Abstract
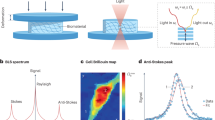
Brillouin microscopy is a technique for mechanical characterization of biological material without contact at high three-dimensional resolution. Here, we introduce dual line-scanning Brillouin microscopy (dLSBM), which improves acquisition speed and reduces irradiation dose by more than one order of magnitude with selective illumination and single-shot analysis of hundreds of points along the incident beam axis. Using tumor spheroids, we demonstrate the ability to capture the sample response to rapid mechanical perturbations as well as the spatially resolved evolution of the mechanical properties in growing spheroids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available in the paper and its Supplementary Notes 1 and 2. Source data are provided with this paper.
Change history
17 April 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41592-023-01877-0
References
Wang, N., Tytell, J. D. & Ingber, D. E. Mechanotransduction at a distance: mechanically coupling the extracellular matrix with the nucleus. Nat. Rev. Mol. Cell Biol. 10, 75–82 (2009).
Miller, C. J. & Davidson, L. A. The interplay between cell signalling and mechanics in developmental processes. Nat. Rev. Genet. 14, 733–744 (2013).
Bao, G. & Suresh, S. Cell and molecular mechanics of biological materials. Nat. Mater. 2, 715–725 (2003).
Kennedy, B. F., Wijesinghe, P. & Sampson, D. D. The emergence of optical elastography in biomedicine. Nature Photonics 11, 215–221 (2017).
Scarcelli, G. et al. Noncontact three-dimensional mapping of intracellular hydromechanical properties by Brillouin microscopy. Nat. Methods 12, 1132–1134 (2015).
Prevedel, R., Diz-Muñoz, A., Ruocco, G. & Antonacci, G. Brillouin microscopy: an emerging tool for mechanobiology. Nat. Methods 16, 969–977 (2019).
Zhang, J. & Scarcelli, G. Mapping mechanical properties of biological materials via an add-on Brillouin module to confocal microscopes. Nat. Protoc. 16, 1251–1275 (2021).
Gouveia, R. M. et al. Assessment of corneal substrate biomechanics and its effect on epithelial stem cell maintenance and differentiation. Nat. Commun. 10, 1496 (2019).
Margueritat, J. et al. High-frequency mechanical properties of tumors measured by Brillouin light scattering. Phys. Rev. Lett. 122, 018101 (2019).
Wisniewski, E. O. et al. Dorsoventral polarity directs cell responses to migration track geometries. Sci. Adv. 6, eaba6505 (2020).
Dil, J. G. Brillouin scattering in condensed matter. Reports on Progress in Physics 45, 285–334 (1982).
Antonacci, G. et al. Recent progress and current opinions in Brillouin microscopy for life science applications. Biophys. Rev. 12, 615–624 (2020).
Remer, I., Shaashoua, R., Shemesh, N., Ben-Zvi, A. & Bilenca, A. High-sensitivity and high-specificity biomechanical imaging by stimulated Brillouin scattering microscopy. Nat. Methods 17, 913–916 (2020).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Zhang, J., Fiore, A., Yun, S.-H., Kim, H. & Scarcelli, G. Line-scanning Brillouin microscopy for rapid non-invasive mechanical imaging. Sci. Rep. 6, 35398 (2016).
Tanner, K. & Gottesman, M. M. Beyond 3D culture models of cancer. Sci. Transl. Med. 7, 283ps289 (2015).
Han, Y. L. et al. Cell swelling, softening and invasion in a three-dimensional breast cancer model. Nat. Phys. 16, 101–108 (2020).
Zhou, E. et al. Universal behavior of the osmotically compressed cell and its analogy to the colloidal glass transition. Proc. Natl. Acad. Sci. USA 106, 10632–10637 (2009).
Scarcelli, G. & Yun, S. H. Reply to ‘Water content, not stiffness, dominates Brillouin spectroscopy measurements in hydrated materials’. Nat. Methods 15, 562–563 (2018).
Taubenberger, A. V. et al. 3D microenvironment stiffness regulates tumor spheroid growth and mechanics via p21 and ROCK. Adv. Biosyst. 3, 1900128 (2019).
Cross, S. E., Jin, Y.-S., Rao, J. & Gimzewski, J. K. Nanomechanical analysis of cells from cancer patients. Nat. Nanotechnol. 2, 780–783 (2007).
