Abstract
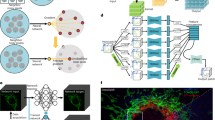
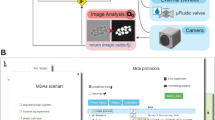
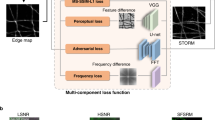
A common goal of fluorescence microscopy is to collect data on specific biological events. Yet, the event-specific content that can be collected from a sample is limited, especially for rare or stochastic processes. This is due in part to photobleaching and phototoxicity, which constrain imaging speed and duration. We developed an event-driven acquisition framework, in which neural-network-based recognition of specific biological events triggers real-time control in an instant structured illumination microscope. Our setup adapts acquisitions on-the-fly by switching between a slow imaging rate while detecting the onset of events, and a fast imaging rate during their progression. Thus, we capture mitochondrial and bacterial divisions at imaging rates that match their dynamic timescales, while extending overall imaging durations. Because event-driven acquisition allows the microscope to respond specifically to complex biological events, it acquires data enriched in relevant content.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data contained in this manuscript and the training data for the model used can be found at https://doi.org/10.5281/zenodo.5548354. Source data are provided with this paper.
Code availability
All code used in this project is available at https://github.com/LEB-EPFL/EDA. The Python plugin described can be found at https://github.com/wl-stepp/eda_plugin.
References
Laissue, P. P., Alghamdi, R. A., Tomancak, P., Reynaud, E. G. & Shroff, H. Assessing phototoxicity in live fluorescence imaging. Nat. Methods 14, 657–661 (2017).
Scherf, N. & Huisken, J. The smart and gentle microscope. Nat. Biotechnol. 33, 815–818 (2015).
Grimm, J. B. & Lavis, L. D. Caveat fluorophore: an insiders’ guide to small-molecule fluorescent labels. Nat. Methods 19, 149–158 (2022).
Rodriguez, E. A. et al. The growing and glowing toolbox of fluorescent and photoactive proteins. Trends Biochem. Sci. 42, 111–129 (2017).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Jin, L. et al. Deep learning enables structured illumination microscopy with low light levels and enhanced speed. Nat. Commun. 11, 1934 (2020).
Ouyang, W., Aristov, A., Lelek, M., Hao, X. & Zimmer, C. Deep learning massively accelerates super-resolution localization microscopy. Nat. Biotechnol. 36, 460–468 (2018).
Chen, J. et al. Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes. Nat. Methods 18, 678–687 (2021).
Hoebe, R. A. et al. Controlled light-exposure microscopy reduces photobleaching and phototoxicity in fluorescence live-cell imaging. Nat. Biotechnol. 25, 249–253 (2007).
Hoebe, R. A., Van der Voort, H. T. M., Stap, J., Van Noorden, C. J. F. & Manders, E. M. M. Quantitative determination of the reduction of phototoxicity and photobleaching by controlled light exposure microscopy. J. Microsc. 231, 9–20 (2008).
Heine, J. et al. Adaptive-illumination STED nanoscopy. Proc. Natl Acad. Sci. USA 114, 9797–9802 (2017).
Dreier, J. et al. Smart scanning for low-illumination and fast RESOLFT nanoscopy in vivo. Nat. Commun. 10, 556 (2019).
Chu, K. K., Lim, D. & Mertz, J. Enhanced weak-signal sensitivity in two-photon microscopy by adaptive illumination. Opt. Lett. 32, 2846–2848 (2007).
Li, B., Wu, C., Wang, M., Charan, K. & Xu, C. An adaptive excitation source for high-speed multiphoton microscopy. Nat. Methods 17, 163–166 (2020).
Pinkard, H. et al. Learned adaptive multiphoton illumination microscopy for large-scale immune response imaging. Nat. Commun. 12, 1916 (2021).
Chakrova, N., Canton, A. S., Danelon, C., Stallinga, S. & Rieger, B. Adaptive illumination reduces photobleaching in structured illumination microscopy. Biomed. Opt. Express 7, 4263 (2016).
