Abstract
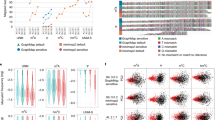
Replication of eukaryotic genomes is highly stochastic, making it difficult to determine the replication dynamics of individual molecules with existing methods. We report a sequencing method for the measurement of replication fork movement on single molecules by detecting nucleotide analog signal currents on extremely long nanopore traces (D-NAscent). Using this method, we detect 5-bromodeoxyuridine (BrdU) incorporated by Saccharomyces cerevisiae to reveal, at a genomic scale and on single molecules, the DNA sequences replicated during a pulse-labeling period. Under conditions of limiting BrdU concentration, D-NAscent detects the differences in BrdU incorporation frequency across individual molecules to reveal the location of active replication origins, fork direction, termination sites, and fork pausing/stalling events. We used sequencing reads of 20–160 kilobases to generate a whole-genome single-molecule map of DNA replication dynamics and discover a class of low-frequency stochastic origins in budding yeast. The D-NAscent software is available at https://github.com/MBoemo/DNAscent.git.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw and processed Illumina and MinION data are available from NCBI GEO under accession number GSE121941.
Code availability
The D-NAscent software can be accessed at https://github.com/MBoemo/DNAscent.git.
References
Gilbert, D. M. Evaluating genome-scale approaches to eukaryotic DNA replication. Nat. Rev. Genet. 11, 673–684 (2010).
Mechali, M. Eukaryotic DNA replication origins: many choices for appropriate answers. Nat. Rev. Mol. Cell Biol. 11, 728–738 (2010).
Raghuraman, M. K. et al. Replication dynamics of the yeast genome. Science 294, 115–121 (2001).
Hawkins, M. et al. High-resolution replication profiles define the stochastic nature of genome replication initiation and termination. Cell Rep. 5, 1132–1141 (2013).
Tuduri, S., Tourriere, H. & Pasero, P. Defining replication origin efficiency using DNA fiber assays. Chromosome Res. 18, 91–102 (2010).
Lacroix, J. et al. Analysis of DNA replication by optical mapping in nanochannels. Small. 12, 5963–5970 (2016).
De Carli, F. et al. High‐throughput optical mapping of replicating DNA. Small Methods 2, 1800146 (2018).
Simpson, J. T. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat. Methods 14, 407–410 (2017).
Rand, A. C. et al. Mapping DNA methylation with high-throughput nanopore sequencing. Nat. Methods 14, 411–413 (2017).
Hagenkort, A. et al. dUTPase inhibition augments replication defects of 5-Fluorouracil. Oncotarget 8, 23713–23726 (2017).
Hua, H. & Kearsey, S. E. Monitoring DNA replication in fission yeast by incorporation of 5-ethynyl-2´-deoxyuridine. Nucl. Acids Res. 39, e60 (2011).
Diermeier-Daucher, S. et al. Cell type specific applicability of 5-ethynyl-2´-deoxyuridine (EdU) for dynamic proliferation assessment in flow cytometry. Cytometry. A. 75, 535–546 (2009).
Talarek, N., Petit, J., Gueydon, E. & Schwob, E. EdU incorporation for FACS and microscopy analysis of DNA replication in budding yeast. Methods Mol. Biol. 1300, 105–112 (2015).
Vernis, L., Piskur, J. & Diffley, J. F. Reconstitution of an efficient thymidine salvage pathway in Saccharomyces cerevisiae. Nucl. Acids Res. 31, e120 (2003).
Peace, J. M., Villwock, S. K., Zeytounian, J. L., Gan, Y. & Aparicio, O. M. Quantitative BrdU immunoprecipitation method demonstrates that Fkh1 and Fkh2 are rate-limiting activators of replication origins that reprogram replication timing in G1 phase. Genome Res. 26, 365–375 (2016).
Loman, N. J., Quick, J. & Simpson, J. T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 12, 733–735 (2015).
Magiera, M. M., Gueydon, E. & Schwob, E. DNA replication and spindle checkpoints cooperate during S phase to delay mitosis and preserve genome integrity. J. Cell. Biol. 204, 165–175 (2014).
Alvino, G. M. et al. Replication in hydroxyurea: it’s a matter of time. Mol. Cell. Biol. 27, 6396–6406 (2007).
