Abstract
Super-resolution microscopies have become an established tool in biological research. However, imaging throughput remains a main bottleneck in acquiring large datasets required for quantitative biology. Here we describe multifocal flat illumination for field-independent imaging (mfFIFI). By integrating mfFIFI into an instant structured illumination microscope (iSIM), we extend the field of view (FOV) to >100 × 100 µm2 while maintaining high-speed, multicolor, volumetric imaging at double the diffraction-limited resolution. We further extend the effective FOV by stitching adjacent images for fast live-cell super-resolution imaging of dozens of cells. Finally, we combine our flat-fielded iSIM with ultrastructure expansion microscopy to collect three-dimensional (3D) images of hundreds of centrioles in human cells, or thousands of purified Chlamydomonas reinhardtii centrioles, per hour at an effective resolution of ~35 nm. Classification and particle averaging of these large datasets enables 3D mapping of posttranslational modifications of centriolar microtubules, revealing differences in their coverage and positioning.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The datasets generated and analyzed in this study are available from the corresponding authors upon request. Sample datasets have been made available on Zenodo (https://zenodo.org/record/3613906#.Xib5J1NKhTY).
Code availability
The wave optics simulation platform for optimizing mfFIFI is provided as Supplementary Software 1. Updated versions can be found on GitHub (https://github.com/dmahecic/simmla_w_mfFIFI). The custom MATLAB source code for aligning 3D particles is available at https://github.com/dfortun2/U-ExM37.
References
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Gustafsson, M. G. L. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Hell Stefan, W. & Jan, W. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Tanaami, T. et al. High-speed 1-frame/ms scanning confocal microscope with a microlens and Nipkow disks. Appl. Opt. 41, 4704 (2002).
Bingen, P., Reuss, M., Engelhardt, J. & Hell, S. W. Parallelized STED fluorescence nanoscopy. Opt. Express 19, 23716 (2011).
Chmyrov, A. et al. Nanoscopy with more than 100,000 ‘doughnuts’. Nat. Methods 10, 737–740 (2013).
York, A. G. et al. Instant super-resolution imaging in live cells and embryos via analog image processing. Nat. Methods 10, 1122–1126 (2013).
Schulz, O. et al. Resolution doubling in fluorescence microscopy with confocal spinning-disk image scanning microscopy. Proc. Natl Acad. Sci. USA 110, 21000–21005 (2013).
Douglass, K. M., Sieben, C., Archetti, A., Lambert, A. & Manley, S. Super-resolution imaging of multiple cells by optimized flat-field epi-illumination. Nat. Photonics 10, 705–708 (2016).
Archetti, A. et al. Waveguide-PAINT offers an open platform for large field-of-view super-resolution imaging. Nat. Commun. 10, 1267 (2019).
Chen, F., Tillberg, P. W. & Boyden, E. S. Expansion microscopy. Science 347, 543–548 (2015).
Gao, R. et al. Cortical column and whole-brain imaging with molecular contrast and nanoscale resolution. Science 363, eaau8302 (2019).
Cahoon, C. K. et al. Superresolution expansion microscopy reveals the three-dimensional organization of the Drosophila synaptonemal complex. Proc. Natl Acad. Sci. USA 114, 6857–6866 (2017).
Gao, M. et al. Expansion stimulated emission depletion microscopy (ExSTED). ACS Nano 12, 4178–4185 (2018).
Xu, H. et al. Molecular organization of mammalian meiotic chromosome axis revealed by expansion STORM microscopy. Proc. Natl Acad. Sci. USA 116, 18423–18428 (2019).
Mahecic, D., Testa, I., Griffié, J. & Manley, S. Strategies for increasing the throughput of super-resolution microscopies. Curr. Opin. Chem. Biol. 51, 84–91 (2019).
Sacconi, L. et al. Multiphoton multifocal microscopy exploiting a diffractive optical element. Opt. Lett. 28, 1918 (2003).
Coelho, S. et al. Multifocal multiphoton microscopy with adaptive optical correction. Multiphot. Microsc. Biomed. Sci. XIII 8588, 858817 (2013).
Cooper, D. J. F. Imaging distal end of multimode fiber. US Patent 867,017,8B2 (2014).
