Abstract
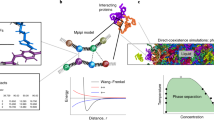
The coarse-grained Martini force field is widely used in biomolecular simulations. Here we present the refined model, Martini 3 (http://cgmartini.nl), with an improved interaction balance, new bead types and expanded ability to include specific interactions representing, for example, hydrogen bonding and electronic polarizability. The updated model allows more accurate predictions of molecular packing and interactions in general, which is exemplified with a vast and diverse set of applications, ranging from oil/water partitioning and miscibility data to complex molecular systems, involving protein–protein and protein–lipid interactions and material science applications as ionic liquids and aedamers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Force-field parameters and procedures (for example, tutorials) are publicly available at http://cgmartini.nl. Simulation data (for example, trajectories) supporting the results of this paper are available from the corresponding authors upon reasonable request.
Code availability
Martinize2 (for which the manuscript is in preparation) and Martinate codes used in this work are publicly available at https://github.com/marrink-lab/. For more detailed information, see Supplementary Codes.
References
Bottaro, S. & Lindorff-Larsen, K. Biophysical experiments and biomolecular simulations: a perfect match? Science 361, 355–360 (2018).
Ingólfsson, H. I. et al. The power of coarse graining in biomolecular simulations. Wiley Interdiscip. Rev. Comput. Mol. Sci. 4, 225–248 (2014).
Marrink, S. J., De Vries, A. H. & Mark, A. E. Coarse grained model for semiquantitative lipid simulations. J. Phys. Chem. B 108, 750–760 (2004).
Marrink, S. J., Risselada, H. J., Yefimov, S., Tieleman, D. P. & de Vries, A. H. The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 111, 7812–7824 (2007).
Uusitalo, J. J., Ingólfsson, H. I., Akhshi, P., Tieleman, D. P. & Marrink, S. J. Martini coarse-grained force field: extension to DNA. J. Chem. Theory Comput. 11, 3932–3945 (2015).
Abellón-Ruiz, J. et al. Structural basis for maintenance of bacterial outer membrane lipid asymmetry. Nat. Microbiol. 2, 1616–1623 (2017).
Yen, H. Y. et al. PtdIns(4,5)P2 stabilizes active states of GPCRs and enhances selectivity of G-protein coupling. Nature 559, 423–427 (2018).
Van Eerden, F. J., Melo, M. N., Frederix, P. W. J. M., Periole, X. & Marrink, S. J. Exchange pathways of plastoquinone and plastoquinol in the photosystem II complex. Nat. Commun. 8, 15214 (2017).
Vögele, M., Köfinger, J. & Hummer, G. Hydrodynamics of diffusion in lipid membrane simulations. Phys. Rev. Lett. 120, 268104 (2018).
Agostino, M. D., Risselada, H. J., Lürick, A., Ungermann, C. & Mayer, A. A tethering complex drives the terminal stage of SNARE-dependent membrane fusion. Nature 551, 634–638 (2017).
Jeena, M. T. et al. Mitochondria localization induced self-assembly of peptide amphiphiles for cellular dysfunction. Nat. Commun. 8, 26 (2017).
Jiang, Z. et al. Subnanometre ligand-shell asymmetry leads to Janus-like nanoparticle membranes. Nat. Mater. 14, 912–917 (2015).
Maingi, V. et al. Stability and dynamics of membrane-spanning DNA nanopores. Nat. Commun. 8, 14784 (2017).
Frederix, P. W. J. M. et al. Exploring the sequence space for (tri-)peptide self-assembly to design and discover new hydrogels. Nat. Chem. 7, 30–37 (2015).
Bochicchio, D., Salvalaglio, M. & Pavan, G. M. Into the dynamics of a supramolecular polymer at submolecular resolution. Nat. Commun. 8, 147 (2017).
Stark, A. C., Andrews, C. T. & Elcock, A. H. Toward optimized potential functions for protein-protein interactions in aqueous solutions: osmotic second virial coefficient calculations using the MARTINI coarse-grained force field. J. Chem. Theory Comput. 9, 4176–4185 (2013).
