Abstract
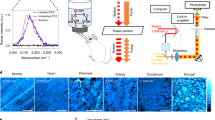
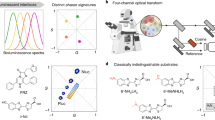
Simultaneous spatial mapping of the activity of multiple enzymes in a living system can elucidate their functions in health and disease. However, methods based on monitoring fluorescent substrates are limited. Here, we report the development of nitrile (C≡N)-tagged enzyme activity reporters, named nitrile chameleons, for the peak shift between substrate and product. To image these reporters in real time, we developed a laser-scanning mid-infrared photothermal imaging system capable of imaging the enzymatic substrates and products at a resolution of 300 nm. We show that when combined, these tools can map the activity distribution of different enzymes and measure their relative catalytic efficiency in living systems such as cancer cells, Caenorhabditis elegans, and brain tissues, and can be used to directly visualize caspase–phosphatase interactions during apoptosis. Our method is generally applicable to a broad category of enzymes and will enable new analyses of enzymes in their native context.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data related to the work are available in the article and supplementary information in this paper. The original data for the figures are available at https://doi.org/10.5281/zenodo.10085096.
References
Baruch, A., Jeffery, D. A. & Bogyo, M. Enzyme activity: it’s all about image. Trends Cell Biol. 14, 29–35 (2004).
Patel, T., Gores, G. J. & Kaufmann, S. H. The role of proteases during apoptosis. FASEB J. 10, 587–597 (1996).
Zhang, J. et al. Fluorogenic probes for disease-relevant enzymes. Chem. Soc. Rev. 48, 683–722 (2019).
Chyan, W. & Raines, R. T. Enzyme-activated fluorogenic probes for live-cell and in vivo imaging. ACS Chem. Biol. 13, 1810–1823 (2018).
Xing, B., Khanamiryan, A. & Rao, J. Cell-permeable near-infrared fluorogenic substrates for imaging β-lactamase activity. J. Am. Chem. Soc. 127, 4158–4159 (2005).
Grimm, J. B., Heckman, L. M. & Lavis, L. D. The chemistry of small-molecule fluorogenic probes. Prog. Mol. Biol. Transl. Sci. 113, 1–34 (2013).
Kwan, D. H. et al. Self-immobilizing fluorogenic imaging agents of enzyme activity. Angew. Chem. Int. Ed. Engl. 50, 300–303 (2011).
Johnson, D. S., Weerapana, E. & Cravatt, B. F. Strategies for discovering and derisking covalent, irreversible enzyme inhibitors. Future Med. Chem. 2, 949–964 (2010).
Noe, M. C. & Gilbert, A. M. Targeted covalent enzyme inhibitors. Annu. Rep. Med. Chem. 47, 413–439 (2012).
He, H. et al. Enzymatic noncovalent synthesis. Chem. Rev. 120, 9994–10078 (2020).
Zhou, J. et al. Enzyme-instructed self-assembly for spatiotemporal profiling of the activities of alkaline phosphatases on live cells. Chem 1, 246–263 (2016).
Liu, H. W. et al. In situ localization of enzyme activity in live cells by a molecular probe releasing a precipitating fluorochrome. Angew. Chem. Int. Ed. Engl. 56, 11788–11792 (2017).
Gao, Y. et al. Imaging self-assembly dependent spatial distribution of small molecules in a cellular environment. Langmuir 29, 15191–15200 (2013).
Zanetti-Domingues, L. C., Tynan, C. J., Rolfe, D. J., Clarke, D. T. & Martin-Fernandez, M. Hydrophobic fluorescent probes introduce artifacts into single molecule tracking experiments due to non-specific binding. PLoS One 8, e74200 (2013).
Bao, K. et al. Charge and hydrophobicity effects of NIR fluorophores on bone-specific imaging. Theranostics 5, 609–617 (2015).
Kim, B. J. & Xu, B. Enzyme-instructed self-assembly for cancer therapy and imaging. Bioconjug. Chem. 31, 492–500 (2020).
Razgulin, A., Ma, N. & Rao, J. Strategies for in vivo imaging of enzyme activity: an overview and recent advances. Chem. Soc. Rev. 40, 4186–4216 (2011).
Hamilton, B. R. et al. Mapping enzyme activity on tissue by functional mass spectrometry imaging. Angew. Chem. Int. Ed. Engl. 59, 3855–3858 (2020).
Xia, Q., Yin, J., Guo, Z. & Cheng, J.-X. Mid-infrared photothermal microscopy: principle, instrumentation, and applications. J. Phys. Chem. B 126, 8597–8613 (2022).
