Abstract
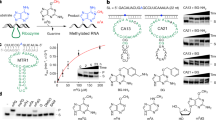
Chemical modifications to messenger RNA are increasingly recognized as a critical regulatory layer in the flow of genetic information, but quantitative tools to monitor RNA modifications in a whole-transcriptome and site-specific manner are lacking. Here we describe a versatile platform for directed evolution that rapidly selects for reverse transcriptases that install mutations at sites of a given type of RNA modification during reverse transcription, allowing for site-specific identification of the modification. To develop and validate the platform, we evolved the HIV-1 reverse transcriptase against N1-methyladenosine (m1A). Iterative rounds of selection yielded reverse transcriptases with both robust read-through and high mutation rates at m1A sites. The optimal evolved reverse transcriptase enabled detection of well-characterized m1A sites and revealed hundreds of m1A sites in human mRNA. This work develops and validates the reverse transcriptase evolution platform, and provides new tools, analysis methods and datasets to study m1A biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw and processed m1A-IP-seq and m1A-quant-seq data are available at NCBI Gene Expression Omnibus, accession number GSE123365. The DNA sequence of RT-1306 is shown in Supplementary Table 1, and the plasmid for bacterial expression of RT-1306 is available on Addgene with the ID 131521. The data that support the findings of this study are available from the corresponding author upon request.
Code availability
Processing scripts for synthetic m1A oligonucleotide library, m1A-IP-seq and m1A-quant-seq are available in the Supplementary Data.
Change history
02 October 2019
When this Article was originally published, Supplementary Tables 1–5 were missing; they have now been added.
References
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Frye, M., Harada, B. T., Behm, M. & He, C. RNA modifications modulate gene expression during development. Science 361, 1346 (2018).
Jonkhout, N. et al. The RNA modification landscape in human disease. RNA 23, 1754–1769 (2017).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Dominissini, D. et al. The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature 530, 441 (2016).
Li, X., Xiong, X. & Yi, C. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods 14, 23 (2016).
Li, X. et al. Transcriptome-wide mapping reveals reversible and dynamic N1-methyladenosine methylome. Nat. Chem. Biol. 12, 311 (2016).
Zheng, G. et al. Efficient and quantitative high-throughput tRNA sequencing. Nat. Methods 12, 835 (2015).
Cozen, A. E. et al. ARM-seq: AlkB-facilitated RNA methylation sequencing reveals a complex landscape of modified tRNA fragments. Nat. Methods 12, 879 (2015).
Li, X. et al. Base-resolution mapping reveals distinct m1A methylome in nuclear- and mitochondrial-encoded transcripts. Mol. Cell 68, 993–1005.e1009 (2017).
Schwartz, S. & Motorin, Y. Next-generation sequencing technologies for detection of modified nucleotides in RNAs. RNA Biol. 14, 1124–1137 (2017).
Mohr, S. et al. Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing. RNA 19, 958–970 (2013).
Zhao, C., Liu, F. & Pyle, A. M. An ultraprocessive, accurate reverse transcriptase encoded by a metazoan group II intron. RNA 24, 183–195 (2018).
Ellefson, J. W. et al. Synthetic evolutionary origin of a proofreading reverse transcriptase. Science 352, 1590 (2016).
Huber, C., von Watzdorf, J. & Marx, A. 5-methylcytosine-sensitive variants of Thermococcus kodakaraensis DNA polymerase. Nucleic Acids Res. 44, 9881–9890 (2016).
Harcourt, E. M., Ehrenschwender, T., Batista, P. J., Chang, H. Y. & Kool, E. T. Identification of a selective polymerase enables detection of N6-methyladenosine in RNA. J. Am. Chem. Soc. 135, 19079–19082 (2013).
Helm, M., Giegé, R. & Florentz, C. A Watson−Crick base-pair-disrupting methyl group (m1A9) is sufficient for cloverleaf folding of human mitochondrial tRNALys. Biochemistry 38, 13338–13346 (1999).
Anderson, J. T. & Droogmans, L. in Fine-Tuning of RNA Functions by Modification and Editing (ed Grosjean, H.) 121–139 (Springer Berlin Heidelberg, 2005).
Saikia, M., Fu, Y., Pavon-Eternod, M., He, C. & Pan, T. Genome-wide analysis of N1-methyl-adenosine modification in human tRNAs. RNA 16, 1317–1327 (2010).
Safra, M. et al. The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature 551, 251 (2017).
