Abstract
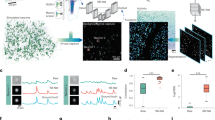
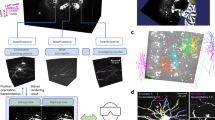
Reading out neuronal activity from three-dimensional (3D) functional imaging requires segmenting and tracking individual neurons. This is challenging in behaving animals if the brain moves and deforms. The traditional approach is to train a convolutional neural network with ground-truth (GT) annotations of images representing different brain postures. For 3D images, this is very labor intensive. We introduce ‘targeted augmentation’, a method to automatically synthesize artificial annotations from a few manual annotations. Our method (‘Targettrack’) learns the internal deformations of the brain to synthesize annotations for new postures by deforming GT annotations. This reduces the need for manual annotation and proofreading. A graphical user interface allows the application of the method end-to-end. We demonstrate Targettrack on recordings where neurons are labeled as key points or 3D volumes. Analyzing freely moving animals exposed to odor pulses, we uncover rich patterns in interneuron dynamics, including switching neuronal entrainment on and off.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data for key-point tracking are freely available for download and use at: https://drive.google.com/drive/folders/1_qVKLS09Cb4HuZeHzCG63ftxX1cgUimO. Videos illustrating key-point tracking are freely available for download and use at: https://drive.google.com/drive/folders/1uMUJu0Jed5HA1f3NbpZIAPDqPdxzl4su. Sample data for 3D volume tracking and videos are freely available for download and use at: https://drive.google.com/drive/folders/1-El9nexOvwNGAJw6uFFENGY1DqQ7tvxH. All data have additionally been deposited at: https://doi.org/10.5281/zenodo.10008744.
Code availability
The code is available under the MIT license at: https://github.com/rahi-lab/targettrack
References
Dupre, C. & Yuste, R. Non-overlapping neural networks in Hydra vulgaris. Curr. Biol. 27, 1085–1097 (2017).
Kato, S. et al. Global brain dynamics embed the motor command sequence of Caenorhabditis elegans. Cell 163, 656–669 (2015).
Lemon, W. C. et al. Whole-central nervous system functional imaging in larval Drosophila. Nat. Commun. 6, 7924 (2015).
Mann, K., Gallen, C. L. & Clandinin, T. R. Whole-brain calcium imaging reveals an intrinsic functional network in Drosophila. Curr. Biol. 27, 2389–2396 (2017).
Venkatachalam, V. et al. Pan-neuronal imaging in roaming Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 113, E1082–E1088 (2016).
Schrödel, T., Prevedel, R., Aumayr, K., Zimmer, M. & Vaziri, A. Brain-wide 3D imaging of neuronal activity in Caenorhabditis elegans with sculpted light. Nat. Methods 10, 1013–1020 (2013).
Nguyen, J. P. et al. Whole-brain calcium imaging with cellular resolution in freely behaving Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 113, E1074–E1081 (2016).
Prevedel, R. et al. Simultaneous whole-animal 3D imaging of neuronal activity using light-field microscopy. Nat. Methods 11, 727–730 (2014).
Voleti, V. et al. Real-time volumetric microscopy of in vivo dynamics and large-scale samples with SCAPE 2.0. Nat. Methods 16, 1054–1062 (2019).
Hallinen, K. M. et al. Decoding locomotion from population neural activity in moving C. elegans. eLife 10, e66135 (2021).
Susoy, V. et al. Natural sensory context drives diverse brain-wide activity during C. elegans mating. Cell 184, 5122–5137 (2021).
Marques, J. C., Li, M., Schaak, D., Robson, D. N. & Li, J. M. Internal state dynamics shape brainwide activity and foraging behaviour. Nature 577, 239–243 (2020).
Toyoshima, Y. et al. Accurate automatic detection of densely distributed cell nuclei in 3D space. PLoS Comput. Biol. 12, e1004970 (2016).
Ma, J. & Yuille, A. Nonrigid point set registration by preserving global and local structures. IEEE Trans. Image Proc. 25, 53–62 (2016).
Nguyen, J. P., Linder, A. N., Plummer, G. S., Shaevitz, J. W. & Leifer, A. M. Automatically tracking neurons in a moving and deforming brain. PLoS Comput. Biol. 13, e1005517 (2017).
Chaudhary, S., Lee, S. A., Li, Y., Patel, D. S. & Lu, H. Graphical-model framework for automated annotation of cell identities in dense cellular images. eLife 10, e60321 (2021).