Mahecic, D. et al. Mitochondrial membrane tension governs fission. Cell Rep. 35, 108947 (2021).
Lambert, A. et al. Constriction rate modulation can drive cell size control and homeostasis in C. crescentus. iScience 4, 180–189 (2018).
Zhou, X. et al. Mechanical crack propagation drives millisecond daughter cell separation in Staphylococcus aureus. Science 348, 574–578 (2015).
Edelstein, A. D. et al. Advanced methods of microscope control using μManager software. J. Biol. Methods 1, e10 (2014).
Pinkard, H., Stuurman, N., Corbin, K., Vale, R. & Krummel, M. F. Micro-Magellan: open-source, sample-adaptive, acquisition software for optical microscopy. Nat. Methods 13, 807–809 (2016).
Almada, P. et al. Automating multimodal microscopy with NanoJ-Fluidics. Nat. Commun. 10, 1223 (2019).
Yan, X. et al. High-content imaging-based pooled CRISPR screens in mammalian cells. J. Cell Biol. 220, e202008158 (2021).
York, A. G. et al. Instant super-resolution imaging in live cells and embryos via analog image processing. Nat. Methods 10, 1122–1126 (2013).
Mahecic, D. et al. Homogeneous multifocal excitation for high-throughput super-resolution imaging. Nat Methods 17, 726–733 (2020).
Twig, G. et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J 27, 433–446 (2008).
Sekh, A. A. et al. Physics-based machine learning for subcellular segmentation in living cells. Nat. Mach. Intell. 3, 1071–1080 (2021).
Fischer, C. A. et al. MitoSegNet: easy-to-use deep learning segmentation for analyzing mitochondrial morphology. iScience 23, 101601 (2020).
Lihavainen, E., Mäkelä, J., Spelbrink, J. N. & Ribeiro, A. S. Mytoe: automatic analysis of mitochondrial dynamics. Bioinformatics 28, 1050–1051 (2012).
Peng, J.-Y. et al. Automatic morphological subtyping reveals new roles of caspases in mitochondrial dynamics. PLoS Comput. Biol. 7, 14 (2011).
Valente, A. J., Maddalena, L. A., Robb, E. L., Moradi, F. & Stuart, J. A. A simple ImageJ macro tool for analyzing mitochondrial network morphology in mammalian cell culture. Acta Histochem. 119, 315–326 (2017).
Leonard, A. P. et al. Quantitative analysis of mitochondrial morphology and membrane potential in living cells using high-content imaging, machine learning, and morphological binning. Biochim. Biophys. Acta 1853, 348–360 (2015).
Lefebvre, A. E. Y. T., Ma, D., Kessenbrock, K., Lawson, D. A. & Digman, M. A. Automated segmentation and tracking of mitochondria in live-cell time-lapse images. Nat. Methods 18, 1091–1102 (2021).
Smirnova, E., Griparic, L., Shurland, D.-L. & van der Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Kamerkar, S. C., Kraus, F., Sharpe, A. J., Pucadyil, T. J. & Ryan, M. T. Dynamin-related protein 1 has membrane constricting and severing abilities sufficient for mitochondrial and peroxisomal fission. Nat. Commun 9, 5239 (2018).
Kleele, T. et al. Distinct fission signatures predict mitochondrial degradation or biogenesis. Nature 593, 435–439 (2021).
Ugarte-Uribe, B., Müller, H.-M., Otsuki, M., Nickel, W. & García-Sáez, A. J. Dynamin-related protein 1 (Drp1) promotes structural intermediates of membrane division. J. Biol. Chem. 289, 30645–30656 (2014).
Ronneberger, O., Fischer, P. & Brox, T. in U-Net: Convolutional Networks for Biomedical Image Segmentation. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015 (eds. Navab, N., Hornegger, J., et al.) 234–241 (Springer International Publishing, 2015).
Andersson, S. G. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–140 (1998).
