Abstract
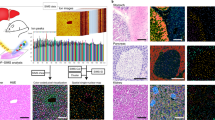
A growing appreciation of the importance of cellular metabolism and revelations concerning the extent of cell–cell heterogeneity demand metabolic characterization of individual cells. We present SpaceM, an open-source method for in situ single-cell metabolomics that detects >100 metabolites from >1,000 individual cells per hour, together with a fluorescence-based readout and retention of morpho-spatial features. We validated SpaceM by predicting the cell types of cocultured human epithelial cells and mouse fibroblasts. We used SpaceM to show that stimulating human hepatocytes with fatty acids leads to the emergence of two coexisting subpopulations outlined by distinct cellular metabolic states. Inducing inflammation with the cytokine interleukin-17A perturbs the balance of these states in a process dependent on NF-κB signaling. The metabolic state markers were reproduced in a murine model of nonalcoholic steatohepatitis. We anticipate SpaceM to be broadly applicable for investigations of diverse cellular models and to democratize single-cell metabolomics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All MALDI-imaging MS data as well as metabolite and lipid annotations and images are publicly available through METASPACE (https://metaspace2020.eu/project/Rappez_2021_SpaceM). The MALDI-imaging MS data and LC–MS/MS datasets are available in the MetaboLights repository under accession number MTBLS78. The microscopy data are available at the European Bioinformatics Institute BioStudies repository under accession number S-BSST369. Source data are provided with this paper.
Code availability
The SpaceM codebase is accessible as Supplementary Software and on GitHub (https://github.com/alexandrovteam/SpaceM). The spatio-molecular matrices and the code for their downstream processing, including generation of the main figures, are available on Google Collaboratory (https://colab.research.google.com/drive/1CKdHDUkGIpAcBzrSfuCodMF_l2xbVAKT).
References
Wellen, K. E. & Thompson, C. B. A two-way street: reciprocal regulation of metabolism and signalling. Nat. Rev. Mol. Cell Biol. 13, 270–276 (2012).
Kim, J. & DeBerardinis, R. J. Mechanisms and implications of metabolic heterogeneity in cancer. Cell Metab. 30, 434–446 (2019).
Johnson, C. H., Ivanisevic, J. & Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 17, 451–459 (2016).
Li, H. et al. The landscape of cancer cell line metabolism. Nat. Med. 25, 850–860 (2019).
Marioni, J. C. & Arendt, D. How single-cell genomics is changing evolutionary and developmental biology. Annu. Rev. Cell Dev. Biol. 33, 537–553 (2017).
Pelkmans, L. Cell biology. Using cell-to-cell variability–a new era in molecular biology. Science 336, 425–426 (2012).
Lee, M.-C. W. et al. Single-cell analyses of transcriptional heterogeneity during drug tolerance transition in cancer cells by RNA sequencing. Proc. Natl Acad. Sci. USA 111, E4726–E4735 (2014).
Russell, A. B., Trapnell, C. & Bloom, J. D. Extreme heterogeneity of influenza virus infection in single cells. eLife 7, e3230 (2018).
Duncan, K. D., Fyrestam, J. & Lanekoff, I. Advances in mass spectrometry based single-cell metabolomics. Analyst 144, 782–793 (2019).
Rubakhin, S. S., Romanova, E. V., Nemes, P. & Sweedler, J. V. Profiling metabolites and peptides in single cells. Nat. Methods 8, S20–S29 (2011).
Ibáñez, A. J. et al. Mass spectrometry-based metabolomics of single yeast cells. Proc. Natl Acad. Sci. USA 110, 8790–8794 (2013).
Qi, M., Philip, M. C., Yang, N. & Sweedler, J. V. Single cell neurometabolomics. ACS Chem. Neurosci. 9, 40–50 (2018).
Ali, A. et al. Single-cell metabolomics by mass spectrometry: advances, challenges, and future applications. Trends Anal. Chem. 120, 115436 (2019).
Gilmore, I. S., Heiles, S. & Pieterse, C. L. Metabolic imaging at the single-cell scale: recent advances in mass spectrometry imaging. Annu. Rev. Anal. Chem. 12, 201–224 (2019).
Lombard-Banek, C. et al. In vivo subcellular mass spectrometry enables proteo-metabolomic single-cell systems biology in a chordate embryo developing to a normally behaving tadpole (X. laevis). Angew. Chem. Int. Ed Engl. https://doi.org/10.1002/anie.202100923 (2021).
