Abstract
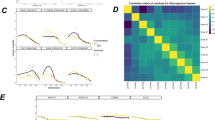
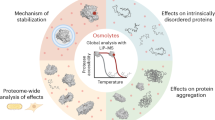
We have used a mass spectrometry-based proteomic approach to compile an atlas of the thermal stability of 48,000 proteins across 13 species ranging from archaea to humans and covering melting temperatures of 30–90 °C. Protein sequence, composition and size affect thermal stability in prokaryotes and eukaryotic proteins show a nonlinear relationship between the degree of disordered protein structure and thermal stability. The data indicate that evolutionary conservation of protein complexes is reflected by similar thermal stability of their proteins, and we show examples in which genomic alterations can affect thermal stability. Proteins of the respiratory chain were found to be very stable in many organisms, and human mitochondria showed close to normal respiration at 46 °C. We also noted cell-type-specific effects that can affect protein stability or the efficacy of drugs. This meltome atlas broadly defines the proteome amenable to thermal profiling in biology and drug discovery and can be explored online at http://meltomeatlas.proteomics.wzw.tum.de:5003/ and http://www.proteomicsdb.org.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Supplementary Information is available in the online version of this manuscript. The mass spectrometry raw files, peptide and protein identification search engine output files and files showing the melting curves for all proteins have been deposited with the proteomeXchange consortium via the PRIDE partner repository under the project name ‘Meltome atlas—thermal proteome stability across the tree of life’ and the data set identifier PXD011929.
The melting curves for all proteins are also available in the online R Shiny App http://meltomeatlas.proteomics.wzw.tum.de:5003/. The human meltome data are also available at https://www.proteomicsdb.org. Files containing data sources for main text figures can be downloaded from https://figshare.com/account/home#/projects/75567. Source data for Figs. 2–6 are presented with the paper.
Code availability
Code for data analysis is available for download at Bioconductor: https://bioconductor.org/packages/release/bioc/html/TPP.html.
References
Leuenberger, P. et al. Cell-wide analysis of protein thermal unfolding reveals determinants of thermostability. Science. 355, eaai7825 (2017).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Chretien, D. et al. Mitochondria are physiologically maintained at close to 50 degrees C. PLoS Biol. 16, e2003992 (2018).
Rai, A. K., Chen, J. X., Selbach, M. & Pelkmans, L. Kinase-controlled phase transition of membraneless organelles in mitosis. Nature 559, 211–216 (2018).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Becher, I. et al. Pervasive protein thermal stability variation during the cell cycle. Cell 173, 1495–1507 e1418 (2018).
Dai, L. et al. Modulation of protein-interaction states through the cell cycle. Cell 173, 1481–1494 e1413 (2018).
Tan, C. S. H. et al. Thermal proximity coaggregation for system-wide profiling of protein complex dynamics in cells. Science 359, 1170–1177 (2018).
Reinhard, F. B. et al. Thermal proteome profiling monitors ligand interactions with cellular membrane proteins. Nat. Methods 12, 1129–1131 (2015).
Siddiqui, K. S. & Cavicchioli, R. Cold-adapted enzymes. Annu. Rev. Biochem. 75, 403–433 (2006).
Mateus, A. et al. Thermal proteome profiling in bacteria: probing protein state in vivo. Mol. Syst. Biol. 14, e8242 (2018).
Samaras, P. et al. ProteomicsDB: a multi-omics and multi-organism resource for life science research. Nucleic Acids Res. 48 (D1), D1153–D1163 (2019).
Schmidt, T. et al. ProteomicsDB. Nucleic Acids Res. 46, 1271–1281 (2018).
Seashore-Ludlow, B., Axelsson, H. & Lundback, T. Perspective on CETSA Literature: toward more quantitative data interpretation. SLAS Disco. 25, 118–126 (2019). 2472555219884524.
Van Derlinden, E. & Van Impe, J. F. Modeling growth rates as a function of temperature: model performance evaluation with focus on the suboptimal temperature range. Int. J. Food Microbiol. 158, 73–78 (2012).
Le, T. T. et al. Hydrophilic lecithins protect milk proteins against heat-induced aggregation. Colloids Surf. B. 60, 167–173 (2007).
Guruharsha, K. G. et al. A protein complex network of Drosophila melanogaster. Cell 147, 690–703 (2011).