Xu, L. et al. Three distinct 3-methylcytidine (m3C) methyltransferases modify tRNA and mRNA in mice and humans. J. Biol. Chem. 292, 14695–14703 (2017).
Patterson, J. T., Nickens, D. G. & Burke, D. H. HIV-1 reverse transcriptase pausing at bulky 2′ adducts is relieved by deletion of the rnase H domain. RNA Biol. 3, 163–169 (2006).
Kohlstaedt, L. A., Wang, J., Friedman, J. M., Rice, P. A. & Steitz, T. A. Crystal structure at 3.5 A resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 256, 1783 (1992).
Huang, H., Chopra, R., Verdine, G. L. & Harrison, S. C. Structure of a Covalently Trapped Catalytic Complex of HIV-1 Reverse Transcriptase: Implications for Drug Resistance. Science 282, 1669 (1998).
Müller, B. et al. Co-expression of the subunits of the heterodimer of HIV-1 reverse transcriptase in Escherichia coli. J. Biol. Chem. 264, 13975–13978 (1989).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Chen, T. et al. Evolution of thermophilic DNA polymerases for the recognition and amplification of C2ʹ-modified DNA. Nat. Chem. 8, 556 (2016).
Betz, K. et al. Structural insights into dna replication without hydrogen bonds. J. Am. Chem. Soc. 135, 18637–18643 (2013).
Diaz, A., Nellore, A. & Song, J. S. CHANCE: comprehensive software for quality control and validation of ChIP-seq data. Genome Biol. 13, R98 (2012).
Fang, S. et al. NONCODEV5: a comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 46, D308–D314 (2017).
Shi, H., Wei, J. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Dominissini, D. & Rechavi, G. Loud and clear epitranscriptomic m1A signals: now in single-base resolution. Mol. Cell 68, 825–826 (2017).
Xiong, X., Li, X., Wang, K. & Yi, C. Perspectives on topology of the human m1A methylome at single nucleotide resolution. RNA 24, 1437–1442 (2018).
Sas-Chen, A. & Schwartz, S. Misincorporation signatures for detecting modifications in mRNA: Not as simple as it sounds. Methods 156, 53–59 (2019).
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75 (2016).
Guo, J. U. & Bartel, D. P. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science 353, (2016).
Le Grice, S. F. J., Cameron, C. E. & Benkovic, S. J. in Methods in Enzymology Vol. 262 (ed. Campbell, J.) 130–144 (Acad. Press, 1995).
Lee, T. S. et al. BglBrick vectors and datasheets: A synthetic biology platform for gene expression. J. Biol. Eng. 5, 12 (2011).
Mishina, Y., Chen, L. X. & He, C. Preparation and characterization of the native iron(II)-containing dna repair AlkB protein directly from Escherichia coli. J. Am. Chem. Soc. 126, 16930–16936 (2004).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 3 (2011).
Bushnell, B. BBMap Package (SourceForge, 2019); https://sourceforge.net/projects/bbmap/
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Lin, Y.-C. et al. Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations. Nat. Commun. 5, 4767 (2014).
Chen, L. Characterization and comparison of human nuclear and cytosolic editomes. Proc. Natl Acad. Sci. USA 110, E2741 (2013).
Rigby, R. A. & Stasinopoulos, D. M. Generalized additive models for location, scale and shape. J. R. Stat. Soc. Ser. C. (Appl. Stat.) 54, 507–554 (2005).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Bailey, T. L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Cui, X. et al. Guitar: An R/Bioconductor package for gene annotation guided transcriptomic analysis of RNA-related genomic features. BioMed. Res. Int. 2016, 8 (2016).
Acknowledgements
We thank the entire staff at the University of Chicago Comprehensive Cancer Center sequencing facility at for performing the Sanger and NGS sequencing measurements; T. Pan for sharing the ModSig libraries; L. Zhang and C. Liu for sharing the protocols for constructing biological RNA libraries; L. Hu and L. Luo for helping with protein purification. This work was supported by the University of Chicago, National Human Genome Research Institute (RM1 HG008935, C.H. and B.C.D.) and the National Institute of General Medical Sciences (R35 GM119840, B.C.D.) of the National Institutes of Health, the University of Chicago Medicine Comprehensive Cancer Center (P30 CA14599), the Chicago Fellows Program (H.Z.) and the MSTP Training Grant (T32GM007281) (C.S.). S.N. is an HHMI fellow of the Damon Runyon Cancer Research Foundation (DRG-2215-15). C.H. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
B.C.D., C.H., and H.Z. conceived the idea of the project and designed the experiments. H.Z. performed the directed-evolution experiments with assistance from S.R., who performed the initial screening experiment on the Broccoli DNA. H.Z. prepared protein samples and carried out biochemical characterizations, NGS library preparations, NGS data processing and analyses. Q.D. synthesized the RNA oligonucleotides used in this study. S.N. and C.S. assisted with cell culturing for biological RNA library preparations. X.C. and Z.Z. assisted with NGS data analysis pipeline, enrichment and statistical analyses. H.Z., C.H., and B.C.D. wrote the manuscript with critical inputs from S.R., Q.D., X.C., Z.Z., S.N. and C.S.