Bai, Y., Yin, J. & Cheng, J.-X. Bond-selective imaging by optically sensing the mid-infrared photothermal effect. Sci. Adv. 7, eabg1559 (2021).
Zhang, D. et al. Depth-resolved mid-infrared photothermal imaging of living cells and organisms with submicrometer spatial resolution. Sci. Adv. 2, e1600521 (2016).
Yin, J. et al. Video-rate mid-infrared photothermal imaging by single-pulse photothermal detection per pixel. Sci. Adv. 9, eadg8814 (2023).
Tai, F. et al. Detecting nitrile-containing small molecules by infrared photothermal microscopy. Analyst 146, 2307–2312 (2021).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Yin, J. et al. Nanosecond-resolution photothermal dynamic imaging via MHZ digitization and match filtering. Nat. Commun. 12, 7097 (2021).
Gao, Y., Shi, J., Yuan, D. & Xu, B. Imaging enzyme-triggered self-assembly of small molecules inside live cells. Nat. Commun. 3, 1033 (2012).
Yi, M. et al. Enzyme responsive rigid-rod aromatics target ‘undruggable’ phosphatases to kill cancer cells in a mimetic bone microenvironment. J. Am. Chem. Soc. 144, 13055–13059 (2022).
Wei, L. et al. Live-cell imaging of alkyne-tagged small biomolecules by stimulated Raman scattering. Nat. Methods 11, 410–412 (2014).
Wang, R. et al. Aggregation enhanced responsiveness of rationally designed probes to hydrogen sulfide for targeted cancer imaging. J. Am. Chem. Soc. 142, 15084–15090 (2020).
Hanczyc, P. & Fita, P. Laser emission of thioflavin T uncovers protein aggregation in amyloid nucleation phase. ACS Photonics 8, 2598–2609 (2021).
Zaguri, D. et al. Kinetic and thermodynamic driving factors in the assembly of phenylalanine-based modules. ACS Nano 15, 18305–18311 (2021).
Fried, S. D. & Boxer, S. G. Measuring electric fields and noncovalent interactions using the vibrational Stark effect. Acc. Chem. Res. 48, 998–1006 (2015).
Feng, Z. et al. Artificial intracellular filaments. Cell Rep. Phys. Sci. 1, 100085 (2020).
Qiu, C. et al. Advanced strategies for overcoming endosomal/lysosomal barrier in nanodrug delivery. Research 6, 0148 (2023).
He, H., Wang, H., Zhou, N., Yang, D. & Xu, B. Branched peptides for enzymatic supramolecular hydrogelation. Chem. Commun. 54, 86–89 (2018).
Guo, J. et al. The ratio of hydrogelator to precursor controls the enzymatic hydrogelation of a branched peptide. Soft Matter 16, 10101–10105 (2020).
He, H. et al. Dynamic continuum of nanoscale peptide assemblies facilitates endocytosis and endosomal escape. Nano Lett. 21, 4078–4085 (2021).
Sztiller‐Sikorska, M., Jakubowska, J., Wozniak, M., Stasiak, M. & Czyz, M. A non‐apoptotic function of caspase-3 in pharmacologically-induced differentiation of K562 cells. Br. J. Pharmacol. 157, 1451–1462 (2009).
Wilhelm, S., Wagner, H. & Häcker, G. Activation of caspase-3-like enzymes in non-apoptotic T cells. Eur. J. Immunol. 28, 891–900 (1998).
Carlile, G. W., Smith, D. H. & Wiedmann, M. Caspase-3 has a nonapoptotic function in erythroid maturation. Blood 103, 4310–4316 (2004).
Zhou, J., Du, X., Li, J., Yamagata, N. & Xu, B. Taurine boosts cellular uptake of small D-peptides for enzyme-instructed intracellular molecular self-assembly. J. Am. Chem. Soc. 137, 10040–10043 (2015).
Andrews, L. D., Zalatan, J. G. & Herschlag, D. Probing the origins of catalytic discrimination between phosphate and sulfate monoester hydrolysis: comparative analysis of alkaline phosphatase and protein tyrosine phosphatases. Biochemistry 53, 6811–6819 (2014).
Li, X. et al. Introducing D-amino acid or simple glycoside into small peptides to enable supramolecular hydrogelators to resist proteolysis. Langmuir 28, 13512–13517 (2012).
Torres, J. et al. Phosphorylation-regulated cleavage of the tumor suppressor PTEN by caspase-3: implications for the control of protein stability and PTEN–protein interactions. J. Biol. Chem. 278, 30652–30660 (2003).
Hallé, M. et al. Caspase-3 regulates catalytic activity and scaffolding functions of the protein tyrosine phosphatase PEST, a novel modulator of the apoptotic response. Mol. Cell. Biol. 27, 1172–1190 (2007).