Belloni, L. et al. Targeting a phospho-STAT3-miRNAs pathway improves vesicular hepatic steatosis in an in vitro and in vivo model. Sci. Rep. 8, 13638 (2018).
Tanner, N. et al. Regulation of drug metabolism by the interplay of inflammatory signaling, steatosis, and xeno-sensing receptors in HepaRG cells. Drug Metab. Dispos. 46, 326–335 (2018).
Herms, A. et al. Cell-to-cell heterogeneity in lipid droplets suggests a mechanism to reduce lipotoxicity. Curr. Biol. 23, 1489–1496 (2013).
Anstee, Q. M., Reeves, H. L., Kotsiliti, E., Govaere, O. & Heikenwalder, M. From NASH to HCC: current concepts and future challenges. Nat. Rev. Gastroenterol. Hepatol. 16, 411–428 (2019).
Malehmir, M. et al. Platelet GPIbα is a mediator and potential interventional target for NASH and subsequent liver cancer. Nat. Med. 25, 641–655 (2019).
Alexandrov, T. Spatial metabolomics and imaging mass spectrometry in the age of artificial intelligence. Annu. Rev. Biomed. Data Sci. 3, 61–87 (2020).
Patterson, N. H., Tuck, M., Van de Plas, R. & Caprioli, R. M. Advanced registration and analysis of MALDI imaging mass spectrometry measurements through autofluorescence microscopy. Anal. Chem. 90, 12395–12403 (2018).
Wolf, M. J. et al. Metabolic activation of intrahepatic CD8+ T cells and NKT cells causes nonalcoholic steatohepatitis and liver cancer via cross-talk with hepatocytes. Cancer Cell 26, 549–564 (2014).
Spandl, J., White, D. J., Peychl, J. & Thiele, C. Live cell multicolor imaging of lipid droplets with a new dye, LD540. Traffic 10, 1579–1584 (2009).
Molenaar, M. R. et al. LION/web: a web-based ontology enrichment tool for lipidomic data analysis. Gigascience 8, giz061 (2019).
Ress, C. & Kaser, S. Mechanisms of intrahepatic triglyceride accumulation. World J. Gastroenterol. 22, 1664–1673 (2016).
Gluchowski, N. L., Becuwe, M., Walther, T. C. & Farese, R. V. Jr. Lipid droplets and liver disease: from basic biology to clinical implications. Nat. Rev. Gastroenterol. Hepatol. 14, 343–355 (2017).
Baiceanu, A., Mesdom, P., Lagouge, M. & Foufelle, F. Endoplasmic reticulum proteostasis in hepatic steatosis. Nat. Rev. Endocrinol. 12, 710–722 (2016).
Lau, A. N. et al. Dissecting cell-type-specific metabolism in pancreatic ductal adenocarcinoma. eLife https://doi.org/10.7554/eLife.56782 (2020).
Rodríguez-Colman, M. J. et al. Interplay between metabolic identities in the intestinal crypt supports stem cell function. Nature 543, 424–427 (2017).
Robinson, J. L. et al. An atlas of human metabolism. Sci. Signal. 13, eaaz1482 (2020).
Daemen, A. et al. Metabolite profiling stratifies pancreatic ductal adenocarcinomas into subtypes with distinct sensitivities to metabolic inhibitors. Proc. Natl Acad. Sci. USA 112, E4410–E4417 (2015).
Guillaume-Gentil, O. et al. Tunable single-cell extraction for molecular analyses. Cell 166, 506–516 (2016).
Liu, R., Pan, N., Zhu, Y. & Yang, Z. T-Probe: an integrated microscale device for online in situ single cell analysis and metabolic profiling using mass spectrometry. Anal. Chem. 90, 11078–11085 (2018).
Cahill, J. F., Kertesz, V. & Van Berkel, G. J. Laser dissection sampling modes for direct mass spectral analysis. Rapid Commun. Mass Spectrom. 30, 611–619 (2016).
Cahill, J. F., Riba, J. & Kertesz, V. Rapid, untargeted chemical profiling of single cells in their native environment. Anal. Chem. 91, 6118–6126 (2019).
Rubakhin, S. S., Lanni, E. J. & Sweedler, J. V. Progress toward single cell metabolomics. Curr. Opin. Biotechnol. 24, 95–104 (2013).
Zenobi, R. Single-cell metabolomics: analytical and biological perspectives. Science 342, 1243259 (2013).