Nielsen, T., Fricke, M., Hellweg, D. & Andresen, P. High efficiency beam splitter for multifocal multiphoton microscopy. J. Microsc. 201, 368–376 (2001).
Zimmermann, M., Lindlein, N., Voelkel, R. & Weible, K. J. Microlens laser beam homogenizer: theory application. In Proc. SPIE 6663, Laser Beam Shaping VIII 666302 (SPIE, 2007).
Voelkel, R. & Weible, K. J. Laser beam homogenizing: limitations and constraints. In Proc. SPIE 7102, Optical Fabrication, Testing, and Metrology III 71020J (eds. Duparré, A. & Geyl, R.) (2008).
Gambarotto, D. et al. Imaging cellular ultrastructures using expansion microscopy (U-ExM). Nat. Methods 16, 71–74 (2019).
Schreiber, P., Kudaev, S., Dannberg, P. & Zeitner, U. D. Homogeneous LED-illumination using microlens arrays. Nonimaging Opt. Effic. Illum. Syst. II 5942, 59420K (2005).
Curd, A. et al. Construction of an instant structured illumination microscope. Methods 88, 37–47 (2015).
Model, M. A. & Blank, J. L. Concentrated dyes as a source of two-dimensional fluorescent field for characterization of a confocal microscope. J. Microsc. 229, 12–16 (2008).
Friedman, J. R. et al. ER tubules mark sites of mitochondrial division. Science 334, 358–362 (2011).
Görlitz, F. et al. easySLM-STED: stimulated emission depletion microscopy with aberration correction, extended field of view and multiple beam scanning. J. Biophotonics 11, 1–11 (2018).
Guichard, P., Chrétien, D., Marco, S. & Tassin, A. M. Procentriole assembly revealed by cryo-electron tomography. EMBO J. 29, 1565–1572 (2010).
Banterle, N. & Gönczy, P. Centriole biogenesis: from identifying the characters to understanding the plot. Annu. Rev. Cell Dev. Biol. 33, 23–49 (2017).
Janke, C. & Bulinski, J. C. Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 12, 773–786 (2011).
Bobinnec, Y. et al. Centriole disassembly in vivo and its effect on centrosome structure and function in vertebrate cells. J. Cell Biol. 143, 1575–1589 (1998).
Xu, Z. et al. Microtubules acquire resistance from mechanical breakage through intralumenal acetylation. Science 356, 328–332 (2017).
Portran, D., Schaedel, L., Xu, Z., Théry, M. & Nachury, M. V. Tubulin acetylation protects long-lived microtubules against mechanical ageing. Nat. Cell Biol. 19, 391–398 (2017).
Abal, M., Keryer, G. & Bornens, M. Centrioles resist forces applied on centrosomes during G2/M transition. Biol. Cell 97, 425–434 (2005).
Wong, Y. L. et al. Reversible centriole depletion with an inhibitor of Polo-like kinase 4. Science 348, 1155–1160 (2015).
Hamel, V. et al. Identification of chlamydomonas central core centriolar proteins reveals a role for human WDR90 in ciliogenesis. Curr. Biol. 27, 2486–2498.e6 (2017).
Fortun, D. et al. Reconstruction from multiple particles for 3D isotropic resolution in fluorescence microscopy. IEEE Trans. Med. Imaging 37, 1235–1246 (2018).
Sieben, C., Banterle, N., Douglass, K. M., Gönczy, P. & Manley, S. Multicolor single-particle reconstruction of protein complexes. Nat. Methods 15, 777–780 (2018).
Jiang, J., Pentelute, B. L., Collier, R. J. & Hong Zhou, Z. Atomic structure of anthrax protective antigen pore elucidates toxin translocation. Nature 521, 545–549 (2015).
Meehl, J. B., Bayless, B. A., Giddings, T. H., Pearson, C. G. & Winey, M. Tetrahymena Poc1 ensures proper intertriplet microtubule linkages to maintain basal body integrity. Mol. Biol. Cell 27, 2394–2403 (2016).
Le Guennec, M. et al. A helical inner scaffold provides a structural basis for centriole cohesion. Sci. Adv. 6, eaaz4137 (2020).
