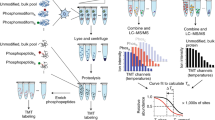
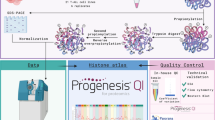
Abstract
Lactylation was initially discovered on human histones. Given its nascence, its occurrence on nonhistone proteins and downstream functional consequences remain elusive. Here we report a cyclic immonium ion of lactyllysine formed during tandem mass spectrometry that enables confident protein lactylation assignment. We validated the sensitivity and specificity of this ion for lactylation through affinity-enriched lactylproteome analysis and large-scale informatic assessment of nonlactylated spectral libraries. With this diagnostic ion-based strategy, we confidently determined new lactylation, unveiling a wide landscape beyond histones from not only the enriched lactylproteome but also existing unenriched human proteome resources. Specifically, by mining the public human Meltome Atlas, we found that lactylation is common on glycolytic enzymes and conserved on ALDOA. We also discovered prevalent lactylation on DHRS7 in the draft of the human tissue proteome. We partially demonstrated the functional importance of lactylation: site-specific engineering of lactylation into ALDOA caused enzyme inhibition, suggesting a lactylation-dependent feedback loop in glycolysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The reprocessed SILAC-based lactylproteome datasets from human MCF-7 cells, the affinity-enriched lactylproteome from the plant fungal pathogen Botrytis cinerea, the affinity-enriched lactylproteome from the protozoan parasite Trypanosoma brucei, the draft map of the Human Proteome and the Meltome Atlas were accessed through the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) with the dataset identifier PXD014870, PXD020746, PXD023011, PXD000561 and PXD011929, respectively. Experimental data collected in this study can be accessed through the Consortium via the iProX partner repository40 with the dataset identifier PXD028488. Uncropped scans of immunoblots and gels shown in Fig. 5d and Supplementary Figs. 7d and 8a can be accessed via https://data.mendeley.com/datasets/4k9y97s7z9/1 on Mendeley Data. Source data are provided with this paper.
References
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Li, L. et al. Glis1 facilitates induction of pluripotency via an epigenome-metabolome-epigenome signalling cascade. Nat. Metab. 2, 882–892 (2020).
Weinert, B. T. et al. Time-resolved analysis reveals rapid dynamics and broad scope of the CBP/p300 acetylome. Cell 174, 231–244 (2018).
Gao, M., Zhang, N. & Liang, W. Systematic analysis of lysine lactylation in the plant fungal pathogen Botrytis cinerea. Front. Microbiol. 11, 594743 (2020).
Zhang, N. et al. Protein lactylation critically regulates energy metabolism in the protozoan parasite Trypanosoma brucei. Front. Cell Dev. Biol. 9, 719720 (2021).
Meng, X., Baine, J. M., Yan, T. & Wang, S. Comprehensive analysis of lysine lactylation in rice (Oryza sativa) grains. J. Agric. Food Chem. 69, 8287–8297 (2021).
Chalkley, R. J. When target-decoy false discovery rate estimations are inaccurate and how to spot instances. J. Proteome Res. 12, 1062–1064 (2013).
Anapindi, K. D. B., Romanova, E. V., Southey, B. R. & Sweedler, J. V. Peptide identifications and false discovery rates using different mass spectrometry platforms. Talanta 182, 456–463 (2018).
Kim, M. S., Zhong, J. & Pandey, A. Common errors in mass spectrometry-based analysis of post-translational modifications. Proteomics 16, 700–714 (2016).
Morelle, W. & Michalski, J. C. Analysis of protein glycosylation by mass spectrometry. Nat. Protoc. 2, 1585–1602 (2007).
Bodenmiller, B., Mueller, L. N., Mueller, M., Domon, B. & Aebersold, R. Reproducible isolation of distinct, overlapping segments of the phosphoproteome. Nat. Methods 4, 231–237 (2007).
Olsen, J. V. et al. Higher-energy C-trap dissociation for peptide modification analysis. Nat. Methods 4, 709–712 (2007).
Trelle, M. B. & Jensen, O. N. Utility of immonium ions for assignment of ε-N-acetyllysine-containing peptides by tandem mass spectrometry. Anal. Chem. 80, 3422–3430 (2008).
Potel, C. M., Lin, M. H., Heck, A. J. R. & Lemeer, S. Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics. Nat. Methods 15, 187–190 (2018).
Lassak, J. et al. Arginine-rhamnosylation as new strategy to activate translation elongation factor P. Nat. Chem. Biol. 11, 266–270 (2015).
Zolg, D. P. et al. ProteomeTools: systematic characterization of 21 post-translational protein modifications by liquid chromatography tandem mass spectrometry (LC–MS/MS) using synthetic peptides. Mol. Cell Proteom. 17, 1850–1863 (2018).
Muroski, J. M., Fu, J. Y., Nguyen, H. H., Ogorzalek Loo, R. R. & Loo, J. A. Leveraging immonium ions for targeting acyl-lysine modifications in proteomic datasets. Proteomics 21, e2000111 (2021).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Gaffney, D. O. et al. Non-enzymatic lysine lactoylation of glycolytic enzymes. Cell Chem. Biol. 27, 206–213 (2020).
Zolg, D. P. et al. Building proteometools based on a complete synthetic human proteome. Nat. Methods 14, 259–262 (2017).