Mahajan, V. et al. Mapping tumor spheroid mechanics in dependence of 3D microenvironment stiffness and degradability by Brillouin microscopy. Cancers 13, 5549 (2021).
Schlüßler, R. et al. Correlative all-optical quantification of mass density and mechanics of subcellular compartments with fluorescence specificity. eLife 11, e68490 (2022).
Zhang, J., Nou, X., Kim, H. & Scarcelli, G. Brillouin flow cytometry for label-free mechanical phenotyping of the nucleus. Lab Chip 17, 663–670 (2017).
Lee, G. Y., Kenny, P. A., Lee, E. H. & Bissell, M. J. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat. Methods 4, 359–365 (2007).
Debnath, J., Muthuswamy, S. K. & Brugge, J. S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Schlüßler, R. et al. Mechanical mapping of spinal cord growth and repair in living zebrafish larvae by Brillouin imaging. Biophys. J. 115, 911–923 (2018).
Acknowledgements
The authors thank G. Zanini for help with the photodamage experiment and J. Xu and S. He for preparing the PDMS sphere. This work was supported by grants from the National Science Foundation (DBI-1942003, CMMI-1929412 to G.S.), the National Institutes of Health (R21CA258008 to G.S., R01EY028666 to G.S., R01HD095520 to G.S., K25HD097288 to J.Z.) and the Intramural Research Program of the National Cancer Institute (to K.T.), the American Cancer Society Institutional Research Grant (1816016) to J.Z. and a Wayne State University Research Grant to J.Z.
Author information
Authors and Affiliations
Contributions
J.Z. and G.S. conceived the project. J.Z., M.N., K.T. and G.S. devised the research plan. J.Z. developed the instrument and performed the experiments. M.N. developed the spheroid protocols and performed the photodamage experiment and AFM measurement. J.Z. and G.S. wrote the manuscript with input from all of the other authors.
Corresponding authors
Ethics declarations
Competing interests
J.Z., M.N and G.S. are inventors of patents related to the Brillouin technology. G.S. is a consultant for Intelon Optics. K.T. declares no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nina Vogt, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detailed schematic of the illumination beam.
a, M, mirror; L1-L2, spherical lens with focal length of 20 mm and 80 mm, respectively; Obj1-Obj2, objective lenses (4×/0.1NA). b, Measured laser spectra with and without Bragg filters. c, Extinction spectrum of the Fabry-Perot etalon used in the setup. The value of zero in b and c represents the wavelength and frequency of the laser light, respectively.
Extended Data Fig. 2 Dual-line illumination.
Brillouin image sections (left panels) and the averaged profile (right panels) along illumination direction of left-line illumination (a), right-line illumination (b), and combined dual-line illumination (c). The red arrow indicates the illumination direction of the beam line. Scale bar, 10 µm. The color bar represents the Brillouin shift with the unit of GHz. The center and error band represents the mean value + /− SD.
Extended Data Fig. 3 Brillouin shift gradient caused by the mismatch of refractive index.
a, Brillouin image of a PDMS sphere immersed in 1% agarose gel. Arrow indicates the propagation of illumination beam. Black dashed line indicates the location of interest for b. Scale bar is 10 µm. b, Shift profile across the PDMS sphere shows the gradient along illumination direction. c, Theoretical calculation model. ‘A’ represent an arbitrary point on the illumination axis, θ is the actual scattering angle collected by the spectrometer at 90° geometry, and α is the azimuthal angle of the sphere. d, The shift gradients from the calculation and the experiment.
Extended Data Fig. 4 Characterization of Brillouin spectrometer.
a, Exemplary raw Brillouin spectrum of DI water acquired by the spectrometer of the dLSBM setup with single-line illumination. b, Brillouin shift is extracted by fitting the spectrum of a single pixel with a Lorentzian function. Peaks represent the Stokes and anti-Stokes components of the Brillouin frequency. c, Representative histogram distribution of the estimated Brillouin shift of the same point after repeated acquisition (n = 300). The solid line is fitting result by Gaussian profile. The standard deviation of the estimated Brillouin shift is 8 MHz. d, Spectral precision of all points on illumination axis. The average is 9.8 MHz. e, Signal-to-noise (SNR) ratio against light energy of dLSBM spectrometer for water sample. The fitted line has a slope of 0.49.f, The measured spectrum of the laser. Circles are measured data, and solid line is Lorentzian fit. g, Spectral precision against light energy of the dLSBM spectrometer in water and spheroid sample. h, Comparison of dLSBM and confocal Brillouin microscopy (CBM) regarding the spectral precision against total light dose for 3D mapping of 100 × 200 × 10 pixels. water is used as sample. The data point of CBM is adapted from ref. 27, and the dash line indicates the shot noise limited operation.