Javanainen, M., Martinez-Seara, H. & Vattulainen, I. Excessive aggregation of membrane proteins in the Martini model. PLoS ONE 12, e0187936 (2017).
Schmalhorst, P. S., Deluweit, F., Scherrers, R., Heisenberg, C.-P. & Sikora, M. Overcoming the limitations of the MARTINI force field in simulations of polysaccharides. J. Chem. Theory Comput. 13, 5039–5053 (2017).
Alessandri, R. et al. Pitfalls of the Martini model. J. Chem. Theory Comput. 15, 5448–5460 (2019).
Uusitalo, J. J., Ingólfsson, H. I., Marrink, S. J. & Faustino, I. Martini coarse-grained force field: extension to RNA. Biophys. J. 113, 246–256 (2017).
Ben-Naim, A. Molecular Theory of Solutions (Oxford Univ. Press, 2006).
Ploetz, E. A., Bentenitis, N. & Smith, P. E. Kirkwood–Buff integrals for ideal solutions. J. Chem. Phys. 132, 164501 (2010).
Zych, A. J. & Iverson, B. L. Synthesis and conformational characterization of tethered, self-complexing 1,5-dialkoxynaphthalene/1,4,5,8-naphthalenetetracarboxylic diimide systems. J. Am. Chem. Soc. 122, 8898–8909 (2000).
Gabriel, G. J. & Iverson, B. L. Aromatic oligomers that form hetero duplexes in aqueous solution. J. Am. Chem. Soc. 124, 15174–15175 (2002).
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012).
Gao, Z. G. & Ijzerman, A. P. Allosteric modulation of A(2A) adenosine receptors by amiloride analogues and sodium ions. Biochem. Pharmacol. 60, 669–676 (2000).
Okur, H. I. et al. Beyond the Hofmeister series: ion-specific effects on proteins and their biological functions. J. Phys. Chem. B 121, 1997–2014 (2017).
Dupont, D., Depuydt, D. & Binnemans, K. Overview of the effect of salts on biphasic ionic liquid/water solvent extraction systems: anion exchange, mutual solubility, and thermomorphic properties. J. Phys. Chem. B 119, 6747–6757 (2015).
Naert, P., Rabaey, K. & Stevens, C. V. Ionic liquid ion exchange: exclusion from strong interactions condemns cations to the most weakly interacting anions and dictates reaction equilibrium. Green Chem. 20, 4277–4286 (2018).
Khan, H. M. et al. Capturing choline-aromatics cation–π interactions in the MARTINI force field. J. Chem. Theory Comput. 16, 2550–2560 (2020).
Tanaka, K., Caaveiro, J. M. M., Morante, K., González-Manãs, J. M. & Tsumoto, K. Structural basis for self-assembly of a cytolytic pore lined by protein and lipid. Nat. Commun. 6, 6337 (2015).
Huang, G., Willems, K., Soskine, M., Wloka, C. & Maglia, G. Electro-osmotic capture and ionic discrimination of peptide and protein biomarkers with FraC nanopores. Nat. Commun. 8, 935 (2017).
Alessandri, R., Uusitalo, J. J., De Vries, A. H., Havenith, R. W. A. & Marrink, S. J. Bulk heterojunction morphologies with atomistic resolution from coarse-grain solvent evaporation simulations. J. Am. Chem. Soc. 139, 3697–3705 (2017).
Chiu, M. Y., Jeng, U. S., Su, C. H., Liang, K. S. & Wei, K. H. Simultaneous use of small- and wide-angle X-ray techniques to analyze nanometerscale phase separation in polymer heterojunction solar cells. Adv. Mater. 20, 2573–2578 (2008).
Petrov, D. & Zagrovic, B. Are current atomistic force fields accurate enough to study proteins in crowded environments? PLoS Comput. Biol. 10, e1003638 (2014).
Højgaard, C. et al. A soluble, folded protein without charged amino acid residues. Biochemistry 55, 3949–3956 (2016).
Ruckenstein, E. & Shulgin, I. L. Effect of salts and organic additives on the solubility of proteins in aqueous solutions. Adv. Colloid Interface Sci. 123–126, 97–103 (2006).