Corresponding authors
Ethics declarations
Competing interests
H.Z., C.H., and B.C.D. have filed a provisional patent application for the RTs described in this manuscript. C.H. is a scientific founder and a member of the scientific advisory board of Accent Therapeutics, Inc.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Screening of single-site mutations in Broccoli aptamer.
a, Screening results for single-site mutations in Broccoli DNA show various levels of decrease in fluorescence during IVT. Fluorescence traces are color coded for the mutations at different sites as shown in the secondary structure; electrophoresis of IVT products is shown on the right to show similar amount of RNA are produced. b, Test of fluorescence intensities of various mutations at U15 of Broccoli aptamer and at the A36 site that is complementary to U15 based on the secondary structure prediction. c, A zoomed-in view of the fluorophore-binding structure in the reference crystal structure of the Spinach RNA aptamer that is sequentially similar to Broccoli RNA.
Supplementary Figure 2 RT-PCR-IVT assay with purified RT and RT in crude lysates.
a, Results from the validation test of the RT-PCR-IVT assay with purified wild-type HIV-RT p66, with varying numbers of PCR cycles is shown on the left; data shown are mean ± s.e.m. from n = 5 independent assays. 8% SDS-PAGE of the purified p66 subunit is shown on the right. b, RT-PCR-IVT assay data (mean ± s.e.m. from n = 2 independent cell cultures for RT lysates) on the positive (U15) and negative (A15) control RNA; crude lysate activity of wild-type HIV-1 RT p66 show a reliable dynamic range for mutation detection with various PCR cycle numbers. Shown on the right are high expression levels of p66 proteins in over-expressed BL21 cell lysates lysed by boiling or lysozyme treatment analyzed by 8% SDS-PAGE, with n = 2 independent cell cultures. c, Screen plate layout of 90 RT variants and 6 control samples on each 96-deep well plate for the directed evolution. Shown on the right are robust data produced from control experiments (no. of repeats = 16) of wild-type RT lysate with positive (P) U15, negative A15 (N) and m1A15 (B) RNA substrates throughout the first round of directed evolution screening. d, Shown on the left are two repeating assays (n = 3 cell culture replicates) for variants (RT-164, RT-176 and RT-395) that show positive responses against m1A15 RNA in comparison to the basal activity by the wild-type RT. The middle panel shows one repeated test from cell culture for variants (RT-110 and RT-142) that show negative responses in the screen. Protein expression levels in the crude cell lysates of four variants via 8% SDS-PAGE are shown on the right in duplicates. This shows that lack of fluorescence responses in RT-110 and RT-142 is not due to lack of protein expression.
Supplementary Figure 3 In vitro characterization of evolved RT variants.
a, Purified wild-type HIV-1 RT p66 and evolved variants RT-733 and RT-1306 are shown with 8% SDS-PAGE. b, RT-PCR-IVT assay data on the positive (U15) and negative (A15) control RNAs with purified wild-type HIV-1 RT p66, and evolved variants RT-733 and RT-1306; data shown represent mean fluorescence intensities ± s.e.m from n = 5 independent assays. c, Sanger sequencing data showing no detectable mutation signatures on the negative control A15 RNA by the wild-type and evolved RT variants. The A15 position is noted between the dashed lines. d, The overall mutation signatures of wild-type HIV-1 RT over the ModSig-m1A library and the sequence-context-dependent mutation rates at the m1A site.
Supplementary Figure 4 Mutation patterns at m1A sites in sequence contexts.
Heatmaps show mutation patterns (A to T, G, or C) of the wild-type and evolved RT variants in 256 sequence contexts in the ModSig-m1A library, in comparison to those in the control ModSig-A library. Yellow boxes are sequence contexts not covered by the sequencing data.