Edelstein, A. D. et al. Advanced methods of microscope control using μManager software. J. Biol. Methods 1, 10 (2014).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Arganda-Carreras, I. et al. Trainable Weka Segmentation: a machine learning tool for microscopy pixel classification. Bioinformatics 33, 2424–2426 (2017).
Mahecic, D. et al. Membrane bending energy and tension govern mitochondrial division. Preprint at bioRxiv https://doi.org/10.1101/255356 (2018).
Klena, N. et al. Isolationand fluorescence imaging for single-particle reconstruction of chlamydomonascentrioles. J. Vis. Exp. 139, 58109 (2018).
Acknowledgements
We thank H. Shroff and A. Curd for their help and advice on the construction of the iSIM; H. Perreten for technical assistance; C. Berz for her help with multi-FOV imaging; R. Kirchner on discussions on the Köhler integrator and C. Sieben and L. Carlini for critical reading of the manuscript. This work is supported by the European Research Council (grants AdG 835322, CENGIN to P. Gönczy and StG 715289 ACCENT to P. Guichard), the Swiss National Science Foundation (SNSF) grants PP00P3_157517 to P. Gu and 182429 to S.M., the MSCA (grant 75200, CARTASSY to N.B.), and the NCCR Chemical Biology (S.M. and P. Gönczy).
Author information
Authors and Affiliations
Contributions
D.M., K.M.D., P. Gönczy, V.H., P. Guichard and S.M. conceived and designed the project. P. Gönczy, V.H., P. Guichard and S.M. supervised the project. D.M., K.M.D. and S.M. designed the multifocal illumination system. D.M. and K.M.D. developed the simulation platform. D.M. built the microscope, performed all experiments and data analysis. D.G. performed the purified C. reinhardtii centriole sample preparation. D.F. performed single particle averaging and reconstruction. N.B. performed the human RPE-1 centriole sample preparation. M.L.G. performed the particle classification and alignment. K.A.I. built and performed experiments on the flat-fielding characterization platform. D.M. and S.M. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The mfFIFI flat-fielding module has been covered in a patent application filed by EPFL, with D. Mahecic, K.M. Douglass and S. Manley as the inventors and is currently pending under patent application number PCT/EP2018/085228.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended Data
Extended Data Fig. 1 mfFIFI development stages.
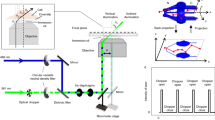
a) A standard Köhler uses a pair of flat-fielding MLAs to split the incoming light into multiple channels, before focusing them in the front-focal plane of the Fourier lens. In the case of a coherent light source, such as lasers, a focusing lens and a rotating diffuser are used to scramble the incoming light and generate an extended partially coherent source, which is then collimated by the collimating lens. Implementing a traditional Köhler integrator with a variable length L3 between the second flat-fielding MLA and the Fourier lens, results in nontelecentric illumination of the excitation MLA. This will in turn cause the pitch of the excitation spots produced by the excitation MLA to vary. b) The telecentric Köhler integrator assures that the pitch of the excitation spots generated by the excitation MLA is conserved, by setting the distance between the second flat-fielding MLA and the Fourier lens to the focal length of the Fourier lens: L3 = FFL. The partially coherent extended source will however limit the capability of the microscope to achieve diffraction-limited excitation at the sample. A possible solution would be to limit the size of the spots by placing a pinhole array in the front focal plane of the excitation MLA, although at a cost to the transmission efficiency. c) The extended design overcomes these problems by introducing a beam expander between the collimating lens and the first flat-fielding MLA, which allows control over how many microlens channels are used to average over in the focus of the Fourier lens. Alternatively, placing a hard aperture to limit the radius of the beam incident on the flat-fielding MLAs would have a similar effect, albeit rejecting much of the incident light.