Zhou, F. X., Cocco, M. J., Russ, W. P., Brunger, A. T. & Engelman, D. M. Interhelical hydrogen bonding drives strong interactions in membrane proteins. Nat. Struct. Biol. 7, 154–160 (2000).
Zhou, F. X., Merianos, H. J., Brunger, A. T. & Engelman, D. M. Polar residues drive association of polyleucine transmembrane helices. Proc. Natl Acad. Sci. USA 98, 2250–2255 (2001).
Grau, B. et al. The role of hydrophobic matching on transmembrane helix packing in cells. Cell Stress 1, 90–106 (2017).
Chen, L., Merzlyakov, M., Cohen, T., Shai, Y. & Hristova, K. Energetics of ErbB1 transmembrane domain dimerization in lipid bilayers. Biophys. J. 96, 4622–4630 (2009).
Artemenko, E. O., Egorova, N. S., Arseniev, A. S. & Feofanov, A. V. Transmembrane domain of EphA1 receptor forms dimers in membrane-like environment. Biochim. Biophys. Acta 1778, 2361–2367 (2008).
Sarabipour, S. & Hristova, K. Glycophorin A transmembrane domain dimerization in plasma membrane vesicles derived from CHO, HEK 293T, and A431 cells. Biochim. Biophys. Acta - Biomembr. 1828, 1829–1833 (2013).
Chen, L., Novicky, L., Merzlyakov, M., Hristov, T. & Hristova, K. Measuring the energetics of membrane protein dimerization in mammalian membranes. J. Am. Chem. Soc. 132, 3628–3635 (2010).
Nash, A., Notman, R. & Dixon, A. M. De novo design of transmembrane helix–helix interactions and measurement of stability in a biological membrane. Biochim. Biophys. Acta - Biomembr. 1848, 1248–1257 (2015).
Finger, C. et al. The stability of transmembrane helix interactions measured in a biological membrane. J. Mol. Biol. 358, 1221–1228 (2006).
Hong, H., Blois, T. M., Cao, Z. & Bowie, J. U. Method to measure strong protein–protein interactions in lipid bilayers using a steric trap. Proc. Natl Acad. Sci. USA 107, 19802–19807 (2010).
Sparr, E. et al. Self-association of transmembrane α-helices in model membranes: importance of helix orientation and role of hydrophobic mismatch. J. Biol. Chem. 280, 39324–39331 (2005).
MacKenzie, K. R., Prestegard, J. H. & Engelman, D. M. Transmembrane helix dimer: structure and implications. Science 276, 131–133 (1997).
Trenker, R., Call, M. E. & Call, M. J. Crystal structure of the glycophorin A transmembrane dimer in lipidic cubic phase. J. Am. Chem. Soc. 137, 15676–15679 (2015).
Domański, J., Sansom, M. S. P., Stansfeld, P. J. & Best, R. B. Balancing force field protein–lipid interactions to capture transmembrane helix–helix association. J. Chem. Theory Comput. 14, 1706–1715 (2018).
Souza, P. C. T., Thallmair, S., Marrink, S. J. & Mera-Adasme, R. An allosteric pathway in copper, zinc superoxide dismutase unravels the molecular mechanism of the G93A amyotrophic lateral sclerosis-linked mutation. J. Phys. Chem. Lett. 10, 7740–7744 (2019).
Brini, E. et al. Systematic coarse-graining methods for soft matter simulations-a review. Soft Matter 9, 2108–2119 (2013).
Foley, T. T., Shell, M. S. & Noid, W. G. The impact of resolution upon entropy and information in coarse-grained models. J. Chem. Phys. 143, 243104 (2015).
Wagner, J. W., Dama, J. F., Durumeric, A. E. P. & Voth, G. A. On the representability problem and the physical meaning of coarse-grained models. J. Chem. Phys. 145, 044108 (2016).
Wörner, S. J., Bereau, T., Kremer, K. & Rudzinski, J. F. Direct route to reproducing pair distribution functions with coarse-grained models via transformed atomistic cross correlations. J. Chem. Phys. 151, 244110 (2019).