Feng, W. et al. Genomic mapping of single-stranded DNA in hydroxyurea-challenged yeasts identifies origins of replication. Nat. Cell Biol. 8, 148–155 (2006).
Poli, J. et al. dNTP pools determine fork progression and origin usage under replication stress. EMBO J. 31, 883–894 (2012).
Siow, C. C., Nieduszynska, S. R., Muller, C. A. & Nieduszynski, C. A. OriDB, the DNA replication origin database updated and extended. Nucl. Acids Res. 40, D682–D686 (2012).
Müller, C. A. et al. The dynamics of genome replication using deep sequencing. Nucl. Acids Res. 42, e3 (2014).
Koc, A., Wheeler, L. J., Mathews, C. K. & Merrill, G. F. Hydroxyurea arrests DNA replication by a mechanism that preserves basal dNTP pools. J. Biol. Chem. 279, 223–230 (2004).
Pasero, P., Bensimon, A. & Schwob, E. Single-molecule analysis reveals clustering and epigenetic regulation of replication origins at the yeast rDNA locus. Genes Dev. 16, 2479–2484 (2002).
Cavanagh, B. L., Walker, T., Norazit, A. & Meedeniya, A. C. Thymidine analogues for tracking DNA synthesis. Molecules 16, 7980–7993 (2011).
Kee, N., Sivalingam, S., Boonstra, R. & Wojtowicz, J. M. The utility of Ki-67 and BrdU as proliferative markers of adult neurogenesis. J. Neurosci. Methods 115, 97–105 (2002).
Dolken, L. et al. High-resolution gene expression profiling for simultaneous kinetic parameter analysis of RNA synthesis and decay. RNA 14, 1959–1972 (2008).
Jain, M., Olsen, H. E., Paten, B. & Akeson, M. The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome. Biol. 17, 239 (2016).
Sekedat, M. D. et al. GINS motion reveals replication fork progression is remarkably uniform throughout the yeast genome. Mol. Syst. Biol. 6, 353 (2010).
On, K. F. et al. Prereplicative complexes assembled in vitro support origin-dependent and independent DNA replication. EMBO J. 33, 605–620 (2014).
Gros, J., Devbhandari, S. & Remus, D. Origin plasticity during budding yeast DNA replication in vitro. EMBO J. 33, 621–636 (2014).
Payne, A., Holmes, N., Rakyan, V. & Loose, M. BulkVis: a graphical viewer for Oxford nanopore bulk FAST5 files. Bioinformatics https://doi.org/10.1093/bioinformatics/bty841 (2018).
Chen, C. et al. Single-cell whole-genome analyses by Linear Amplification via Transposon Insertion (LIANTI). Science 356, 189–194 (2017).
Dileep, V. & Gilbert, D. M. Single-cell replication profiling to measure stochastic variation in mammalian replication timing. Nat. Commun. 9, 427 (2018).
Natsume, T. et al. Kinetochores coordinate pericentromeric cohesion and early DNA replication by Cdc7-Dbf4 kinase recruitment. Mol. Cell 50, 661–674 (2013).
Fang, D. et al. Dbf4 recruitment by forkhead transcription factors defines an upstream rate-limiting step in determining origin firing timing. Genes Dev. 31, 2405–2415 (2017).
Hiraga, S. I. et al. Human RIF1 and protein phosphatase 1 stimulate DNA replication origin licensing but suppress origin activation. EMBO Rep. 18, 403–419 (2017).
Foti, R. et al. Nuclear architecture organized by Rif1 underpins the replication-timing program. Mol. Cell 61, 260–273 (2016).
Donnianni, R. A. & Symington, L. S. Break-induced replication occurs by conservative DNA synthesis. Proc. Natl Acad. Sci. USA 110, 13475–13480 (2013).
Saini, N. et al. Migrating bubble during break-induced replication drives conservative DNA synthesis. Nature 502, 389–392 (2013).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Suzuki, H. & Kasahara, M. Acceleration of nucleotide semi-global alignment with adaptive banded dynamic programming. Preprint at https://www.biorxiv.org/content/10.1101/130633v2 (2017).
Moreno, A. et al. Unreplicated DNA remaining from unperturbed S phases passes through mitosis for resolution in daughter cells. Proc. Natl Acad. Sci. USA 113, E5757–E5764 (2016).
