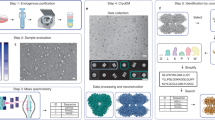
Abstract
Single-particle cryo-electron microscopy (cryo-EM) has become a powerful technique in the field of structural biology. However, the inability to reliably produce pure, homogeneous membrane protein samples hampers the progress of their structural determination. Here, we develop a bottom-up iterative method, Build and Retrieve (BaR), that enables the identification and determination of cryo-EM structures of a variety of inner and outer membrane proteins, including membrane protein complexes of different sizes and dimensions, from a heterogeneous, impure protein sample. We also use the BaR methodology to elucidate structural information from Escherichia coli K12 crude membrane and raw lysate. The findings demonstrate that it is possible to solve high-resolution structures of a number of relatively small (<100 kDa) and less abundant (<10%) unidentified membrane proteins within a single, heterogeneous sample. Importantly, these results highlight the potential of cryo-EM for systems structural proteomics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Atomic coordinates and structure factors have been deposited with accession codes 6WTI (PDB) and EMD-21897 (EMDB) for cytochrome bo3; 6WU0 (PDB) and EMD-21901 (EMDB) for BpHpnN; 6WTZ (PDB) and EMD-21900 (EMDB) for OmpF; 6WU6 (PDB) and EMD-21906 (EMDB) for SQR (3.60 Å); 7JZ3 (PDB) and EMD-22529 (EMDB) for OmpC; 7JZ2 (PDB) and EMD-22528 (EMDB) for SQR (2.50 Å); 7JZ6 (PDB) and EMD-22530 (EMDB) for KatG; and 7JZH (PDB) and EMD-22531 (EMDB) for GadB. Source data are provided with this paper.
References
Vinothkumar, K. R. & Henderson, R. Single particle electron cryomicroscopy: trends, issues and future perspective. Q. Rev. Biophys. 49, 1–25 (2016).
Herzik, M. A.Jr, Wu, M. & Lander, G. C. High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM. Nat. Commun. 10, 1032 (2019).
Ho, C. M. et al. Malaria parasite translocon structure and mechanism of effector export. Nature 561, 70–75 (2018).
Morgan, C. E. et al. Cryo-electron microscopy structure of the Acinetobacter baumannii 70S ribosome and implications for new antibiotic development. mBio 11, e03117–e03119 (2020).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Daligault, H. E. et al. Whole-genome assemblies of 56 Burkholderia species. Genome Announc. 2, e01106–e01114 (2014).
Doughty, D. M. et al. The RND-family transporter, HpnN, is required for hopanoid localization to the outer membrane of Rhodopseudomonas palustris TIE-1. Proc. Natl Acad. Sci. U S A 108, E1045–E1051 (2011).
Sousa, F. L. et al. The superfamily of heme-copper oxygen reductases: types and evolutionary considerations. Biochim. Biophys. Acta 1817, 629–637 (2012).
Abramson, J. et al. The structure of the ubiquinol oxidase from Escherichia coli and its ubiquinone binding site. Nat. Struct. Biol. 7, 910–917 (2000).
Yap, L. L. et al. The quinone-binding sites of the cytochrome bo3 ubiquinol oxidase from Escherichia coli. Biochim. Biophys. Acta 1797, 1924–1932 (2010).
Choi, S. K. et al. Location of the substrate binding site of the cytochrome bo3 ubiquinol oxidase from Escherichia coli. J. Am. Chem. Soc. 139, 8346–8354 (2017).
Kumar, N. et al. Crystal structures of the Burkholderia multivorans hopanoid transporter HpnN. Proc. Natl Acad. Sci. U S A 114, 6557–6562 (2017).
Centers for Disease Control and Prevention. Bioterrorism agents/diseases (U.S. Department of Health and Human Services, 2018); https://emergency.cdc.gov/agent/agentlist-category.asp
Wagar, E. Bioterrorism and the role of the clinical microbiology laboratory. Clin. Microbiol. Rev. 29, 175–189 (2016).
Christopher, G. W., Cieslak, T. J., Pavlin, J. A. & Eitzen, E. M.Jr Biological warfare. A historical perspective. JAMA 278, 412–417 (1997).