Extended Data Fig. 5
Brillouin image reconstruction of the 3D mapping of a spheroid.
Extended Data Fig. 6 Characterization of the spatial resolution of the dLSBM setup.
a, Brillouin measurement across the interface of PDMS and water. b, Measurement of Rayleigh scattering from a 0.5 µm bead.
Extended Data Fig. 7 Comparison of dLSBM and CBM.
a-e, five independent spheroids measured by dLSBM (NA = 0.3) and CBM (NA = 0.4). Scale bar is 5 µm. For each spheroid, the Brillouin shifts of all pixels from the image was plotted into a histogram. The histogram was then fitted by a combination of two Gaussian distributions.
Extended Data Fig. 8 Analysis of subcellular mechanical information of spheroids.
a-d, Co-registered fluorescence and Brillouin images of four representative spheroids. Each sub figure shows the bright field image, fluorescence image of the nuclei, and the Brillouin image acquired by the CBM, respectively. Red line in the image outlines the profile of the spheroid. The plots of ‘H-peak 2’ is the right peak extracted from the curve fitting of the Brillouin shift histogram. The plots of ‘fluor. nucleus’ represents Brillouin shift of the nucleus region. e, Results of all the spheroid samples (n = 14). n.s.: not statistically significant (p = 0.6039). Statistical significance is determined by performing two-sided two-sample t-test, and no adjustment was made. In all boxplots, the central mark indicates the median, and the bottom and top edges indicate the 25th and 75th percentiles, respectively. The whiskers extend to the most extreme data points not considered outliers.
Extended Data Fig. 9 Effect of light illumination on the viability and growth rate of the spheroids.
a, Fluorescent images of the spheroids in control and illuminated groups. Live cells fluoresce in green (Calcein-AM) and dead cells fluoresce in red (EthD-1). b, Spheroid area against day of growth. The upper panel is the representative time-lapse image of a spheroid that is illuminated on Day 2. The points in the lower panel represent the area of spheroids over time. Black curves are best fit for exponential growth. Control group (n = 10), Illuminated group (n = 7).
Extended Data Fig. 10 Temporal change of the projection area of the spheroids under osmotic shock.
Hyperosmotic shock (n = 5), no shock (n = 6), and hypoosmotic shock (n = 5). Error bound indicates ± s.e.m.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2, Supplementary Figs. 1–3.
Supplementary Data 1
Source data for Supplementary Fig. 2.
Supplementary Data 2
Source data for Supplementary Fig. 3.
Supplementary Code 1
Matlab code for spectrum analysis.
Supplementary Code 2
Matlab code for image fusion.
Source data
Source Data Fig. 1
Source data for Fig. 1e,g,h.
Source Data Fig. 2
Source data for Fig. 2d,e,g–j.
Source Data Extended Data Fig. 1
Source data for Extended Data Fig.1b,c.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3b,d.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4b–h.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8e.
Source Data Extended Data Fig. 9
Source data for Extended Data Fig. 9b.
Source Data Extended Data Fig. 10
Source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J., Nikolic, M., Tanner, K. et al. Rapid biomechanical imaging at low irradiation level via dual line-scanning Brillouin microscopy. Nat Methods 20, 677–681 (2023). https://doi.org/10.1038/s41592-023-01816-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-01816-z
This article is cited by
-
Brillouin microscopy
Nature Reviews Methods Primers (2024)
-
Brillouin light scattering anisotropy microscopy for imaging the viscoelastic anisotropy in living cells
Nature Photonics (2024)
-
Mechanical properties of human tumour tissues and their implications for cancer development
Nature Reviews Physics (2024)
-
Label-free Brillouin endo-microscopy for the quantitative 3D imaging of sub-micrometre biology
Communications Biology (2024)
-
Mechanical anisotropy with Brillouin spectroscopy in one shot
Nature Photonics (2024)