Zhang, L. & Vertes, A. Single-cell mass spectrometry approaches to explore cellular heterogeneity. Angew. Chem. Int. Ed. Engl. 57, 4466–4477 (2018).
Comi, T. J., Neumann, E. K., Do, T. D. & Sweedler, J. V. microMS: a Python platform for image-guided mass spectrometry profiling. J. Am. Soc. Mass. Spectrom. 28, 1919–1928 (2017).
Neumann, E. K., Comi, T. J., Rubakhin, S. S. & Sweedler, J. V. Lipid heterogeneity between astrocytes and neurons revealed by single-cell MALDI-MS combined with immunocytochemical classification. Angew. Chem. Int. Ed. Engl. 58, 5910–5914 (2019).
Mereu, E. et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat. Biotechnol. 38, 747–755 (2020).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2(+) cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, https://doi.org/10.1038/s41586-019-1373-2 (2019).
Hall, Z. et al. Lipid zonation and phospholipid remodeling in nonalcoholic fatty liver disease. Hepatology 65, 1165–1180 (2017).
Thiam, A. R. & Beller, M. The why, when and how of lipid droplet diversity. J. Cell Sci. 130, 315–324 (2017).
Araya, J. et al. Increase in long-chain polyunsaturated fatty acid n − 6/n − 3 ratio in relation to hepatic steatosis in patients with non-alcoholic fatty liver disease. Clin. Sci. 106, 635–643 (2004).
Sanders, F. W. B. et al. Hepatic steatosis risk is partly driven by increased de novo lipogenesis following carbohydrate consumption. Genome Biol. 19, 79 (2018).
Saito, K. et al. Characterization of hepatic lipid profiles in a mouse model with nonalcoholic steatohepatitis and subsequent fibrosis. Sci. Rep. 5, 12466 (2015).
Apostolopoulou, M. et al. Specific hepatic sphingolipids relate to insulin resistance, oxidative stress, and inflammation in nonalcoholic steatohepatitis. Diabetes Care 41, 1235–1243 (2018).
Gripon, P. et al. Infection of a human hepatoma cell line by hepatitis B virus. Proc. Natl Acad. Sci. USA 99, 15655–15660 (2002).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Palmer, A. et al. FDR-controlled metabolite annotation for high-resolution imaging mass spectrometry. Nat. Methods 14, 57–60 (2017).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Ovchinnikova, K., Kovalev, V., Stuart, L. & Alexandrov, T. OffsampleAI: artificial intelligence approach to recognize off-sample mass spectrometry images. BMC Bioinf. 21, 129 (2020).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Acknowledgements
We thank C.B. Vibe for her support and feedback on the manuscript, M. Shahraz, M. Ekelhof and A. Palmer for help with MALDI-imaging, A. Eisenbarth for software development, N. Typas for advising on biology and providing access to a microscope, B. El Debs and J. Selkrig for training on microscopy and cell culturing, METASPACE software development team (all EMBL), M. Stanifer and S. Boulant (DKFZ) for training on cell culturing, A. Andersen (Life Science Editors) and T. O’Connor (Helmholtz Center Munich) for scientific editing, S. Seah and C. Merten (EMBL) for providing NIH3T3-GFP, F. Merkel and C. Häring (EMBL) for providing HeLa Kyoto H2B-mCherry. We thank other members of the Thesis Advisory Committee of L.R., A.-C. Gavin (EMBL) and B. Brügger (Heidelberg University). This work was funded in part by the European Union’s Horizon 2020 program under agreement numbers 634402, 777222 (T.A.) and 667273 (M.H.), the DKFZ-MOST cooperation program (M.H., M.S.), Darwin Trust of Edinburgh (S.T.), SFB Transregio grant nos. 179, 209, 1335 and I&I Helmholtz Zukunftsthema (all M.H.), the ERC Consolidator grants HepatoMetaboPath (M.H.) and METACELL (T.A.) and ERC Proof of Concept Faith (M.H.). We thank all the reviewers and the editor for detailed feedback that helped improve the paper.
Author information
Authors and Affiliations
Contributions
L.R. and T.A. conceived the method. L.R. developed the method. S.T. conceived and performed the coculture experiments. K.O. performed the on-sample analysis for hepatocytes. R.M.G. and P.P. performed LC–MS/MS validation. M.S. and M.H conceived the hepatocytes study. M.S. cultured and prepared hepatocytes. L.R. and T.A. conceived and performed data analyses. L.R., M.S., M.H. and T.A. interpreted data. L.R., M.S., M.H. and T.A. wrote the paper. T.A. supervised and coordinated the work.