Extended Data Fig. 2 Optimization of design parameters using the extended simulation platform.
a) Schematic showing the standard Köhler integrator setup, indicating how the different design parameters are defined in the simulation. b) Field amplitude at the front focal plane of the Fourier lens, corresponding to the field incident on the excitation MLA, and c) at the front focal plane of the excitation MLA for different distances L2 of the Fourier lens from the flat-fielding MLAs. The telecentric condition corresponds to L2 = 0. d) Average pitch of the multifocal excitation measured at the front focal plane of the excitation MLA for different values of L2. Dashed line marks the actual pitch of the excitation MLA. e) Schematic representation showing the mfFIFI configuration including the beam contractor and labeling the simulation parameters. The beam contraction factor is set by the inverse of the magnification of the two lenses of the beam contractor: F2/F1. f) Field amplitude at the front focal plane of the Fourier lens and g) the front focal plane of the excitation MLA. h) Trade-off of the beam contraction factor between the spot size (left axis, n = 87 excitation peaks) and the homogeneity of the excitation spots (right axis), quantified by the plateau uniformity; the central mark on the boxplot indicates the median, and the bottom and top edges of the box indicate the 25th and 75th percentiles, respectively. Shown data representative out of >3 similar independent simulations.
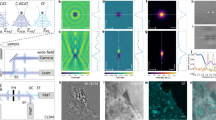
Extended Data Fig. 3 Quantifying excitation homogeneity as function of the scrambling speed of the rotating diffuser.
a) Representative simulated data showing the relative field intensity in the plane of the excitation MLA and b) at the front focal plane of the excitation MLA. c-d) Variation in excitation spot localization, width and amplitude for illumination averaged over different numbers of simulated random wavefronts (N = 909 binned peak variations). f) Real data showing the field intensity in the plane of the excitation MLA and g) at the front focal plane of the excitation MLA. In this case the two fields were not acquired at the same random wavefront, but are intended to give a general idea of what the fields look like. h–j) Variation in excitation spot localization, width and amplitude for illumination averaged over different numbers of random wavefronts acquired experimentally (N = 4248 binned peak variations). k–m) Dependence of the spot localization, width and amplitude as function of the number of iterations over which the illumination is averaged over (representing different scrambling speeds, or acquisition speeds). n–p) Variation in the localization, width and amplitude between individual random wavefronts for simulated (N = 499900 individual peak variations) and experimentally acquired data (N = 973736 individual peak variations).Shown simulated data representative out of >3 similar independent simulations.
Extended Data Fig. 4 Integrating mfFIFI into an instant structured illumination microscope.
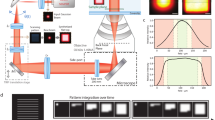
a) Schematic representation of the iSIM setup, showing where the mfFIFI module is integrated into the excitation path prior to the excitation MLA. b) Simulated flat-field incident on the excitation MLA. c) Simulated intensity profile in the front-focal plane of the excitation MLA, showing an array of excitation points (inset). Simulated data representative out of >3 similar independent simulations.
Extended Data Fig. 5 Centriole particles from expanded RPE-1 cells.
Montage of a random subset of unclassified raw particles collected in situ from expanded synchronized human RPE-1 cells stained for acetylated tubulin (yellow) and PolyE (magenta). Scale bar: 1 µm.
Extended Data Fig. 6 Particle shape analysis and PTM coverage.
a) Particle lengths measured along side views of expanded human centrioles in the cellular context. b) Particle diameters measured on top views of human centrioles. c) PolyE coverage for human centrioles with respect to the acetylated tubulin signal along the length of the centriole. d) PTM length coverage measured along side views of purified Chlamydomonas reinhardtii centrioles. e) Particle diameters measured on top views of purified Chlamydomonas reinhardtii centrioles. α-tubulin signal was measured from three different datasets (dual-labeling with acetylated tubulin, GT335 and PolyE), with their individual means marked individually (acetylated tubulin in yellow, α-tubulin in blue, GT335 in orange and PolyE in magenta). f) PTM coverage for purified Chlamydomonas reinhardtii centrioles measured by dividing the length profiles of different PTMs by their respective α-tubulin signal. All scales reflect pre-expansion size. N = 25 top views and N = 41 side views for human centrioles. N = 50 for acetylation dataset, N = 49 for GT335 dataset and N = 50 for PolyE dataset in Chlamydomonas reinhardtii. The center line represents the mean and error bars represent the standard deviation.
Extended Data Fig. 7 Multi-color particle averaging and reconstruction of tubulin PTMs in Chlamydomonas reinhardtii centrioles.
a) Side and top views of α-tubulin reference with different PTMs. Top views are taken from the distal side toward the proximal side. Scale bar: 100 nm.
