Abstract
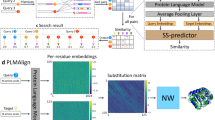
Protein sequence alignment is a key component of most bioinformatics pipelines to study the structures and functions of proteins. Aligning highly divergent sequences remains, however, a difficult task that current algorithms often fail to perform accurately, leaving many proteins or open reading frames poorly annotated. Here we leverage recent advances in deep learning for language modeling and differentiable programming to propose DEDAL (deep embedding and differentiable alignment), a flexible model to align protein sequences and detect homologs. DEDAL is a machine learning-based model that learns to align sequences by observing large datasets of raw protein sequences and of correct alignments. Once trained, we show that DEDAL improves by up to two- or threefold the alignment correctness over existing methods on remote homologs and better discriminates remote homologs from evolutionarily unrelated sequences, paving the way to improvements on many downstream tasks relying on sequence alignment in structural and functional genomics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The Uniref50 dataset is freely available under the Creative Commons Attribution (CC BY 4.0) License from https://www.uniprot.org. We used the 2018-03 release available at ftp.uniprot.org/pub/databases/uniprot/previous_major_releases/release-2018_03/uniref/. Pfam is freely available under the Creative Commons Zero (‘CC0’) licence from https://pfam.xfam.org. We used the Pfam-A seed dataset available at http://ftp.ebi.ac.uk/pub/databases/Pfam/releases/Pfam34.0/Pfam-A.seed.gz. Source data are provided with this paper.
Code availability
The code used in this study is freely available under an Apache v.2.0 license at https://github.com/google-research/google-research/tree/master/dedal. All analysis was carried out in Python v.3.6, with the following libraries: absl-py 0.7, gin-config 0.4, numpy 1.18.4, tensorflow 2.3, tensorflow_datasets 3.0, tensorflow_probability 0.1 and tf-models-official 2.6.
References
Prakash, T. & Taylor, T. D. Functional assignment of metagenomic data: challenges and applications. Brief Bioinform. 13, 711–727 (2012).
Lockless, S. W. & Ranganathan, R. Evolutionarily conserved pathways of energetic connectivity in protein families. Science 286, 295–299 (1999).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PloS ONE 6, e28766 (2011).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Smith, T. F. & Waterman, M. S. et al. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981).
Altschul, S., Gish, W., Miller, W., Myers, E. & Lipman, D. Basic local alignment search tools. J. Mol. Bol. 215, 403–410 (1990).
Pearson, W. R. Rapid and sensitive sequence comparisons with FASTP and FASTA. Meth. Enzymol. 183, 63–98 (1990).
Altschul, S. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Landan, G. & Graur, D. Characterization of pairwise and multiple sequence alignment errors. Gene 441, 141–147 (2009).
Lobb, B., Kurtz, D. A., Moreno-Hagelsieb, G. & Doxey, A. C. Remote homology and the functions of metagenomic dark matter. Front Genet. 6, 234 (2015).
Yu, C.-N. J., Joachims, T., Elber, R. & Pillardy, J. Support vector training of protein alignment models. J. Comput. Biol. 15, 867–880 (2008).
Fitch, W. M. & Smith, T. F. Optimal sequence alignments. Proc. Natl Acad. Sci. USA 80, 1382–1386 (1983).
Waterman, M. S., Eggert, M. & Lander, E. Parametric sequence comparisons. Proc. Natl Acad. Sci. USA 89, 6090–6093 (1992).
Gusfield, D., Balasubramanian, K. & Naor, D. Parametric optimization of sequence alignment. Algorithmica 12, 312–326 (1994).
Waterman, M. S. Parametric and ensemble sequence alignment algorithms. Bull. Math. Biol. 56, 743–767 (1994).
Vingron, M. & Waterman, M. S. Sequence alignment and penalty choice. review of concepts, case studies and implications. J. Mol. Biol. 235, 1–12 (1994).
Gusfield, D. & Stelling, P. Parametric and inverse-parametric sequence alignment with xparal. Methods Enzymol. 266, 481–494 (1996).
Pachter, L. & Sturmfels, B. Parametric inference for biological sequence analysis. Proc. Natl Acad. Sci. USA 101, 16138–16143 (2004).
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. USA 89, 10915–10919 (1992).
Keul, F., Hess, M., Goesele, M. & Hamacher, K. Pfasum: a substitution matrix from pfam structural alignments. BMC Bioinform. 18, 293 (2017).
Sun, F., Fernández-Baca, D. & Yu, W. Inverse Parametric Sequence Alignment. In Computing and Combinatorics (eds Ibarra, O. H. & Zhang, L.) 97–106 (Springer, 2002).
Saigo, H., Vert, J.-P. & Akutsu, T. Optimizing amino acid substitution matrices with a local alignment kernel. BMC Bioinform. 7, 246 (2006).
Kececioglu, J. & Kim, E. Simple and Fast Inverse Alignment. In Research in Computational Molecular Biology (eds Apostolico, A. et al.) 441–455 (Springer, 2006).
Bepler, T. & Berger, B. Learning protein sequence embeddings using information from structure. In 7th International Conference on Learning Representations (ICLR) (Openreview.net, 2019).
