analysis of label-free and 8-plex iTRAQ approach for quantitative tissue proteomic analysis. PLoS One 10, e0137048 (2015).
Sharma, K. et al. Cell type- and brain region-resolved mouse brain proteome. Nat. Neurosci. 18, 1819–1831 (2015).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Li, Z. et al. Systematic comparison of label-free, metabolic labeling, and isobaric chemical labeling for quantitative proteomics on LTQ Orbitrap Velos. J. Proteome Res. 11, 1582–1590 (2012).
Steiner, H. & Tseng, K. Y., eds. Handbook of Basal Ganglia Structure and Function (Academic, Cambridge, Massachusetts, USA, 2017).
Szklarczyk, D. et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43, D447–D452 (2015).
Volkow, N. D. & Morales, M. The brain on drugs: from reward to addiction. Cell 162, 712–725 (2015).
Tessier-Lavigne, M. & Goodman, C. S. The molecular biology of axon guidance. Science 274, 1123–1133 (1996).
Scheiffele, P. Cell-cell signaling during synapse formation in the CNS. Annu. Rev. Neurosci. 26, 485–508 (2003).
Lindner, M., Ng, J. K. M., Hochmeister, S., Meinl, E. & Linington, C. Neurofascin 186 specific autoantibodies induce axonal injury and exacerbate disease severity in experimental autoimmune encephalomyelitis. Exp. Neurol. 247, 259–266 (2013).
Mathey, E. K. et al. Neurofascin as a novel target for autoantibody-mediated axonal injury. J. Exp. Med. 204, 2363–2372 (2007).
Weder, N. et al. Child abuse, depression, and methylation in genes involved with stress, neural plasticity, and brain circuitry. J. Am. Acad. Child Adolesc. Psychiatry 53, 417–424.e5 (2014).
Montalvo-Ortiz, J. L. et al. The role of genes involved in stress, neural plasticity, and brain circuitry in depressive phenotypes: Convergent findings in a mouse model of neglect. Behav. Brain Res. 315, 71–74 (2016).
Kovács, G. G. et al. Natively unfolded tubulin polymerization promoting protein TPPP/p25 is a common marker of alpha-synucleinopathies. Neurobiol. Dis. 17, 155–162 (2004).
Hintiryan, H. et al. The mouse cortico-striatal projectome. Nat. Neurosci. 19, 1100–1114 (2016).
Seyfried, N. T. et al. Quantitative analysis of the detergent-insoluble brain proteome in frontotemporal lobar degeneration using SILAC internal standards. J. Proteome Res. 11, 2721–2738 (2012).
Llinas, R.R., Walton, K.D. & Lang, E.J. Cerebellum. in The Synaptic Organization of the Brain (ed. Shepherd, G. M.) 271–310, https://doi.org/10.1093/acprof:oso/9780195159561.003.0007 (2003).
Namjoshi, S. V. & Raab-Graham, K. F. Screening the molecular framework underlying local dendritic mRNA translation. Front. Mol. Neurosci. 10, 45 (2017).
Holt, C. E. & Schuman, E. M. The central dogma decentralized: new perspectives on RNA function and local translation in neurons. Neuron 80, 648–657 (2013).
Dammer, E. B. et al. Neuron enriched nuclear proteome isolated from human brain. J. Proteome Res. 12, 3193–3206 (2013).
Tagawa, K. et al. Comprehensive phosphoproteome analysis unravels the core signaling network that initiates the earliest synapse pathology in preclinical Alzheimer’s disease brain. Hum. Mol. Genet 24, 540–558 (2015).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Habegger, L. et al. RSEQtools: a modular framework to analyze RNA-Seq data using compact, anonymized data summaries. Bioinformatics 27, 281–283 (2011).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Langfelder, P., Zhang, B. & Horvath, S. Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics 24, 719–720 (2008).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing on JSTOR. J. R. Stat. Soc. B 57, 289–300 (1995).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Uhlen, M. et al. A proposal for validation of antibodies. Nat. Methods 13, 823–827 (2016).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44 (D1), D447–D456 (2016).