Supplementary Figure 5 Biochemical assays and optimization of demethylation activity of purified AlkB enzyme against m1A.
a, Shown are LC-MS/MS data of m1A level before and after AlkB treatment for synthetic m1A15 RNA (left; mean ± s.d. from n = 2 technical replicates) and polyA-enriched RNA from HEK293T cells (right; mean ± s.d. from n = 12 replicates — that is, 3 cell culture replicates and 4 LC–MS/MS injection replicates). b, Schematics for the RT-PCR-IVT assay for detecting AlkB activity using m1A18 RNA. Fluorescence data of positive (m1A18 treated by AlkB) and negative control (no AlkB added or AlkB added together with 5mM EDTA) experiments are shown on the right. Error bars represent s.e.m. from n = 2 independent assays for –AlkB and from n = 6 independent assays for +AlkB and +AlkB+EDTA. c, Optimization of the AlkB reaction condition by the RT-PCR-IVT assay for a 2-h reaction at 37 °C. Fluorescence intensities at 60 min are plotted versus various reaction components with the reaction conditions as 50 mM MES (pH 5.0), 50 μM (NH4)2Fe(SO4)2, 300 μM 2-ketoglutarate, 2 mM l-ascorbic acid, 2 mM MgCl2 and RNase inhibitor unless specified on the plot. Error bars represent s.d. from n = 2 or 3 independent assays. d, Optimization of iron concentration and reaction temperatures by the RT-PCR-IVT assay under the condition 50 mM MES (pH 5.0), 300 μM 2-ketoglutarate, 2 mM l-ascorbic acid and RNase inhibitor and noted concentrations of (NH4)2Fe(SO4)2 on the plot. Shown are mean ± s.d. from n = 2 independent assays. e, Optimization of iron concentration and reaction temperatures by LC–MS/MS assay with total RNA sample from HEK293T cells with small RNA removal. Shown are mean ± s.d. from n = 6 replicates — that is, 2 independent AlkB assays and 3 LC–MS/MS injection replicates.
Supplementary Figure 6 Schematics and quality control of m1A-IP-seq and m1A-quant-seq libraries.
a, Schematic of library-construction procedures with evolved RT-130, and the final sequence composition of the library. b, Representative bioanalyzer analyses of NGS libraries built with RT-1306 for one of the three biological replicates. The library size is ~240 bp for all libraries with and without AlkB treatment. Considering the adaptor dimer is 132 bp, the average size of the fragmented insert is ~100 nt. c, Reproducibility examination based on correlation plots of transcriptome-wide coverage between replicates in both m1A-IP-seq and m1A-quant-seq. Each dot represents the coverages of one RefSeq-annotated RNA transcript (n = 38882) counted with ‘bedtools multicov’. At the sequencing data coverage depth of the current experiments (~100 million reads per library), transcriptome-wide coverages show decent reproducibility between replicates evaluated with Pearson’s correlation coefficient (r).
Supplementary Figure 7 Location of mutation signatures within aligned reads.
a, IGV coverage and alignment tracks of three mRNA and lncRNA m1A sites. The alignment tracks show that mutations (labeled with colors, red for T and blue for C) are located distal to the ends of reads. b, Histograms of distributions of the mutation locations within the reads for all m1A sites (n = 614) called from m1A-IP-seq with P < 0.1 based on beta-binomial regression test in each library replicate. The normalized location of each mutation site is analyzed by the ‘bam-readcount’ program, in which 0 represents mutations come from either end of the read, and 1 represents the center of the read. Histograms show that distributions of read locations of mutations for m1A sites are mostly centered in the middle, rather than ends of aligned reads with mean ± s.d. normalized locations of 614 m1A sites in each library replicate.
Supplementary Figure 8 Characterization of antibody enrichment activity by in vitro assays and sequencing data.
a, Dot blot of m1A antibody (MBL 345–3) against short 5-mer m1A and control m6A and synthetic A oligonucleotides. b, LC–MS/MS assay for quantifying m1A level from biological RNA sample before and after immunoprecipitation with the anti-m1A antibody. Quantified percentages of m1A and m6A in reference to A are shown by mean ± s.d. of n = 3 LC–MS/MS injection replicates. c, Overlay of coverage tracks of 28S rRNA between m1A-IP-seq (‘IP’) and m1A-quant-seq libraries (‘quant’). ~7-fold coverage enrichment is observed at the m1A1322 site together with mutation signatures. d, Overlays of coverage tracks for m1A sites in mRNA and lncRNA. Coverage enrichment of IP libraries is observed for ND5 and MALAT1 sites, however, not for the PRUNE site. e, Lorenz curve analysis of IP libraries as compared to quant libraries without AlkB treatment. Replicate 1 in the m1A-quant-seq is set as the reference for RNA expression level in HEK293T cells; the diagonal line suggests uniform coverage relative to the reference, and more deviated curves suggest more biased coverage distribution. IP libraries show significant biases relative to the quant libraries.