Keseler, I. M. et al. The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 45, 543–550 (2017).
Mukhopadhyay, A., Ray, S. & De, M. Detecting protein complexes in a PPI network: a gene ontology based multi-objective evolutionary approach. Mol. Biosyst. 8, 3036–3048 (2012).
Ori, A. et al. Spatiotemporal variation of mammalian protein complex stoichiometries. Genome Biol. 17, 47 (2016).
Sterner, R. & Liebl, W. Thermophilic adaptation of proteins. Crit. Rev. Biochem. Mol. Biol. 36, 39–106 (2001).
Jaenicke, R. & Bohm, G. The stability of proteins in extreme environments. Curr. Opin. Struct. Biol. 8, 738–748 (1998).
Nick Pace, C., Scholtz, J. M. & Grimsley, G. R. Forces stabilizing proteins. FEBS Lett. 588, 2177–2184 (2014).
Zeldovich, K. B., Berezovsky, I. N. & Shakhnovich, E. I. Protein and DNA sequence determinants of thermophilic adaptation. PLoS Comput. Biol. 3, e5 (2007).
Zhou, X. X., Wang, Y. B., Pan, Y. J. & Li, W. F. Differences in amino acids composition and coupling patterns between mesophilic and thermophilic proteins. Amino Acids 34, 25–33 (2008).
Weingarten, M. D., Lockwood, A. H., Hwo, S. Y. & Kirschner, M. W. A protein factor essential for microtubule assembly. Proc. Natl Acad. Sci. USA 72, 1858–1862 (1975).
Weerkamp, F. et al. Flow cytometric immunobead assay for the detection of BCR-ABL fusion proteins in leukemia patients. Leukemia 23, 1106–1117 (2009).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol. Cell Proteom. 11, M111 014050 (2012).
Zhou, B. et al. Comprehensive, integrated, and phased whole-genome analysis of the primary ENCODE cell line K562. Genome Res. 29, 472–484 (2019).
Gioia, L., Siddique, A., Head, S. R., Salomon, D. R. & Su, A. I. A genome-wide survey of mutations in the Jurkat cell line. BMC Genomics 19, 334 (2018).
Zecha, J. et al. Peptide level turnover measurements enable the study of proteoform dynamics. Mol. Cell Proteom. 17, 974–992 (2018).
Chang, W. K., Yang, K. D., Chuang, H., Jan, J. T. & Shaio, M. F. Glutamine protects activated human T cells from apoptosis by up-regulating glutathione and Bcl-2 levels. Clin. Immunol. 104, 151–160 (2002).
Dundas, S. R., Lawrie, L. C., Rooney, P. H. & Murray, G. I. Mortalin is over-expressed by colorectal adenocarcinomas and correlates with poor survival. J. Pathol. 205, 74–81 (2005).
Kwiatkowski, N. et al. Targeting transcription regulation in cancer with a covalent CDK7 inhibitor. Nature 511, 616–620 (2014).
Liu, H. et al. Impact of IgG Fc-oligosaccharides on recombinant monoclonal antibody structure, stability, safety, and efficacy. Biotechnol. Prog. 33, 1173–1181 (2017).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 34, D668–D672 (2006).
Becher, I. et al. Thermal profiling reveals phenylalanine hydroxylase as an off-target of panobinostat. Nat. Chem. Biol. 12, 908–910 (2016).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Choorapoikayil, S., Overvoorde, J. & den Hertog, J. Deriving cell lines from zebrafish embryos and tumors. Zebrafish 10, 316–325 (2013).
Ruprecht, B., Zecha, J., Zolg, D. P. & Kuster, B. High pH reversed-phase micro-columns for simple, sensitive, and efficient fractionation of proteome and (TMT labeled) phosphoproteome digests. Methods Mol. Biol. 1550, 83–98 (2017).
Yu, P. et al. Trimodal mixed mode chromatography that enables efficient offline two-dimensional peptide fractionation for proteome analysis. Anal. Chem. 89, 8884–8891 (2017).
Ruprecht, B. et al. Hydrophilic strong anion exchange (hSAX) chromatography enables deep fractionation of tissue proteomes. Methods Mol. Biol. 1550, 69–82 (2017).