Noid, W. G., Chu, J. W., Ayton, G. S. & Voth, G. A. Multiscale coarse-graining and structural correlations: connections to liquid-state theory. J. Phys. Chem. B. 111, 4116–4127 (2007).
Wu, Z., Cui, Q. & Yethiraj, A. Driving force for the association of hydrophobic peptides: the importance of electrostatic interactions in coarse-grained water models. J. Phys. Chem. Lett. 2, 1794–1798 (2011).
Jin, J., Yu, A. & Voth, G. A. Temperature and phase transferable bottom-up coarse-grained models. J. Chem. Theory Comput. 16, 6823–6842 (2020).
Yesylevskyy, S. O., Schäfer, L. V., Sengupta, D. & Marrink, S. J. Polarizable water model for the coarse-grained MARTINI force field. PLoS Comput. Biol. 6, e1000810 (2010).
Michalowsky, J., Schäfer, L. V., Holm, C. & Smiatek, J. A refined polarizable water model for the coarse-grained MARTINI force field with long-range electrostatic interactions. J. Chem. Phys. 146, 054501 (2017).
Marrink, S. J. & Tieleman, D. P. Perspective on the Martini model. Chem. Soc. Rev. 42, 6801–22 (2013).
Bruininks, B. M. H., Souza, P. C. T. & Marrink, S. J. in Biomolecular Simulations: Methods and Protocols (eds Bonomi, M. & Camilloni, C.) 105–127 (Humana Press Inc., 2019).
Liu, J. et al. Enhancing molecular n-type doping of donor-acceptor copolymers by tailoring side chains. Adv. Mater. 30, 1704630 (2018).
Vazquez-Salazar, L. I., Selle, M., de Vries, A., Marrink, S. J. & Souza, P. C. T. Martini coarse-grained models of imidazolium-based ionic liquids: from nanostructural organization to liquid–liquid extraction. Green Chem. 22, 7376–7386 (2020).
Souza, P. C. T. et al. Protein–ligand binding with the coarse-grained Martini model. Nat. Commun. 11, 3714 (2020).
López, C. A. et al. Martini coarse-grained force field: extension to carbohydrates. J. Chem. Theory Comput. 5, 3195–3210 (2009).
Monticelli, L. et al. The MARTINI coarse-grained force field: extension to proteins. J. Chem. Theory Comput. 4, 819–834 (2008).
Grunewald, F., Rossi, G., de Vries, A. H., Marrink, S. J. & Monticelli, L. Transferable MARTINI model of poly(ethylene oxide). J. Phys. Chem. B 122, 7436–7449 (2018).
de Jong, D. H. et al. Improved parameters for the martini coarse-grained protein force field. J. Chem. Theory Comput. 9, 687–97 (2013).
Herzog, F. A., Braun, L., Schoen, I. & Vogel, V. Improved side chain dynamics in MARTINI simulations of protein–lipid interfaces. J. Chem. Theory Comput. 12, 2446–2458 (2016).
Poma, A. B., Cieplak, M. & Theodorakis, P. E. Combining the MARTINI and structure-based coarse-grained approaches for the molecular dynamics studies of conformational transitions in proteins. J. Chem. Theory Comput. 13, 1366–1374 (2017).
Periole, X., Cavalli, M., Marrink, S.-J. & Ceruso, M. A. Combining an elastic network with a coarse-grained molecular force field: structure, dynamics, and intermolecular recognition. J. Chem. Theory Comput. 5, 2531–2543 (2009).
Wassenaar, T. A., Ingólfsson, H. I., Böckmann, R. A., Tieleman, D. P. & Marrink, S. J. Computational lipidomics with insane: a versatile tool for generating Custom membranes for molecular simulations. J. Chem. Theory Comput. 11, 2144–2155 (2015).
Melo, M. N., Ingólfsson, H. I. & Marrink, S. J. Parameters for Martini sterols and hopanoids based on a virtual-site description. J. Chem. Phys. 143, 243152 (2015).