Morton, J. T. et al. Protein structural alignments from sequence. Preprint at bioRxiv https://doi.org/10.1101/2020.11.03.365932 (2020).
Petti, S. et al. End-to-end learning of multiple sequence alignments with differentiable Smith-Waterman. Bioinformatics https://doi.org/10.1093/bioinformatics/btac724 (2022).
Vaswani, A. et al. Attention is all you need. In Proc. of the 31st International Conference on Neural Information Processing Systems (eds Guyon, I. et al.), 5998–6008 (Curran Associates, Inc., 2017).
Suzek, B. E. et al. Uniref clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. In Proc. 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, NAACL-HLT 2019 Vol. 1 (Long and Short Papers) (eds Burstein, J. et al.) 4171–4186 (Association for Computational Linguistics, 2019).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl Acad. Sci. USA 118, e2016239118 (2021).
Abadi, M. et al. TensorFlow: large-scale machine learning on heterogeneous systems (tensorflow.org, 2015); https://www.tensorflow.org/
Müller, T. & Vingron, M. Modeling amino acid replacement. J. Comput. Biol. 7, 761–776 (2000).
Müller, T., Spang, R. & Vingron, M. Estimating amino acid substitution models: a comparison of dayhoff’s estimator, the resolvent approach and a maximum likelihood method. Mol. Biol. Evol. 19, 8–13 (2002).
Mensch, A. & Blondel, M. Differentiable dynamic programming for structured prediction and attention. In Proc. 35th International Conference on Machine Learning Vol. 80 (eds Dy, J. & Krause, A.) 3462–3471 (PMLR, 2018).
Berthet, Q. et al. Learning with differentiable perturbed optimizers. In Proc. of the 34th International Conference on Neural Information Processing Systems (eds Larochelle, H. et al.) 9508–9519 (Curran Associates, Inc., 2020).
Rao, R. et al. Evaluating protein transfer learning with TAPE. Adv. Neural Inf. Process. Syst. 32, 9689–9701 (2019).
The UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Steinegger, M. & Söding, J. Mmseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Collins, M. Discriminative training methods for hidden Markov models: theory and experiments with perceptron algorithms. In EMNLP ’02: Proc. ACL-02 Conference on Empirical Methods in Natural Language Processing Vol. 10, 1–8 (Association for Computational Linguistics, 2002).
Lafferty, J., McCallum, A. & Pereira, F. Conditional random fields: probabilistic models for segmenting and labeling sequence data. In Proc. 18th International Conf. on Machine Learning (eds Brodley, C. & Danyluk, A.) 282–289 (Morgan Kaufmann, 2001).
Blondel, M., Martins, A. F. & Niculae, V. Learning with Fenchel–Young losses. J. Mach. Learn. Res. 21, 1–69 (2020).
Karlin, S. & Altschul, S. F. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc. Natl Acad. Sci. USA 87, 2264–2268 (1990).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. In Proc. 3rd International Conference on Learning Representations, ICLR 2015, Conference Track Proceedings (eds Bengio, Y. & LeCun, Y.) (ICLR, 2015).
Raghava, G. P. & Barton, G. J. Quantifications of the variation in percentage identity for protein sequence alignments. BMC Bioinform. 7, 415 (2006).
Doolittle, R. F. Similar amino acid sequences: chance or common ancestry? Science 214, 149–159 (1981).
Acknowledgements
We thank D. Belanger, D. Dohan and A. Gane for preprocessing the data files and implementing the input pipelines for UniRef50 and M. Bileschi for providing feedback to improve the manuscript and guidance for navigating the Pfam database.
Author information
Authors and Affiliations
Contributions
F.L.-L. contributed to the development of the method, implemented most of the code, designed and ran most of the experiments and drafted the manuscript. Q.B. contributed to the development of the method, to the code and to the experiments. M.B. contributed to the development of the method and to the code. O.T. designed and implemented large parts of the code and contributed to the experiments. J.P.-V. designed the initial project, contributed to the method development and drafted the manuscript. All authors provided regular feedback to all aspects of the work, reviewed code and contributed to the writing of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
This study was funded by Google LLC. F.L.-L., Q.B., M.B., O.T. and J.-P.V. were employees of Google LLC and owned Alphabet stock as part of the standard compensation package during the making of this study.
Peer review
Peer review information
Nature Methods thanks Sean Eddy and Rama Ranganathan for their contribution to the peer review of this work. Primary Handling Editor: Arunima Singh, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Information.
Supplemental Data
Statistical Source Data for Supplementary Figs. 2, 3, 5, 6 and 8–10.
Source data
Source Data Fig. 2
Statistical Source Data for Fig. 2.
Source Data Fig. 4
Statistical Source Data for Fig. 4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Llinares-López, F., Berthet, Q., Blondel, M. et al. Deep embedding and alignment of protein sequences. Nat Methods 20, 104–111 (2023). https://doi.org/10.1038/s41592-022-01700-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01700-2
This article is cited by
-
Protein embedding based alignment
BMC Bioinformatics (2024)
-
PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
Nature Communications (2024)
-
Protein remote homology detection and structural alignment using deep learning
Nature Biotechnology (2023)