Acknowledgements
We thank S. Leslie and D. Li for discussions. Data were generated as part of the PsychENCODE Consortium, supported by U01MH103339, U01MH103365, U01MH103392, U01MH103340, U01MH103346, R01MH105472, R01MH094714, R01MH105898, R21MH102791, R21MH105881, R21MH103877 and P50MH106934 awarded to S. Akbarian (Icahn School of Medicine at Mount Sinai), G. Crawford (Duke University), S. Dracheva (Icahn School of Medicine at Mount Sinai), P. Farnham (University of Southern California), M.B.G. (Yale University), D. Geschwind (University of California, Los Angeles), T.M. Hyde (Lieber Institute for Brain Development), A. Jaffe (Lieber Institute for Brain Development), J.A. Knowles (University of Southern California), C. Liu (University of Illinois at Chicago), D. Pinto (Icahn School of Medicine at Mount Sinai), N.S. (Yale University), P. Sklar (Icahn School of Medicine at Mount Sinai), M. State (University of California, San Francisco), P. Sullivan (University of North Carolina), F. Vaccarino (Yale University), S. Weissman (Yale University), K. White (University of Chicago) and P. Zandi (Johns Hopkins University). This work was supported by the Yale/NIDA Neuroproteomics Centre (DA018343-12), by NIA grant AG047270-02, by NIMH grant MH110926, by NIH SIG grants 1S10OD019967-0 and 1S10ODOD018034-01, and by the State of Connecticut, Department of Mental Health & Addiction Services. B.C.C. was supported by a 2014 NARSAD Young Investigator Grant from the Brain & Behavior Research Foundation.
Author information
Authors and Affiliations
Contributions
B.C.C. designed the experiments, performed the experiments, analyzed the data and wrote the manuscript. R.R.K. designed the experiments, analyzed the data and wrote the manuscript. J.E.K. performed the experiments. E.Z.V. performed the experiments. M.P. contributed to tissue and sample processing. A.M.M.S. contributed to tissue and sample processing. T.T.L. designed the experiments and wrote the manuscript. M.B.G. contributed to RNA-seq data generation and provided computational resources. N.S. designed the experiments, contributed to tissue and sample processing, contributed to RNA-seq data generation and wrote the manuscript. A.C.N. designed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Number of proteins detected by highly fractionated proteomics across all regions
Histogram quantifies proteins detected directly by MS/MS (black) versus those detected by the “match between runs” feature (grey) in the fractionated regional samples
Supplementary Figure 2 Number of peptides detected per protein shows no clear inter-regional variability
The average number of peptides detected per protein was 9.2. For simplicity in these histograms, proteins with more than 10 peptides were set to have exactly 10 peptides. The distribution of peptides/protein is similar across all 7 fractionated brain regions
Supplementary Figure 3 Numbers of proteins detected in each single shot sample. The use of “match between runs” results in an approximately 50% increase in the number of proteins identified by single shot proteomics
Proteins directly detected by MS/MS are shown in black, whilst those identified by “match between runs” are in grey. Samples are shown in alphabetical order, and include the data from fractionated brain regions
Supplementary Figure 4 Batch effect identified by sample correlations is corrected by ComBat
A) Clustering of all single shot samples shows a clear batch effect. B) Clustering post correction by ComBat shows correction of the batch effect. C) Technical replicates of a single mixed region control were run evenly spaced throughout the LC-MS/MS runs. Correlation between these technical replicates is strongly improved as a result of batch correction
Supplementary Figure 5 Clustering all samples subjected to MS/MS using proteins significantly differentially expressed between brain regions revealed expected bulk differences between brain regions
This is a fully labelled zoomable version of the main Fig 2B
Supplementary Figure 6 Clustering all proteins significantly differentially expressed between regions reveals consistent patterns of expression that favour region-specific enrichment or region-specific depletion in abundance
Box and whisker plots show region specific patterns of protein expression across 33 gene clusters. The center line indicates the median, limits indicate the IQR, and the whiskers either 1.5* the IQR or the min/max value if it falls within 1.5* the IQR. Expression values from individual samples are shown as dots. Genes belonging to each cluster can be seen in Table 5B and Fig S7
Supplementary Figure 7 Clustering all proteins significantly differentially expressed between regions reveals consistent patterns of expression that favour region-specific enrichment or region-specific depletion in abundance
Heatmap shows expression levels of all differentially expressed genes across all single shot samples. The dendrogram above depicts the clusters of proteins that share patterns of expression across the regions. Cluster 0 is a group of proteins with no shared expression across the regions. Proteins belonging to this cluster are depicted with black arms on the dendrogram, and are not numbered below the heatmap. The heatmap shows that many clusters are dominated by expression changes in the cerebellum compared to other regions