Supplementary Figure 9 Data processing pipeline used in the current study.
Shown processing scripts are reported in the Supplementary Data.
Supplementary Figure 10 Statistical features of identified m1A sites from m1A-IP-seq.
a, Enriched motif found surrounding n = 281 m1A sites identified in m1A-IP-seq, with P < 0.05 based on the beta-binomial regression test in mRNA and lncRNA. The enrichment score of the shown motif has an E-value of 9.2 × 10–5, calculated by the MEME Suite program (Methods), contributed from 68 out of 281 input sequences. b, Distribution of transcript localizations of 215 m1A sites in mRNA is shown by the pie chart and Metagene plot processed by the ‘Guitar’ R package (Methods).
Supplementary Figure 11 Comparisons of mutation signatures captured by targeted library and m1A-quant-seq.
a, Number of sites identified by m1A-quant-seq, with P < 0.05 based on the beta binomial statistical test. b, Mutations captured by the targeted library for PRUNE and MALAT1 m1A sites are shown in IGV coverage traces, with and without AlkB treatment. c, Comparisons of mutation rates and AlkB sensitivities for the PRUNE and MALAT1 sites detected by targeted library and m1A-quant-seq. Error bars represent the s.d. of n = 3 technical targeted library replicates for targeted libraries and n = 3 cell culture replicates for m1A-quant-seq overlaid, with individual data points shown by dots.
Supplementary Figure 12 Data comparisons between current and previously reported datasets.
a, Comparison of transcriptome coverages (mean ± s.d.), with coverage depth per transcript cut-offs at 100 reads and 10 reads. Error bars represent the s.d. from n = 3 library replicates for RT1306, and from n = 2 library replicates for TGIRT1 and TGIRT2. b, Venn diagram of transcripts that are covered by at least 100 reads. c, Correlation plot of sequencing depth per transcript between one replicate in m1A-quant-seq and one replicate in TGIRT111 or TGIRT219 sequencing data without antibody enrichment, evaluated by Pearson’s r. Plotted are total n = 38882 annotated RNA transcripts in the RefSeq database. d, Comparison of mutation signatures captured by RT1306 and TGIRT1 without and with m1A antibody enrichment. e, Correlation plot and Pearson’s r values of mutation rates of overlapping mutation sites between RT1306 and TGIRT1, without or with m1A antibody enrichment. f, Comparison of mutation signatures that are AlkB sensitive (averaged demethylation ratio (DMR) > 10%) between RT1306 and TGIRT1 with m1A antibody enrichment, evaluated by Pearson’s r. Shown are Venn diagrams for overlapping of AlkB-sensitive mutation sites between the two datasets, correlation plot of DMR of 340 overlapping mutation sites and a Venn diagram of antibody-enriched peaks called by MeRIPtools with fold of enrichment (FC) > 2 (Methods) for RT1306 and TGIRT1.
Supplementary Information
Supplementary Information
Supplementary Figs. 1–12, Supplementary Notes and Supplementary References.
Supplementary Tables
Supplementary Tables 1–5.
Rights and permissions
About this article
Cite this article
Zhou, H., Rauch, S., Dai, Q. et al. Evolution of a reverse transcriptase to map N1-methyladenosine in human messenger RNA. Nat Methods 16, 1281–1288 (2019). https://doi.org/10.1038/s41592-019-0550-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0550-4
This article is cited by
-
MSCAN: multi-scale self- and cross-attention network for RNA methylation site prediction
BMC Bioinformatics (2024)
-
Research progress of N1-methyladenosine RNA modification in cancer
Cell Communication and Signaling (2024)
-
Age-related noncanonical TRMT6–TRMT61A signaling impairs hematopoietic stem cells
Nature Aging (2024)
-
BID-seq for transcriptome-wide quantitative sequencing of mRNA pseudouridine at base resolution
Nature Protocols (2024)
-
RNA modifications in physiology and disease: towards clinical applications
Nature Reviews Genetics (2024)