Mongelard, F. & Bouvet, P. Nucleolin: a multiFACeTed protein. Trends Cell Biol. 17, 80–86 (2007).
Jarzab, A. et al. Meltome atlas-thermal proteome stability across the tree of life. Nat. Methods 17, 495–503 (2020).
Huang, J. X. et al. High throughput discovery of functional protein modifications by hotspot thermal profiling. Nat. Methods 16, 894–901 (2019).
Zhang, Z. J., Pedicord, V. A., Peng, T. & Hang, H. C. Site-specific acylation of a bacterial virulence regulator attenuates infection. Nat. Chem. Biol. 16, 95–103 (2020).
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Zemanová, L., Kirubakaran, P., Pato, I. H., Štambergová, H. & Vondrášek, J. The identification of new substrates of human DHRS7 by molecular modeling and in vitro testing. Int. J. Biol. Macromol. 105, 171–182 (2017).
Moreno-Yruela, C. et al. Class I histone deacetylases (HDAC1-3) are histone lysine delactylases. Sci. Adv. 8, eabi6696 (2022).
Wang, M. & Lin, H. Understanding the function of mammalian sirtuins and protein lysine acylation. Annu. Rev. Biochem. 90, 245–285 (2021).
Huang, H. et al. Landscape of the regulatory elements for lysine 2-hydroxyisobutyrylation pathway. Cell Res. 28, 111–125 (2018).
Bian, Y. et al. Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS. Nat. Commun. 11, 157 (2020).
Meier, F. et al. Online parallel accumulation-serial fragmentation (PASEF) with a novel trapped ion mobility mass spectrometer. Mol. Cell. Proteom. 17, 2534–2545 (2018).
Frese, C. K. et al. Improved peptide identification by targeted fragmentation using CID, HCD and ETD on an LTQ-Orbitrap Velos. J. Proteome Res. 10, 2377–2388 (2011).
Bekker-Jensen, D. B. et al. An optimized shotgun strategy for the rapid generation of comprehensive human proteomes. Cell Syst. 4, 587–599 (2017).
Costa Leite, T., Da Silva, D., Guimarães Coelho, R., Zancan, P. & Sola-Penna, M. Lactate favours the dissociation of skeletal muscle 6-phosphofructo-1-kinase tetramers down-regulating the enzyme and muscle glycolysis. Biochem. J. 408, 123–130 (2007).
Prus, G., Hoegl, A., Weinert, B. T. & Choudhary, C. Analysis and interpretation of protein post-translational modification site stoichiometry. Trends Biochem. Sci. 44, 943–960 (2019).
Beausoleil, S. A., Villén, J., Gerber, S. A., Rush, J. & Gygi, S. P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, 1285–1292 (2006).
Ma, B. et al. PEAKS: powerful software for peptide de novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom. 17, 2337–2342 (2003).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Franken, H. et al. Thermal proteome profiling for unbiased identification of direct and indirect drug targets using multiplexed quantitative mass spectrometry. Nat. Protoc. 10, 1567–1593 (2015).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (grant 82173783 to H.Y., grants 81930109, 81720108032 to H.H. and grant 82104050 to N.W.), the National Key Research and Development Program of China (2021YFA1301300 to H.H.), the Fundamental Research Funds for the Central Universities (2632022YC03 to H.Y.), the Natural Science Foundation of Jiangsu Province (BK20210692 to N.W.), the Overseas Expertise Introduction Project for Discipline Innovation (G20582017001 to H.H.) and Sanming Project of Medicine in Shenzhen (SZSM201801060 to H.H.). We thank B. Shan, W. Chen and W. Li from PEAKS Studio and S. Sun from Northwest University for useful discussions.
Author information
Authors and Affiliations
Contributions
H.Y., H.H., G.W. and Nanxi Wang conceived the project. N. Wan, Nian Wang, H.Y., H.H. and Nanxi Wang designed the experiments and analyzed the data. N. Wan, Nian Wang, S.Y., H.Z., Y.K. performed the proteomics experiments. D.W. established lactylation methods. S.Y. and L.L. contributed to data analysis. Nanxi Wang, R.T., S.T., H.L. and W.L. contributed to genetic code expansion and plasmid construction. X.W. and C.S. contributed to metabolomics experiments. H.Y., H.H., G.D., N. Wan and Nanxi Wang wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks W. Andy Tao and the other, anonymous, reviewers for their contribution to the peer review of this work. Arunima Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9, Supplementary Tables 1–7, Supplementary Notes 1–5 and Supplementary References.
Supplementary Table
Supplementary Tables 1–7
Source data
Source Data Fig. 5
Unprocessed Western Blots and/or gels
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wan, N., Wang, N., Yu, S. et al. Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome. Nat Methods 19, 854–864 (2022). https://doi.org/10.1038/s41592-022-01523-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01523-1
This article is cited by
-
Ubiquitous protein lactylation in health and diseases
Cellular & Molecular Biology Letters (2024)
-
Lactylation stabilizes DCBLD1 activating the pentose phosphate pathway to promote cervical cancer progression
Journal of Experimental & Clinical Cancer Research (2024)
-
Immunological profile of lactylation-related genes in Crohn’s disease: a comprehensive analysis based on bulk and single-cell RNA sequencing data
Journal of Translational Medicine (2024)
-
Comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo using highly responsive biosensors
Nature Protocols (2024)
-
Acetyl-methyllysine marks chromatin at active transcription start sites
Nature (2023)