Supplementary Figure 8 Striatally enriched clusters contain interacting proteins with roles in dopaminergic signalling and drug addiction
. A) Network diagram showing proteins from clusters 26 and 31. These clusters are strongly enriched for protein:protein interactions (adj. p value = 1.55E-15). Coloured edges represent different forms of interaction evidence; experimentally determined (pink), coexpressed (black), curated databases (blue) and text mining (green). The node size represents the extent to which the protein structure has been solved. B) Clusters 26 and 31 are significantly enriched for a number of Biological Process ontology terms. C) KEGG pathway analysis shows significant enrichment for expected pathways, including stimulant addiction and dopaminergic synapse
Supplementary Figure 9 Clustering all proteins significantly differentially expressed over developmental period reveals proteins enriched shortly after birth (period 8) and proteins more gently increasing or decreasing in abundance over the time-course
The center line indicates the median, limits indicate the IQR, and the whiskers either 1.5* the IQR or the min/max value if it falls within 1.5* the IQR. Individual samples are shown as dots on the box and whisker plots. Proteins belonging to each cluster can be seen in Table 5
Supplementary Figure 10a RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10b RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10c RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10d RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10e RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10f RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10g RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 10h RNA vs protein fold-change comparison of all pairs of brain regions
These scatterplots are identically defined to those in Fig 5A except that here genes are labelled to show individual gene names observable at high magnification. A) CBC vs all B) MD vs all C) STR vs all D) AMY vs all E) HIP vs all F) V1C vs all G) DFC vs all. Genes are coloured based on their agreement or disagreement between the RNA and protein measurements; genes for which the protein variability between regions is <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction but variable magnitude of change (≥2-fold) between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. H) Zoomable, gene labelled versions of the scatter plots in Figure 5c
Supplementary Figure 11 Summary quantifications of all RNA vs protein fold-change scatter plots
A) Per region-pair counts of the number of genes in each of the colour categories defined in Fig 5A; genes are coloured based on their agreement or disagreement between RNA and protein; genes for which the protein variability between regions was <2-fold of that reported at the RNA-level were considered consistent (green and grey points). Purple coloured genes are those with consistent direction, but variable magnitude, of change between the regions at the protein and RNA level, while red genes disagree in the direction of change between RNA and protein. Blue and orange genes vary between regions according to protein but not RNA and vice-versa. B) Comparison of absolute log2 fold-changes between each region and all others as reported by RNA (blue) and protein (red) show that in general the protein-level fold changes are greater. For example, the DFC plot shows the distribution, over all genes, of fold changes by RNA and by protein of the DFC with each of the 6 other regions. The center line indicates the median, limits indicate the IQR, and the whiskers either 1.5* the IQR or the min/max value if it falls within 1.5* the IQR. Outlying genes are not shown in this plot to highlight the increased median and 75th percentile of the protein fold-change distributions
Supplementary Figure 12 DFC/V1C comparison in detail
A) The scatterplot is identical to the corresponding panel in Fig 5A, except that individual gene names are labelled. Proteins used for validation in parts C & D are highlighted with pink font. B) Protein-protein interaction network, produced by STRING (medium stringency), of the genes upregulated in DFC in protein only (blue points, right hand side Fig S12A). Proteins used for validation in parts C & D are highlighted with pink font/circles. C) Immunoblotting of the 5 adult DFC and V1C samples shows enrichment of GRM2/3, CNR1 and PDE4D in the DFC over the V1C. Note that the blots shown in this figure are cropped images, and that CNR1 and NTRK3 labelling were performed on the same cut membrane, and thus have the same GAPDH control. D) Quantification of the immunoblots (values normalized to GAPDH) shows significant enrichment of GRM2/3, CNR1 & PDE4D in DFC over V1C by twotailed paired student’s T-test (n = 5 biological replicates per group, d.f. = 4, t = 5.842, 7.006, 3.067 respectively)
Supplementary Figure 13 Summary quantifications of all fold-change scatter plots for human vs mouse