Lagache, T., Hanson, A., Pérez-Ortega, J. E., Fairhall, A. & Yuste, R. Tracking calcium dynamics from individual neurons in behaving animals. PLoS Comput. Biol. 17, e1009432 (2021).
Wen, C. et al. 3DeeCellTracker, a deep learning-based pipeline for segmenting and tracking cells in 3D time lapse images. eLife 10, e59187 (2021).
Yu, X. et al. Fast deep neural correspondence for tracking and identifying neurons in C. elegans using semi-synthetic training. eLife 10, e66410 (2021).
Moen, E. et al. Deep learning for cellular image analysis. Nat. Methods 16, 1233–1246 (2019).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In Proc. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015 (Eds. Navab, N. et al.) 234–241 (Springer International Publishing, 2015).
Jian, B. & Vemuri, B. C. Robust point set registration using Gaussian mixture models. IEEE T. Pattern. Anal. 33, 1633–1645 (2011).
Chen, L.-C., Papandreou, G., Kokkinos, I., Murphy, K. & Yuille, A. L. Deeplab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Trans. Pattern Anal. Mach. Iintell. 40, 834–848 (2018).
Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. 3D U-Net: learning dense volumetric segmentation from sparse annotation. In Proc. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2016 (Eds. Ourselin, S. et al.) 424–432 (Springer International Publishing, 2016).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In Proc. 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 3431–3440 (IEEE, 2015).
Masci, J., Meier, U., Cireşan, D. & Schmidhuber, J. Stacked convolutional auto-encoders for hierarchical feature extraction. In Proc. International Conference on Artificial Neural Networks 52–59 (Springer, 2011).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Myronenko, A. & Song, X. Point set registration: coherent point drift. IEEE Trans. Pattern Anal. Mach. Intell. 32, 2262–2275 (2010).
Gatti, A. A. & Khallaghi, S. Pycpd: Pure numpy implementation of the coherent point drift algorithm. J. Open Source Softw. 7, 4681 (2022).
Alvarez, L., Sánchez, J. & Weickert, J. in Scale-Space Theories in Computer Vision (eds. Nielsen, M. et al.) 235–246 (Springer, 1999).
Zach, C., Pock, T. & Bischof, H. in Pattern Recognition (eds. Hamprecht, F. A. et al.) 214–223 (Springer, 2007).
Rahi, S. J. et al. Oscillatory stimuli differentiate adapting circuit topologies. Nat. Methods 14, 1010–1016 (2017).
Bouchard, M. B. et al. Swept confocally-aligned planar excitation (SCAPE) microscopy for high-speed volumetric imaging of behaving organisms. Nat. Photonics 9, 113–119 (2015).
Dietler, N. et al. A convolutional neural network segments yeast microscopy images with high accuracy. Nat. Commun. 11, 5723 (2020).
Acknowledgements
A.D., K.K., M.B.-K. and S.J.R. were supported by the École Polytechnique Fédérale de Lausanne (EPFL), a Helmut-Horten Foundation grant, the Swiss Data Science Center grant no. C20-12, SNSF grant no. CRSK-3_190526 and an EPFL Interdisciplinary Seed Fund grant awarded to S.J.R. C.F.P., V.S. and A.D.T.S. were supported by National Institutes of Health grant no. R01NS113119-01 awarded to A.D.T.S. We thank N. Greensmith and M. Minder for help developing the coarse volumetric tracking and the GUI, A. Lin for help constructing strains, M. Schmidt and A. Gross for help collecting and analyzing data and O. Peter for early tests of published methods.
Author information
Authors and Affiliations
Contributions
A.D.T.S., C.F.P., K.K., M.B.-K. and S.J.R. conceived the project. C.F.P., K.K., M.B.-K. and V.S. collected the data. C.F.P. developed the neural network, suggested targeted augmentation and implemented the method for key points. C.L.J. and M.B.-K. adapted the method for 3D volumes. C.F.P. and M.B.-K. ran the evaluations. C.F.P., M.B.-K. and A.D. developed the GUI. A.D.T.S., C.F.P., M.B.-K. and S.J.R. wrote the manuscript. A.D.T.S. and S.J.R. initiated and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Nina Vogt, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Results, Notes 1–4, Table 1, Figs. 1–10 and references.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Park, C.F., Barzegar-Keshteli, M., Korchagina, K. et al. Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation. Nat Methods 21, 142–149 (2024). https://doi.org/10.1038/s41592-023-02096-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-02096-3