Nierman, W. C. et al. Structural flexibility in the Burkholderia mallei genome. Proc. Natl Acad. Sci. U S A 101, 14246–14251 (2004).
Schweizer, H. P. Mechanisms of antibiotic resistance in Burkholderia pseudomallei: implications for treatment of melioidosis. Future Microbiol. 7, 1389–1399 (2012).
Malott, R. J., Steen-Kinnaird, B. R., Lee, T. D. & Speert, D. P. Identification of hopanoid biosynthesis genes involved in polymyxin resistance in Burkholderia multivorans. Antimicrob. Agents Chemother. 56, 464–471 (2012).
Malott, R. J. et al. Fosmidomycin decreases membrane hopanoids and potentiates the effects of colistin on Burkholderia multivorans clinical isolates. Antimicrob. Agents Chemother. 58, 5211–5219 (2014).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656 (2003).
Cowan, S. W. et al. Crystal structures explain functional properties of two E. coli porins. Nature 358, 727–733 (1992).
Yankovskaya, V. et al. Architecture of succinate dehydrogenase and reactive oxygen species generation. Science 299, 700–704 (2003).
Baslé, A., Rummel, G., Storici, P., Rosenbusch, J. P. & Schirmer, T. Crystal structure of osmoporin OmpC from E. coli at 2.0 Å. J. Mol. Biol. 362, 933–942 (2006).
Carpena, X., Melik-Adamyan, W., Loewen, P. C. & Fita, I. Structure of the C-terminal domain of the catalase–peroxidase KatG from Escherichia coli. Acta Crystallogr. D Biol. Crystallogr. 60, 1824–1832 (2004).
Capitani, G. et al. Crystal structure and functional analysis of Escherichia coli glutamate decarboxylase. EMBO J. 22, 4027–4037 (2003).
Chorev, D. S. et al. Protein assemblies ejected directly from native membranes yield complexes for mass spectrometry. Science 362, 829–834 (2018).
Ho, C. M. et al. Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu. Nat. Methods 17, 79–85 (2020).
Kastritis, P. L. et al. Capturing protein communities by structural proteomics in a thermophilic eukaryote. Mol. Syst. Biol. 13, 936 (2017).
Yi, X., Verbeke, E. J., Chang, Y., Dickinson, D. J. & Taylor, D. W. Electron microscopy snapshots of single particles from single cells. J. Biol. Chem. 294, 1602–1608 (2019).
Schmidli, C. et al. Microfluidic protein isolation and sample preparation for high-resolution cryo-EM. Proc. Natl Acad. Sci. U S A 116, 15007–15012 (2019).
Long, F. et al. Crystal structures of the CusA efflux pump suggest methionine-mediated metal transport. Nature 467, 484–488 (2010).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Terwilliger, T. C., Ludtke, S. J., Read, R. J., Adams, P. D. & Afonine, P. V. Improvement of cryo-EM maps by density modification. Nat. Methods 17, 923–927 (2020).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Acknowledgements
We thank P. A. Klenotic for proofreading the manuscript. We are grateful to the Cryo-Electron Microscopy Core at the CWRU School of Medicine and K. Li for access to the sample preparation and Cryo-EM instrumentation. We thank D. Wu and T. El-Baba for help with proteomics analysis. This work was supported by NIH grant R01AI145069 (E.W.Y.) and MRC grant MR/N020413/1 (C.V.R.). This research was supported in part by the National Cryo-EM Facility of the National Cancer Institute at the Frederick National Laboratory for Cancer Research under contract HSSN261200800001E.
Author information
Authors and Affiliations
Contributions
C.-C.S., M.L., C.E.M. and E.W.Y. designed the cryo-EM experiments. J.R.B. and C.V.R. designed the nMS and proteomics experiments. C.-C.S., M.L. and C.E.M. made the BpHpnN, crude cell membrane and raw cell lysate samples and performed the cryo-EM experiments. J.R.B. performed the nMS and proteomics experiments. E.W.Y. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Arunima Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Flowchart of the ‘Build and Retrieve’ (BaR) iterative method.
Extended Data Fig. 2 Native mass spectrometry (nMS) and proteomics analysis suggest that the sample of B. pseudomallei HpnN was co-purified with several other proteins.