Ruprecht, B. et al. in Proteomics: Methods and Protocols (eds Comai, L. et al.) 69–82 (Springer, 2017).
Hahne, H. et al. DMSO enhances electrospray response, boosting sensitivity of proteomic experiments. Nat. Methods 10, 989–991 (2013).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Yachdav, G. et al. PredictProtein-an open resource for online prediction of protein structural and functional features. Nucleic Acids Res. 42, 337–343 (2014).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Acknowledgements
The data reported here are tabulated in the main paper and Supplementary Information. We thank P. Giansanti for providing the mouse samples; A. Hubauer and S. Petzoldt for technical assistance; M. Übelacker for assistance with bacterial cultivation; I. Cornella-Taracido (Merck Research Laboratories, Boston, MA), C. Siegl, S. Dieter and H. Glimm (NCT Heidelberg, Germany); A. Augustin (Hoffmann-La Roche, Basel, Switzerland) and colleagues for providing access to their meltome data; and M. Bernhofer, J. Reeb and B. Rost for computing and providing protein structure predictions. A.J. is grateful for funding from the Alexander von Humboldt foundation, J.Z. for funding from the German Consortium for Translational Cancer Research, P.S. for funding from SAP, J.M. for funding from the German Science Foundation (DFG-SFB924) and M.W. for funding from the ProteomeTools project (BMBF grant no. 031L0008A).
Author information
Authors and Affiliations
Contributions
A.J. planned and performed experiments, contributed to data analysis, interpretation and manuscript writing. N.K. contributed to data analysis, interpretation and manuscript writing, wrote the shiny app. T.H. contributed to data analysis, interpretation and manuscript writing. M. Moerch contributed to planning and performing experiments. J.Z. performed and analyzed experiments. N.L. performed experiments. Y.B. contributed to data acquisition. E.M. and M. Maschberger performed experiments. G.S. contributed to data acquisition. I.B. and C.D. performed experiments. P.S. contributed to building the data repository ProteomicsDB. J.M. performed experiments. B.S. performed experiments. A.A. contributed to data analysis and interpretation. T.W. performed experiments. M.B. contributed to data analysis and interpretation. M.W. contributed to building the data repository ProteomicsDB. M.K. contributed to data analysis and interpretation. S.L. contributed to data analysis and interpretation. W.L. contributed to data analysis and interpretation. H.H. conceived the idea, supervised the studies, contributed to data interpretation and manuscript writing. M.M.S. conceived the idea, supervised the studies and contributed to data interpretation and manuscript writing. B.K. conceived the idea, supervised the studies and contributed to data interpretation and manuscript writing.
Corresponding authors
Ethics declarations
Competing interests
H.H. is cofounder, shareholder and CEO of OmicScouts GmbH. M.W. and B.K. are cofounders and shareholders of OmicScouts GmbH and msAId GmbH. M.W. and B.K. have no operational role in either company. T.W. and M.B. are employees and shareholders of GlaxoSmithKline. M.M.S. is a shareholder of GlaxoSmithKline.
Additional information
Peer review information Editor recognition statement: Arunima Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17 and Note.
Supplementary Table 1
Supplementary Table 1
Supplementary Table 2
Supplementary Table 2
Supplementary Table 3
Supplementary Table 3
Supplementary Table 4
Supplementary Table 4
Source data
Source Data Figure 2
Figure 2
Source Data Figure 3
Figure 3
Source Data Figure 4
Figure 4
Source Data Figure 5
Figure 5
Source Data Figure 6
Figure 6
Rights and permissions
About this article
Cite this article
Jarzab, A., Kurzawa, N., Hopf, T. et al. Meltome atlas—thermal proteome stability across the tree of life. Nat Methods 17, 495–503 (2020). https://doi.org/10.1038/s41592-020-0801-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0801-4
This article is cited by
-
Tapioca: a platform for predicting de novo protein–protein interactions in dynamic contexts
Nature Methods (2024)
-
Codon language embeddings provide strong signals for use in protein engineering
Nature Machine Intelligence (2024)
-
Chemoproteomic mapping of the glycolytic targetome in cancer cells
Nature Chemical Biology (2023)
-
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Nature Communications (2023)
-
The CDK7 inhibitor CT7001 (Samuraciclib) targets proliferation pathways to inhibit advanced prostate cancer
British Journal of Cancer (2023)