Corresponding authors
Ethics declarations
Competing interests
L.R. and T.A. are the inventors on a patent application describing a spatial single-cell metabolomics method (application in the EU EP3610267A1, US US20200057049A1, Canada CA3059818A1, Australia AU2018252185A1, World Intellectual Property Organization (Patent Cooperation Treaty) WO2018189365A1).
Additional information
Peer review information Nature Methods thanks Young Jin Lee, Peter Nemes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Procedure of detection of laser ablation marks in post-MALDI microscopy images.
The post-MALDI microscopy image is manually cropped around the baltion marks. The Fourier Transform is computed, followed by a low pass filter. The inverse Fourier Transform generates a denoised image with features having spatial frequencies associated with the ablation marks becoming more pronounced. This enables a robust histogram-based thresholding to segment individual ablation marks and determine their centroid coordinates. Every centroid is used as a seed for a region growing algorithm to determine the edges of each ablation mark.
Extended Data Fig. 2 Registration workflow of pre- and post-MALDI microscopy images.
Pen marks are drawn with a black sharpie on the glass slide at the edges of the culturing area. These black pen marks are visible in both the pre- and post-MALDI microscopy images. Histogram-based thresholding is applied to both microscopy images to extract the penmarks areas followed by an edge detection that detects pixels on the edge of the pen marks. This generates more than 400.000 individual features for each microscopy image. A random subset of 5000 features from both pre- (in blue) and post-MALDI (in red) images are used as fiducials to estimate the coordinate transformation for image registration. The overlap of both pre- and post-MALDI fiducials is illustrated. The estimated coordinate transformation from post- to pre-MALDI is applied to the ablation mark coordinates (shown in green) in order to estimate their position in the pre-MALDI microscopy image.
Extended Data Fig. 3 Procedure of the indexation of the segmented ablation marks.
The indexation of the segmented ablation marks involved fitting a theoretical rectangular grid to the ablation marks. In A, the grid angle is estimated by minimizing the number of non-overlapping ablation mark coordinates after projection of the X axis for different rotation angles. In B, the center of the grid is estimated from the extrema of the ablation mark coordinates. The spacing of the grid nodes is estimated in C by minimizing the mean distance for each grid note to the nearest ablation mark. The re-indexing in D is done by choosing the closest ablation mark coordinates from the grid nodes constructed using the parameters defined before (the grid nodes are shown in red, their nearest ablation mark coordinates are shown in black). In E, the ablation mark coordinates are color-coded by their index. An illustration of the different steps for fitting a grid onto the ablation mark coordinates as well as the re-indexing is shown in F. In G, the re-indexed ablation marks are shown.
Extended Data Fig. 4 Details of the SpaceM processing.
The SpaceM processing is composed of two parts: filtering ablation marks and normalizing metabolite intensities across single cells. First, for each cell, its touching ablation marks are filtered based on their area, their sampling proportions (proportion of the ablation mark area sampling any cell) and their sampling specificity (the proportion of their sampling area shared with the cell of interest). Ablation marks sampling predominantly extracellular areas or too many cells at the same time are filtered out (as illustrated here for the ablation mark II and III). Second, for a given metabolite, its intensity in a cell is calculated as a weighted mean of the metabolite intensities from the filtered ablation marks sampling that cell. The intensities are divided by the sampling proportion to account for differences in amount of sampled cellular material between ablation marks. To increase the contribution of ablation marks which sample the cell of interest more than other ablation marks, their intensities are weighted by the sampling specificity. ‘area(a)’ stands for the area of an ablation mark a; ‘sampling area(a)’ stands for the intracellular area of ablation mark ‘a’; ‘area(c)’ stands for the area of a cell ‘c’; all areas are computed in microscopy pixels.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Rights and permissions
About this article
Cite this article
Rappez, L., Stadler, M., Triana, S. et al. SpaceM reveals metabolic states of single cells. Nat Methods 18, 799–805 (2021). https://doi.org/10.1038/s41592-021-01198-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01198-0
This article is cited by
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
A fast-acting lipid checkpoint in G1 prevents mitotic defects
Nature Communications (2024)
-
Dysregulated cellular metabolism in atherosclerosis: mediators and therapeutic opportunities
Nature Metabolism (2024)
-
Integrating single-cell multi-omics and prior biological knowledge for a functional characterization of the immune system
Nature Immunology (2024)
-
Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
Nature Methods (2024)