Cai, Y. et al. Environment-sensitive fluorescent supramolecular nanofibers for imaging applications. Anal. Chem. 86, 2193–2199 (2014).
Yang, C., Chang, J., Gorospe, M. & Passaniti, A. Protein tyrosine phosphatase regulation of endothelial cell apoptosis and differentiation. Cell Growth Differ. 7, 161–172 (1996).
Halle, M., Tremblay, M. L. & Meng, T. C. Protein tyrosine phosphatases: emerging regulators of apoptosis. Cell Cycle 6, 2773–2781 (2007).
Sangwan, V. et al. Protein-tyrosine phosphatase 1B deficiency protects against Fas-induced hepatic failure. J. Biol. Chem. 281, 221–228 (2006).
Van Hoof, C. & Goris, J. Phosphatases in apoptosis: to be or not to be, PP2A is in the heart of the question. Biochim. Biophys. Acta 1640, 97–104 (2003).
Chen, M. J., Dixon, J. E. & Manning, G. Genomics and evolution of protein phosphatases. Sci. Signal. 10, eaag1796 (2017).
Brantley, S. J. et al. Discovery of small molecule inhibitors for the C. elegans caspase CED-3 by high-throughput screening. Biochem. Biophys. Res. Commun. 491, 773–779 (2017).
Stergiou, L., Doukoumetzidis, K., Sendoel, A. & Hengartner, M. The nucleotide excision repair pathway is required for UV-C-induced apoptosis in Caenorhabditis elegans. Cell Death Differ. 14, 1129–1138 (2007).
Fonta, C., Négyessy, L., Renaud, L. & Barone, P. Areal and subcellular localization of the ubiquitous alkaline phosphatase in the primate cerebral cortex: evidence for a role in neurotransmission. Cereb. Cortex 14, 595–609 (2004).
Bardet, P.-L. et al. A fluorescent reporter of caspase activity for live imaging. Proc. Natl Acad. Sci. USA 105, 13901–13905 (2008).
Ye, D. et al. Bioorthogonal cyclization-mediated in situ self-assembly of small-molecule probes for imaging caspase activity in vivo. Nat. Chem. 6, 519–526 (2014).
Van de Bittner, G. C., Bertozzi, C. R. & Chang, C. J. Strategy for dual-analyte luciferin imaging: in vivo bioluminescence detection of hydrogen peroxide and caspase activity in a murine model of acute inflammation. J. Am. Chem. Soc. 135, 1783–1795 (2013).
Ntziachristos, V., Tung, C.-H., Bremer, C. & Weissleder, R. Fluorescence molecular tomography resolves protease activity in vivo. Nat. Med. 8, 757–760 (2002).
Fujioka, H. et al. Multicolor activatable Raman probes for simultaneous detection of plural enzyme activities. J. Am. Chem. Soc. 142, 20701–20707 (2020).
Shi, L. et al. Mid-infrared metabolic imaging with vibrational probes. Nat. Methods 17, 844–851 (2020).
Acknowledgements
This work was supported by R35GM136223 and R33CA261726 to J.-X.C., R01CA142746 to B.X. and grant 2023-321163 from the Chan Zuckerberg Initiative Donor-Advised Fund at the Silicon Valley Community Foundation. Source data are provided with this paper.
Author information
Authors and Affiliations
Contributions
J.-X.C. and H.H. conceived the study. H.H. synthesized the probes, performed the experiments and drafted the paper. J.Y. developed the laser-scanning MIP microscope. J.Y. and M.L. helped with the MIP imaging. M.Y. and X.T. helped in the probe synthesis. M.Z. helped in the culturing and MIP imaging of C. elegans. C.V.P.D., Y.L. and Z.D. carried out the extraction of mice brains, tissue sectioning and tissue imaging. B.X. provided material support on synthesis and made intellectual contributions to experiment design.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Lingyan Shi, Stephen Weber and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Molecular synthesis and identification, Supplementary Figs. 1–21, Supplementary Text and Supplementary Schemes 1–3.
Supplementary Video 1
Live cell MIP imaging of the dynamic cellular movements at amide II.
Supplementary Video 2
3D reconstruction of phosphatase activity in C. elegans.
Supplementary Video 3
3D reconstruction of caspase activity in C. elegans.
Source data
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, H., Yin, J., Li, M. et al. Mapping enzyme activity in living systems by real-time mid-infrared photothermal imaging of nitrile chameleons. Nat Methods 21, 342–352 (2024). https://doi.org/10.1038/s41592-023-02137-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-02137-x