Per region-pair counts of the number of genes in each of the colour categories defined in Fig 6B; genes are coloured based on their agreement or disagreement between mouse and human; genes for which the human variability between regions was within 2-fold of that reported for mouse were considered consistent (green and grey points). Purple coloured genes are those with consistent direction, but variable magnitude, of change between the regions in human vs mouse, while red genes disagree even in the direction of change between the species. Blue and orange genes vary between human regions but not mouse and vice-versa
Supplementary Figure 14a Human protein vs mouse protein fold-change comparison of all pairs of brain regions
The scatterplots are identically defined to those in Fig 6B except that here only significantly differentially expressed genes from the human are included and genes are labelled to show individual gene names. A) CBC vs all. B) MD vs all. C) STR vs all. D) HIP vs all. E) DFC vs all
Supplementary Figure 14b Human protein vs mouse protein fold-change comparison of all pairs of brain regions
The scatterplots are identically defined to those in Fig 6B except that here only significantly differentially expressed genes from the human are included and genes are labelled to show individual gene names. A) CBC vs all. B) MD vs all. C) STR vs all. D) HIP vs all. E) DFC vs all
Supplementary Figure 14c Human protein vs mouse protein fold-change comparison of all pairs of brain regions
The scatterplots are identically defined to those in Fig 6B except that here only significantly differentially expressed genes from the human are included and genes are labelled to show individual gene names. A) CBC vs all. B) MD vs all. C) STR vs all. D) HIP vs all. E) DFC vs all
Supplementary Figure 14d Human protein vs mouse protein fold-change comparison of all pairs of brain regions
The scatterplots are identically defined to those in Fig 6B except that here only significantly differentially expressed genes from the human are included and genes are labelled to show individual gene names. A) CBC vs all. B) MD vs all. C) STR vs all. D) HIP vs all. E) DFC vs all
Supplementary Figure 14e Human protein vs mouse protein fold-change comparison of all pairs of brain regions
The scatterplots are identically defined to those in Fig 6B except that here only significantly differentially expressed genes from the human are included and genes are labelled to show individual gene names. A) CBC vs all. B) MD vs all. C) STR vs all. D) HIP vs all. E) DFC vs all
Supplementary Figure 15 Un-cropped versions of the immunoblots in Figure S12
Note that all GADPH blots were visualised using Licor 800 antibodies, and only the 25 kDa ladder band is visible at this wavelength. A) mGlur2/3 and the corresponding GADPH blot. B) CNR1 and the corresponding GAPDH control. This blot was then trimmed and re-probed for NTRK3. C) PDE4D immunoblot and corresponding GAPDH control. D) NTRK3 immunoblot. See B) for GAPDH control
Supplementary Information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Table 11
Supplementary Table 1
Metadata for all 77 BrainSpan samples subjected to MS/MS for this study.
Supplementary Table 2
Peptide-level data obtained from heavily fractionated per-region MS/MS.
Supplementary Table 3
Protein-level summary of the fractionated per-region and single-shot MS/MS.
Supplementary Table 4
Label-free protein quantification (LFQ) of all single-shot samples.
Supplementary Table 5
Results of the proteomic spatiotemporal differential expression analysis.
Supplementary Table 6
Protein and RNA expression data for genes expressed in both datasets.
Supplementary Table 7
Inter-regional protein and RNA abundance and differential expression summary.
Supplementary Table 8
Summary of the RNA vs. protein differential consistency of each gene in accordance with the definitions introduced in Fig. 5.
Supplementary Table 9
Complete ontology and gene-set enrichment analysis results consistent with the definitions introduced in Fig. 5.
Supplementary Table 10
Inter-regional human and mouse protein abundance summary.
Supplementary Software
Supplementary Software
Rights and permissions
About this article
Cite this article
Carlyle, B.C., Kitchen, R.R., Kanyo, J.E. et al. A multiregional proteomic survey of the postnatal human brain. Nat Neurosci 20, 1787–1795 (2017). https://doi.org/10.1038/s41593-017-0011-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-017-0011-2
This article is cited by
-
ADNP dysregulates methylation and mitochondrial gene expression in the cerebellum of a Helsmoortel–Van der Aa syndrome autopsy case
Acta Neuropathologica Communications (2024)
-
Proteomics of mouse brain endothelium uncovers dysregulation of vesicular transport pathways during aging
Nature Aging (2024)
-
Remodeling of the postsynaptic proteome in male mice and marmosets during synapse development
Nature Communications (2024)
-
SMPD1 expression profile and mutation landscape help decipher genotype–phenotype association and precision diagnosis for acid sphingomyelinase deficiency
Hereditas (2023)
-
Spatiotemporal proteomic atlas of multiple brain regions across early fetal to neonatal stages in cynomolgus monkey
Nature Communications (2023)