Kilian, N. et al. Assessing photodamage in live-cell STED microscopy. Nat. Methods 15, 755–756 (2018).
Eisenstein, M. Smart solutions for automated imaging. Nat. Methods 17, 1075–1079 (2020).
Waithe, D. et al. Object detection networks and augmented reality for cellular detection in fluorescence microscopy. J. Cell. Biol 219, e201903166 (2020).
Kechkar, A., Nair, D., Heilemann, M., Choquet, D. & Sibarita, J.-B. Real-time analysis and visualization for single-molecule based super-resolution microscopy. PLoS ONE 8, e62918 (2013).
Royer, L. A. et al. Adaptive light-sheet microscopy for long-term, high-resolution imaging in living organisms. Nat. Biotechnol. 34, 1267–1278 (2016).
Štefko, M., Ottino, B., Douglass, K. M. & Manley, S. Autonomous illumination control for localization microscopy. Opt. Express 26, 30882 (2018).
Durand, A. et al. A machine learning approach for online automated optimization of super-resolution optical microscopy. Nat. Commun. 9, 5247 (2018).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017).
Cohen, A. E. & Moerner, W. E. Method for trapping and manipulating nanoscale objects in solution. Appl. Phys. Lett. 86, 093109 (2005).
Ely, B. Genetics of Caulobacter crescentus. Methods Enzymol. 204, 372–384 (1991).
Schrader, J. M. & Shapiro, L. Synchronization of Caulobacter crescentus for investigation of the bacterial cell cycle. J. Vis. Exp. 98, e52633 (2015).
Fiolka, R., Shao, L., Rego, E. H., Davidson, M. W. & Gustafsson, M. G. L. Time-lapse two-color 3D imaging of live cells with doubled resolution using structured illumination. Proc. Natl Acad. Sci USA 109, 5311–5315 (2012).
Czech, E., Aksoy, B. A., Aksoy, P. & Hammerbacher, J. Cytokit: a single-cell analysis toolkit for high dimensional fluorescent microscopy imaging. BMC Bioinformatics 20, 448 (2019).
Acknowledgements
We thank H. Perreten for technical support with cell culture and plasmid construction, and L. Casini and J. Collier (University of Lausanne) for sharing plasmids and protocols for the bacterial experiments. Imaging data used for training the neural network in this publication were produced in collaboration with the Advanced Imaging Center (AIC), a facility jointly supported by the Gordon and Betty Moore Foundation and HHMI at HHMI’s Janelia Research Campus. We thank L. Shao and T.-L. Chew at Janelia AIC for their help with SIM imaging. This work was supported by the Swiss National Science Foundation project grant (SNSF; 182429, to S.M., D.M. and W.L.S.), and National Centre for Competence in Research (NCCR Chemical Biology, to S.M. and D.M.); and the European Union’s H2020 program under the European Research Council (ERC; CoG 819823 Piko, to S.M. and C.Z.), and the Marie Skłodowska-Curie Fellowships (890169 BALTIC, to J.G.). M.W. is supported by the EPFL School of Life Sciences and a generous foundation represented by CARIGEST SA.
Author information
Authors and Affiliations
Contributions
D.M., J.G. and S.M. conceived and designed the project. D.M., M.W. and S.M. supervised the project. D.M. collected the training data and performed the experiments on mitochondrial division. D.M. and M.W. implemented the neural network for event detection. W.L.S. and D.M. implemented the EDA framework and performed data analysis. W.L.S. developed the Python plugin for Micro-Manager and its documentation. W.L.S performed the experiments on C. crescentus. C.Z. cultivated the C. crescentus strains and prepared samples for imaging. W.L.S prepared the figures. D.M., W.L.S. and S.M. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Example maximum event score regions which triggered EDA.
The event score output by the neural network can also be used to extract events of high interest from the datasets, after the acquisition is complete. Here, events that triggered EDA in different datasets are shown. The highest event score was used to define a region of interest around the event, representing a time and location of highest interest in the sample. Some regions appear twice, when the neural network event score was high enough to trigger EDA multiple times. Frames are shown in no specific order. Scale bars: 1 μm.
