Abstract
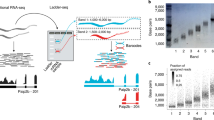
Most approaches to transcript quantification rely on fixed reference annotations; however, the transcriptome is dynamic and depending on the context, such static annotations contain inactive isoforms for some genes, whereas they are incomplete for others. Here we present Bambu, a method that performs machine-learning-based transcript discovery to enable quantification specific to the context of interest using long-read RNA-sequencing. To identify novel transcripts, Bambu estimates the novel discovery rate, which replaces arbitrary per-sample thresholds with a single, interpretable, precision-calibrated parameter. Bambu retains the full-length and unique read counts, enabling accurate quantification in presence of inactive isoforms. Compared to existing methods for transcript discovery, Bambu achieves greater precision without sacrificing sensitivity. We show that context-aware annotations improve quantification for both novel and known transcripts. We apply Bambu to quantify isoforms from repetitive HERVH-LTR7 retrotransposons in human embryonic stem cells, demonstrating the ability for context-specific transcript expression analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The SG-NEx samples are available through GitHub (https://github.com/GoekeLab/sg-nex-data), ENA (PRJEB44348) and AWS open data (https://registry.opendata.aws/sgnex/). Processed data associated with figures and tables are available on code ocean55. The PacBio data are available through SRA (SRP036136). The Arabidopsis data are available through ENA (PRJEB32782).
Code availability
Bambu is a R package for transcript discovery and quantification across multiple samples that is maintained on Bioconductor at https://www.bioconductor.org/packages/bambu/. The source code and a detailed documentation are available on GitHub at https://github.com/GoekeLab/bambu/. We used the BambuManuscriptRevision branch version of Bambu for analysis performed in this manuscript (https://github.com/GoekeLab/bambu/tree/BambuManuscriptRevision). All analysis code is available on Code Ocean55.
References
Matlin, A. J., Clark, F. & Smith, C. W. J. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Blencowe, B. J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Ben-Dov, C., Hartmann, B., Lundgren, J. & Valcárcel, J. Genome-wide analysis of alternative pre-mRNA splicing. J. Biol. Chem. 283, 1229–1233 (2008).
Graveley, B. R. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17, 100–107 (2001).
Nilsen, T. W. & Graveley, B. R. Expansion of the eukaryotic proteome by alternative splicing. Nature 463, 457–463 (2010).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Wang, D. et al. A deep proteome and transcriptome abundance atlas of 29 healthy human tissues. Mol. Syst. Biol. 15, e8503 (2019).
Gonzàlez-Porta, M., Frankish, A., Rung, J., Harrow, J. & Brazma, A. Transcriptome analysis of human tissues and cell lines reveals one dominant transcript per gene. Genome Biol. 14, R70 (2013).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Deschamps-Francoeur, G., Simoneau, J. & Scott, M. S. Handling multi-mapped reads in RNA-seq. Comput. Struct. Biotechnol. J. 18, 1569–1576 (2020).
Sarkar, H., Srivastava, A., Bravo, H. C., Love, M. I. & Patro, R. Terminus enables the discovery of data-driven, robust transcript groups from RNA-seq data. Bioinformatics 36, i102–i110 (2020).
Pardo-Palacios, F. et al. Systematic assessment of long-read RNA-seq methods for transcript identification and quantification. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-777702/v1 (2021).
Tang, A. D. et al. Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns. Nat. Commun. 11, 1438 (2020).
Wyman, D. et al. A technology-agnostic long-read analysis pipeline for transcriptome discovery and quantification. Preprint at bioRxiv https://doi.org/10.1101/672931 (2020).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278 (2019).
Prjibelski, A. D. et al. Accurate isoform discovery with IsoQuant using long reads. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01565-y (2023).
Soneson, C., Matthes, K. L., Nowicka, M., Law, C. W. & Robinson, M. D. Isoform prefiltering improves performance of count-based methods for analysis of differential transcript usage. Genome Biol. 17, 12 (2016).
Workman, R. E. et al. Nanopore native RNA sequencing of a human poly(A) transcriptome. Nat. Methods 16, 1297–1305 (2019).
Kuo, R. I. et al. Illuminating the dark side of the human transcriptome with long read transcript sequencing. BMC Genomics 21, 751 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Wick, R. R., Judd, L. M. & Holt, K. E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20, 129 (2019).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via theEMAlgorithm. J. R. Stat. Soc. 39, 1–22 (1977).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Eddelbuettel, D. et al. Rcpp: Seamless R and C++ integration. J. Stat. Softw. 40, 1–18 (2011).
Eddelbuettel, D. Seamless R and C++ Integration with Rcpp. (Springer, 2013).