(a) SDS–PAGE of the purified sample where the gel bands were sliced and subjected to tryptic digestion to identify the proteins. Proteomics analysis clearly indicated the presence of HpnN, OmpF and different components of succinate dehydrogenase (SdhA and SdhB) and cytochrome bo3 oxidase (Subunit I, Subunit II and Subunit III). The protein bands were quantified and replicated three times using ImageJ (imagej.net), showing the abundance of 60.8% for BpHpnN, 9.4% for SdhA + GlmS, 9.7% for subunit I, 6.4% for ArnC + subunit II+PfkA + GatD + TDH+ OmpF, 6.7% for SdhB and 1.9% for subunit III. (b) Native mass spectra show several charge state distributions whose deconvoluted masses are shown in the Table. Among them, the 93,548 Da can be readily assigned to monomeric HpnN and the 133,853 Da to dimeric glucosamine-6-phosphate synthase (GlmS). While the masses 65,233 Da and 75,031 Da can be assigned to SdhA bound to FAD and Subunit I bound to heme b respectively based on the data from (a). Among the proteins identified in (a), PfkA and TDH are known to exist as tetramers. Therefore, the masses 139,385 Da and 149,545 Da can be assigned to the PfkA and TDH tetramers. The collisionally induced dissociated products shown in (c) further supported this assignment.
Extended Data Fig. 3 2D classes of the cryo-EM images.
The 2D classification indicates that there are at least five different proteins coexist in the nanodisc sample.
Extended Data Fig. 4 Cryo-EM analysis of the E. coli cytochrome bo3 complex.
(a) BaR processing flowchart. (b) Representative 2D classes. (c) Fourier Shell Correlation (FSC) curves. (d) Sharpened cryo-EM map of the cytochrome bo3 complex viewed in the membrane plane. (e) Sharpened cryo-EM map of the cytochrome bo3 complex viewed from the cytoplasmic side. (f) Local EM density map of cytochrome bo3.
Extended Data Fig. 5 Cryo-EM analysis of the B. pseudomallei HpnN transporter.
(a) BaR processing flowchart. (b) Representative 2D classes. (c) Fourier Shell Correlation (FSC) curves. (d and e) Sharpened cryo-EM maps of the BpHpnN transporter viewed in the membrane plane. (f) Local EM density map of BpHpnN.
Extended Data Fig. 6 Cryo-EM analysis of the E. coli OmpF porin channel.
(a) BaR processing flowchart. (b) Representative 2D classes. (c) Fourier Shell Correlation (FSC) curves. (d) Sharpened cryo-EM map of the OmpF porin viewed in the membrane plane and from the periplasmic side. (e) Sharpened cryo-EM map of the OmpF viewed from the exterior side. (f) Local EM density map of OmpF.
Extended Data Fig. 7 Cryo-EM analysis of the E. coli SQR complex.
(a) BaR processing flowchart. (b) Representative 2D classes. (c) Fourier Shell Correlation (FSC) curves. (d) Sharpened cryo-EM map of the SQR complex viewed in the membrane plane. (e) Sharpened cryo-EM map of the SQR complex viewed from the cytoplasmic side. (f) Local EM density map of SQR.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Supplementary Tables 1–5.
Source data
Source Data Extended Data Fig. 2
Full scan image for Extended Data Figure 2a.
Rights and permissions
About this article
Cite this article
Su, CC., Lyu, M., Morgan, C.E. et al. A ‘Build and Retrieve’ methodology to simultaneously solve cryo-EM structures of membrane proteins. Nat Methods 18, 69–75 (2021). https://doi.org/10.1038/s41592-020-01021-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-01021-2
This article is cited by
-
Bridging structural and cell biology with cryo-electron microscopy
Nature (2024)
-
Broadening environmental research in the era of accurate protein structure determination and predictions
Frontiers of Environmental Science & Engineering (2024)
-
Exploring ND-011992, a quinazoline-type inhibitor targeting quinone reductases and quinol oxidases
Scientific Reports (2023)
-
Cryo-EM structure of TMEM63C suggests it functions as a monomer
Nature Communications (2023)
-
DNA is loaded through the 9-1-1 DNA checkpoint clamp in the opposite direction of the PCNA clamp
Nature Structural & Molecular Biology (2022)