Extended Data Fig. 2 Bleaching behavior of a mitochondria sample during EDA imaging.
The different modes of imaging can clearly be seen in the bleaching curve represented by the signal-to-noise ratio calculated from the intensity inside the mitochondria compared to the signal outside of the mitochondria. For some parts with low frame rate, even a slight recovery of signal can be observed. (representative of n = 4 independent experiments).
Extended Data Fig. 3 EDA delivers additional frames during events of interest.
Top row: mitochondrial division as it would have been recorded with the slow fixed imaging rate without EDA. Vertical frames: additional frames captured thanks to EDA switching to the fast imaging speed showing more detail of the dynamics of the event. Both the final constriction state and the fade of the DRP1 peak can be observed with higher temporal resolution, enhancing the relevant content of the dataset. This division event can also be seen in Supplementary Video 3.0. (Scale bars: 1 μm, representative of N = 33 events in n = 4 independent experiments).
Extended Data Fig. 4 EDA imaging of synchronized bacteria populations.
C. crescentus, the strain used in this study, were synchronized via density centrifugation to obtain a population of cells that are all at the beginning of their cell cycle (G0, swarmer). This leads to a time lag before the next divisions take place. As they are synchronized, many bacteria in the sample will then divide at the same time. We used EDA to sense the onset of divisions in the sample and increase imaging speed during the divisions for high SNR and temporal resolution. We tested different times between images for fast and slow speeds, as well as different threshold event scores (gray band). a, slow: 9 min, fast 3 min. b and c, slow: 12 min, fast 2 min. (Scale bar: 1 μm, n = 4 independent synchronizations).
Supplementary information
Supplementary Information
Supplementary Notes 1–5.
Supplementary Video 1.0
Real-time video of a 300 × 300 px FOV of mitochondria (gray) and DRP1 (red). 3:40 min recorded time, fast frame rate 3.8 fps, slow frame rate 0.2 fps.
Supplementary Video 1.1
Slow motion movie corresponding to Supplementary Video 1.0.
Supplementary Video 2.0
Real-time video of a originally 128 × 128 px FOV of mitochondria (gray) and DRP1 (red). 2:04 min recorded time, fast frame rate 3.8 fps, slow frame rate 0.2 fps.
Supplementary Video 2.1
Slow motion video corresponding to Supplementary Video 2.0.
Supplementary Video 3.0
Real-time video of a originally 128 × 128 px FOV of mitochondria (gray) and DRP1 (red). 1:12 min recorded time, fast frame rate 3.8 fps, slow frame rate 0.2 fps.
Supplementary Video 3.1
Slow motion video corresponding to Supplementary Video 3.0.
Supplementary Video 4.0
Real-time video of a 856 × 856 px FOV of C. crescentus (gray) and FtsZ (red). 3:54 h recorded time, fast frames 3 min, slow frames 9 mins.
Supplementary Video 4.1
Slow motion corresponding to Supplementary Video 4.0.
Source data
Source Data Fig. 2
Data points for EDA plot in Fig. 2c.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Data points for intensity contrast plot.
Source Data Extended Data Fig. 4
Data points for EDA plots in Extended Data Fig. 4.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mahecic, D., Stepp, W.L., Zhang, C. et al. Event-driven acquisition for content-enriched microscopy. Nat Methods 19, 1262–1267 (2022). https://doi.org/10.1038/s41592-022-01589-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01589-x
This article is cited by
-
Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Nature Methods (2024)
-
Deep model predictive control of gene expression in thousands of single cells
Nature Communications (2024)
-
EfficientBioAI: making bioimaging AI models efficient in energy and latency
Nature Methods (2024)
-
Computational drug development for membrane protein targets
Nature Biotechnology (2024)
-
Smart lattice light-sheet microscopy for imaging rare and complex cellular events
Nature Methods (2024)