López, C. A., Sovova, Z., van Eerden, F. J., de Vries, A. H. & Marrink, S. J. Martini force field parameters for glycolipids. J. Chem. Theory Comput. 9, 1694–1708 (2013).
Carpenter, T. S. et al. Capturing phase behavior of ternary lipid mixtures with a refined Martini coarse-grained force field. J. Chem. Theory Comput. 14, 6050–6062 (2018).
de Jong, D. H., Baoukina, S., Ingólfsson, H. I. & Marrink, S. J. Martini straight: boosting performance using a shorter cutoff and GPUs. Comput. Phys. Commun. 199, 1–7 (2016).
Hockney, R. W., Goel, S. P. & Eastwood, J. W. Quiet high-resolution computer models of a plasma. J. Comput. Phys. 14, 148–158 (1974).
Páll, S. & Hess, B. A flexible algorithm for calculating pair interactions on SIMD architectures. Comput. Phys. Commun. 184, 2641–2650 (2013).
Verlet, L. Computer ‘experiments’ on classical fluids. I. Thermodynamical properties of Lennard–Jones molecules. Phys. Rev. 159, 98–103 (1967).
Tironi, I. G., Sperb, R., Smith, P. E. & Van Gunsteren, W. F. A generalized reaction field method for molecular dynamics simulations. J. Chem. Phys. 102, 5451–5459 (1995).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Wassenaar, T. A., Ingólfsson, H. I., Prieß, M., Marrink, S. J. & Schäfer, L. V. Mixing MARTINI: electrostatic coupling in hybrid atomistic-coarse-grained biomolecular simulations. J. Phys. Chem. B. 117, 3516–3530 (2013).
Wassenaar, T. A. et al. High-throughput simulations of dimer and trimer assembly of membrane proteins. The DAFT approach. J. Chem. Theory Comput. 11, 2278–91 (2015).
Humphrey, W., Dalke, A. & Schulten, K. VMD—visual molecular dynamics. J. Molec. Graph. 14, 33–38 (1996).
Gowers, R. J. et al. MDAnalysis: a Python package for the rapid analysis of molecular dynamics simulations. in Proc. 15th Python Sci. Conference 98–105 (2016).
Acknowledgements
We thank all members of the S.J.M. group and also external users for testing Martini 3 in its open-beta version. In particular, we thank C. F. E. Schroer, P. W. J. M. Frederix, W. Pezeshkian, M. N. Melo, H. I. Ingólfsson, M. Tsanai, M. König, P. A. Vainikka, T. Zijp, L. Gaifas, J. H. van der Woude, M. Espinoza Cangahuala, M. Scharte, J. Cruiming, L. M. van der Sleen, V. Verduijn, A. H. Beck Frederiksen, B. Schiøtt, M. Sikora, P. Schmalhorst, R. A. Moreira, A. B. Poma, K. Pluhackova, C. Arnarez, C. A. López, E. Jefferys and M. S. P. Sansom for their preliminary tests with a lot of different systems including aedamers, sugars, amino acids, deep eutectic solvents, lipids, peptides and proteins. We also thank the Center for Information Technology of the University of Groningen for providing access to the Peregrine high-performance computing cluster. We acknowledge the National Computing Facilities Foundation of The Netherlands Organization for Scientific Research (NWO), CSC–IT Center for Science Ltd (Espoo, Finland) and CINES (France) for providing computing time. Work in the S.J.M. group was supported by an European Research Council advanced grant no. ‘COMP-MICR-CROW-MEM’. R.A. thanks the NWO (Graduate Programme Advanced Materials, no. 022.005.006) for financial support. L.M. acknowledges the Institut National de la Santé et de la Recherche Medicale and the Agence Nationale de la Recherche for funding (grant no. ANR-17-CE11-0003) and GENCI-CINES for computing time (grant no. A0060710138). S.T. acknowledges the support from the European Commission via a Marie Skłodowska-Curie Actions individual fellowship (MicroMod-PSII, grant agreement no. 748895). M.J. thanks the Emil Aaltonen foundation for financial support. I.V. thanks the Academy of Finland (Center of Excellence program (grant no. 307415)), Sigrid Juselius Foundation, the Helsinki Institute of Life Science fellow program and the HFSP (research grant no. RGP0059/2019). R.B.B. and J.D. were supported by the intramural research program of the NIDDK, NIH. Their work used the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov). H.M.-S. acknowledges the Czech Science Foundation (grant no. 19-19561S). J.B. acknowledges funding from the TOP grant from S.J.M. (NWO) and the EPSRC program grant no. EP/P021123/1. Work in D.P.T.’s group is supported by the Natural Sciences and Engineering Research Council (Canada) and Compute Canada, funded by the Canada Foundation for Innovation. D.P.T. acknowledges further support from the Canada Research Chairs program. N.R. acknowledges funding from the Norwegian Research Council (FRIMEDBIO nos. 251247 and 288008) and computational resources from UNINETT SIGMA2 AS (grant no. NN4700K). H.M.K. acknowledges funding from the University of Calgary through the ‘Eyes High Postdoctoral Fellowship’ program.