Ge, X. Q., Jackson, D. A. & Blow, J. J. Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. Genes Dev. 21, 3331–3341 (2007).
Ibarra, A., Schwob, E. & Mendez, J. Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc. Natl Acad. Sci. USA 105, 8956–8961 (2008).
Ge, X. Q. & Blow, J. J. Chk1 inhibits replication factory activation but allows dormant origin firing in existing factories. J. Cell. Biol. 191, 1285–1297 (2010).
Besnard, E. et al. Unraveling cell type-specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Nat. Struct. Mol. Biol. 19, 837–844 (2012).
Conti, C. et al. Replication fork velocities at adjacent replication origins are coordinately modified during DNA replication in human cells. Mol. Biol. Cell. 18, 3059–3067 (2007).
Jackson, D. A. & Pombo, A. Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J. Cell. Biol. 140, 1285–1295 (1998).
Müller, C. A. & Nieduszynski, C. A. DNA replication timing influences gene expression level. J. Cell. Biol. 216, 1907–1914 (2017).
Demczuk, A. et al. Regulation of DNA replication within the immunoglobulin heavy-chain locus during B cell commitment. PLoS Biol. 10, e1001360 (2012).
Gabrieli, T. et al. Selective nanopore sequencing of human BRCA1 by Cas9-assisted targeting of chromosome segments (CATCH). Nucl. Acids Res. 46, e87 (2018).
Jiang, W. et al. Cas9-Assisted Targeting of CHromosome segments CATCH enables one-step targeted cloning of large gene clusters. Nat. Commun. 6, 8101 (2015).
Acknowledgements
We are grateful to J. Diffley (Francis Crick Institute, UK) and E. Schwob (Institut de Génétique Moléculaire de Montpellier, France) for kindly providing strains; A. Williams and R. Busby for Illumina NextSeq support; M. Maj for assistance with flow cytometry; J. Caesar for IT infrastructure support; and Microsoft and NVIDIA for computing resources. We thank Nieduszynski group members for helpful discussion and advice, and A. Carr, A. Donaldson, S. Hiraga and D. Sherratt for critical reading of the manuscript. This work was supported by Biotechnology and Biological Sciences Research Council grant BB/N016858/1 (to C.A.M. and C.A.N.) and Wellcome Trust Investigator Award 110064/Z/15/Z (to C.A.N.). J.T.S. is supported by the Ontario Institute for Cancer Research through funds provided by the Government of Ontario and the Government of Canada through Genome Canada and Ontario Genomics (OGI-136). P.S. is funded by a Medical Research Council studentship. P.S. and S.K. are funded by Ludwig Cancer Research. C.A.M is a Queen’s College Extraordinary Junior Research Fellow in Physiology. M.A.B. is a St. Cross College Emanoel Lee Junior Research Fellow.
Author information
Authors and Affiliations
Contributions
C.A.N., C.A.M. and M.A.B. designed the study. C.A.M. designed and performed the experiments. M.A.B. designed and implemented the analoge training and detection software. C.A.M. and M.A.B. undertook data analysis. P.S. undertook mass spectrometry, supervised by B.M.K. and S.K. C.A.N. and J.T.S. supervised the study. C.A.N., C.A.M. and M.A.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J.T.S. receives research funding from ONT and has received travel support to attend and speak at meetings organized by ONT.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 and Supplementary Tables 2 and 3
Supplementary Table 1
Information relating to D-NAscent, mass spectrometry and BrdU-seq samples.
Rights and permissions
About this article
Cite this article
Müller, C.A., Boemo, M.A., Spingardi, P. et al. Capturing the dynamics of genome replication on individual ultra-long nanopore sequence reads. Nat Methods 16, 429–436 (2019). https://doi.org/10.1038/s41592-019-0394-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0394-y
This article is cited by
-
Detection of ribonucleotides embedded in DNA by Nanopore sequencing
Communications Biology (2024)
-
Genome-wide measurement of DNA replication fork directionality and quantification of DNA replication initiation and termination with Okazaki fragment sequencing
Nature Protocols (2023)
-
Long-read sequencing in the era of epigenomics and epitranscriptomics
Nature Methods (2023)
-
DeepRepeat: direct quantification of short tandem repeats on signal data from nanopore sequencing
Genome Biology (2022)
-
Purification of mammalian telomeric DNA for single-molecule analysis
Nature Protocols (2022)