R Core Team. R: a language and environment for statistical computing. (R Foundation for Statistical Computing, 2021).
Chen, T. & Guestrin, C. XGBoost: A Scalable Tree Boosting System. in Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (Association for Computing Machinery, 2016).
Hardwick, S. A. et al. Spliced synthetic genes as internal controls in RNA sequencing experiments. Nat. Methods 13, 792–798 (2016).
Chen, Y. et al. A systematic benchmark of Nanopore long read RNA sequencing for transcript level analysis in human cell lines. Preprint at bioRxiv https://doi.org/10.1101/2021.04.21.440736 (2021).
Pertea, G. & Pertea, M. GFF Utilities: GffRead and GffCompare. F1000Res. 9, 304 (2020).
Aken, B. L. et al. The Ensembl gene annotation system. Database 2016, baw093 (2016).
Parker, M. T. et al. Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification. eLlife 9, e49658 (2020).
Berardini, T. Z. et al. The Arabidopsis information resource: Making and mining the ‘gold standard’ annotated reference plant genome. Genesis 53, 474–485 (2015).
Tilgner, H., Grubert, F., Sharon, D. & Snyder, M. P. Defining a personal, allele-specific, and single-molecule long-read transcriptome. Proc. Natl Acad. Sci. USA 111, 9869–9874 (2014).
Gleeson, J. et al. Accurate expression quantification from nanopore direct RNA sequencing with NanoCount. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab1129 (2021).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Hu, Y. et al. LIQA: long-read isoform quantification and analysis. Genome Biol. 22, 182 (2021).
Tian, L. et al. Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing. Genome Biol. 22, 310 (2021).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 21, 423–425 (2014).
Kelley, D. & Rinn, J. Transposable elements reveal a stem cell-specific class of long noncoding RNAs. Genome Biol. 13, R107 (2012).
Göke, J. & Ng, H. H. CTRL+INSERT: retrotransposons and their contribution to regulation and innovation of the transcriptome. EMBO Rep. 17, 1131–1144 (2016).
Berrens, R. V. et al. Locus-specific expression of transposable elements in single cells with CELLO-seq. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01093-1 (2021).
Semenick, D. Tests and measurements: the t-test. J. Strength Cond. 12, 36 (1990).
Massey, F. J. The Kolmogorov–Smirnov test for goodness of fit. J. Am. Stat. Assoc. 46, 68–78 (1951).
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker. RepeatMasker http://repeatmasker.org (1996).
Soneson, C. et al. A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes. Nat. Commun. 10, 3359 (2019).
Troskie, R.-L. et al. Long-read cDNA sequencing identifies functional pseudogenes in the human transcriptome. Genome Biol. 22, 146 (2021).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Mulroney, L. et al. Identification of high confidence human poly(A) RNA isoform scaffolds using nanopore sequencing. RNA https://doi.org/10.1261/rna.078703.121 (2021).
Chen, Y., Sim, A., Lee, J., Goeke, J. Bambu (Source Code) https://codeocean.com/capsule/3893005/tree/v2 (2023).
Acknowledgements
This work was supported by funding from the Agency for Science, Technology and Research (A*STAR) and the National Medical Research Council. M.L. was supported by R01 HG009937.
Author information
Authors and Affiliations
Contributions
Y.C. and A.S. designed and implemented the computational method. J.G. conceived the project. Y.C., A.S. and J.G. designed the study and experiments and analyzed data. Y.K.W., K.Y., J.J.X.L. and M.H.L. contributed to the implementation of the computational method. M.L. contributed to the design of the computation method. Y.C., A.S. and J.G. organized and wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
J.G. received travel and accommodation expenses to speak at the Oxford Nanopore Community Meeting 2018. All other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Chung Chau HON and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling editors: Lei Tang and Lin Tang, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Text, Supplementary Text Figs. 1–5, Supplementary Text Tables 1–7 and Supplementary Notes.
Supplementary Table
Description of samples used for data analysis and annotation information on novel retrotransposon-derived transcripts based on Bambu.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Sim, A., Wan, Y.K. et al. Context-aware transcript quantification from long-read RNA-seq data with Bambu. Nat Methods 20, 1187–1195 (2023). https://doi.org/10.1038/s41592-023-01908-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-01908-w
This article is cited by
-
Isoform-specific RNA structure determination using Nano-DMS-MaP
Nature Protocols (2024)
-
Importance of pre-mRNA splicing and its study tools in plants
Advanced Biotechnology (2024)
-
SQANTI-SIM: a simulator of controlled transcript novelty for lrRNA-seq benchmark
Genome Biology (2023)
-
Benchmarking long-read RNA-sequencing analysis tools using in silico mixtures
Nature Methods (2023)