Author information
Authors and Affiliations
Contributions
P.C.T.S. and S.J.M. conceived the project with suggestions from R.A., A.H.V., J.B. and S.T. P.C.T.S. generated and optimized all bead parameters. P.C.T.S., R.A. and J.B. generated the topology and bonded parameters of all CG models with suggestions from S.T. and I.F. P.C.T.S., R.A., A.H.V. and F.G. performed the simulations and analysis involving transfer free energies, solvent and polymer properties. P.C.T.S., S.T., J.B. and J.M. performed the simulations and analysis involving lipid bilayers. P.C.T.S., I.F. and R.A. performed the simulations and analysis involving nucleobases. P.C.T.S., I.P. and A.H.V. generated the models and performed the simulations and analysis involving aedamers. P.C.T.S. and F.G. generated the models and performed the simulations and analysis involving ionic liquids and ionic water solutions. R.A. generated the models and performed the simulations and analysis involving bulk heterojunctions, with suggestions from L.M. regarding the fullerene model. P.C.T.S., J.B., H.A., R.A., B.M.H.B., S.T., J.M., V.N., X.P., M.J., H.M.K., J.D., V.C. and H.M.-S. performed the simulations and analysis involving amino acids, peptides and proteins. J.B., T.A.W., P.C.K. and S.T. developed some tools and scripts used to generate the CG models and to run the molecular dynamics simulations. L.M., R.B.B., P.T., N.R., I.V., A.H.V. and S.J.M. provided guidance and supervision in the studies performed by their respective group members and collaborators. P.C.T.S. and S.J.M. wrote the main manuscript, with contributions from all the authors. P.C.T.S. prepared the figures with contributions from R.A., B.M.H.B., H.M.K. and A.H.V. P.C.T.S. wrote the Methods with contributions from all the authors. P.C.T.S. wrote the Supplementary Information, with contributions from all the authors. All the authors revised and approved the final version of the manuscript, Methods and Supplementary Information.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks the anonymous reviewers for their contribution for the peer review of this work. Arunima Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes, Results and Codes.
Supplementary Table 1
Comparison between simulation and experimental results
Rights and permissions
About this article
Cite this article
Souza, P.C.T., Alessandri, R., Barnoud, J. et al. Martini 3: a general purpose force field for coarse-grained molecular dynamics. Nat Methods 18, 382–388 (2021). https://doi.org/10.1038/s41592-021-01098-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01098-3
This article is cited by
-
Vesicle protrusion induced by antimicrobial peptides suggests common carpet mechanism for short antimicrobial peptides
Scientific Reports (2024)
-
The juxtamembrane linker of synaptotagmin 1 regulates Ca2+ binding via liquid-liquid phase separation
Nature Communications (2024)
-
Synthesis of lipid-linked precursors of the bacterial cell wall is governed by a feedback control mechanism in Pseudomonas aeruginosa
Nature Microbiology (2024)
-
From structural polymorphism to structural metamorphosis of the coat protein of flexuous filamentous potato virus Y
Communications Chemistry (2024)
-
Micropolarity governs the structural organization of biomolecular condensates
Nature Chemical Biology (2024)