Extended Data Fig. 8 Expanded purified Chlamydomonas reinhardtii centrioles.
a–c) Montages of maximum intensity projections of a subset of centriole particles collected with dual-staining for a) acetylated tubulin, b) PolyE and c) GT335 with α-tubulin used as reference in each case. Inset in b) shows examples of PolyE twisting as single color maximum intensity projections. Scale bars: 1 µm.
Extended Data Fig. 9 Particle classification prior to reconstruction.
Dendrograms generated via hierarchical classification in 10 groups. On top, the average volume of all particles is displayed with 2 different orientations. For each group, the average volume is shown with the same 2 orientations, the tubulin signal dark-coloured and the (a, b) acetylated, (c, d) glutamylated and (e, f) polyglutamylated signal light-coloured.
Extended Data Fig. 10 Twist of polyglutamylated tubulin along Chlamydomonas reinhardtii centriole.
a) Schematic illustration of centriolar microtubule triplets and expected radial and tangential displacements between microtubules in neighbouring triplets (inset) viewed from the proximal end. b) Schematic representation of XY planes between the shift and twist models of PolyE shift along the proximal-distal centriole axis and their predictions (cross sections viewed from the proximal end). c) Cross section of the reconstruction YZ profile showing the barrel diameter with α-tubulin and PolyE signal. d) Radial position of PolyE signal relative to α-tubulin, measured at different positions along the reconstructed centriole for all 9 microtubule triplets, from the proximal to the distal end. Points mark the mean of the N = 9 triplet measurements and error bars their standard deviation. e) The relative radial PolyE shift was calculated with respect to the α-tubulin signal by taking the difference between their radial positions, determined by Gaussian fitting. The profiles were measured from top views of the reconstructed centriole (inset). f, g) The relative tangential shift of PolyE is measured from the tangential projection (inset) with respect to α-tubulin at the f) distal and g) proximal end.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Tables 1 and 2, captions for Videos 1–5 and Discussion.
Supplementary Video 1
Simulation showing how integrating over an increasing number of random wavefronts achieves homogeneous illumination of the excitation MLA (left) and hence a homogeneous array of excitation spots in the front focal plane of the excitation MLA (right). Simulation repeated five times with similar results.
Supplementary Video 2
Intensity in the front focal plane of the excitation MLA at different positions of the rotating diffuser. The video should represent what the excitation looks like ‘in real time’, without any effect of temporal averaging.
Supplementary Video3
Experimental data showing how integrating over an increased number of random wavefronts generates a homogeneous array of excitation spots in the front focal plane of the excitation MLA. Simulated integration repeated five times with similar results.
Supplementary Video 4
Large FOV imaging of Cos-7 cells transfected with KDEL-RFP. Exposure time, 100 ms. Experiment repeated 13 times with similar results.
Supplementary Video 5
Multi-FOV imaging of mitochondria-ER contacts in Cos-7 cells transfected with Mito-GFP and KDEL-RFP. The acquisition was iterated over a grid of 2 × 2 FOVs and stitched back into a Video with an effective temporal resolution of 2 s. Experiment repeated three times with similar results.
Supplementary Software
testBE.ipynb.txt and testL2.ipynb.txt
Rights and permissions
About this article
Cite this article
Mahecic, D., Gambarotto, D., Douglass, K.M. et al. Homogeneous multifocal excitation for high-throughput super-resolution imaging. Nat Methods 17, 726–733 (2020). https://doi.org/10.1038/s41592-020-0859-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0859-z
This article is cited by
-
In situ architecture of Opa1-dependent mitochondrial cristae remodeling
The EMBO Journal (2024)
-
Enhanced detection of fluorescence fluctuations for high-throughput super-resolution imaging
Nature Photonics (2023)
-
Field-dependent deep learning enables high-throughput whole-cell 3D super-resolution imaging
Nature Methods (2023)
-
iU-ExM: nanoscopy of organelles and tissues with iterative ultrastructure expansion microscopy
Nature Communications (2023)
-
Event-driven acquisition for content-enriched microscopy
Nature Methods (2022)