Abstract
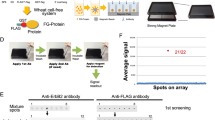
Western blotting (WB) is widely used to test antibody specificity, but the assay has low throughput and precision. Here we used preparative gel electrophoresis to develop a capture format for WB. Fractions with soluble, size-separated proteins facilitated parallel readout with antibody arrays, shotgun mass spectrometry (MS) and immunoprecipitation followed by MS (IP-MS). This pipeline provided the means for large-scale implementation of antibody validation concepts proposed by an international working group on antibody validation (IWGAV).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Supplementary Tables 2–9 contain all data, and an overview is presented in Supplementary Table 1. Raw MS data were uploaded to PRIDE under accessions PXD005945 and PXD010510. Line charts with PAGE-MAP data for antibodies that passed validation and MS data for the intended targets were uploaded to https://www.benchsci.com. Identifier data for all antibodies tested in the study can be obtained on reasonable request from the corresponding author.
References
Algenäs, C. et al. Biotechnol. J. 9, 435–445 (2014).
Berglund, L. et al. Mol. Cell. Proteomics 7, 2019–2027 (2008).
Bradbury, A. & Plückthun, A. Nature 518, 27–29 (2015).
Baker, M. Nature 527, 545–551 (2015).
Baker, M. Nature 521, 274–276 (2015).
Lund-Johansen, F. & Browning, M. D. Nat. Methods 14, 215 (2017).
Uhlen, M. et al. Nat. Methods 13, 823–827 (2016).
Witkowski, C. & Harkins, J. J. Vis. Exp. 2009, 1842 (2009).
Slaastad, H. et al. Proteomics. 11, 4578–4582 (2011).
Wu, W. et al. Mol. Cell. Proteomics 8, 245–257 (2009).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Mol. Cell. Proteomics 11, M111.014050 (2012).
Gholami, A. M. et al. Cell Rep. 4, 609–620 (2013).
Rieckmann, J. C. et al. Nat. Immunol. 18, 583–593 (2017).
Marcon, E. et al. Nat. Methods 12, 725–731 (2015).
Mellacheruvu, D. et al. Nat. Methods 10, 730–736 (2013).
Treindl, F. et al. Nat. Commun. 7, 12852 (2016).
Stuchlý, J. et al. Cytometry A 81, 120–129 (2012).
Acknowledgements
The authors thank J. Olweus, K. Tasken, K.-J. Malmberg and E. Marcon for critical reading. HeLa cells were a kind gift from M.S. Rødland (Oslo University Hospital, Oslo, Norway). This work was funded by grants from the KG Jebsen Foundation to the KG Jebsen Centre for Immunotherapy of Cancer and the KG Jebsen Inflammation Research Centre, Helse-Sør-Øst, Novo-Nordisk Foundation, and the Norwegian Research Council.
Author information
Authors and Affiliations
Contributions
K.S., A.M., M.I. and F.T. conducted experiments (PAGE-MAP), analyzed data and prepared figures. S.K. and M.T carried out DigiWest experiments and data analysis. T.K., J.S. and W.Z. analyzed data. T.A.N., M.E.S. and G.A.d.S. performed mass spectrometry experiments. L.H. performed statistical analysis. F.L.-J. designed the experiments, analyzed data, contributed to figure preparation and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.T. is associated with NMI TT GmbH, a company that sells DigiWest analysis as a service. W.Z. is the founder of Seekquence, a company that sells information about research reagents including antibodies.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Antibody sensitivity is linked to reproducibility.
The box plots show signal-to-noise ratios of antibodies that clustered according to specificity in one experiment only or in both experiments. Whiskers indicate lower and upper quartiles. The Venn diagram shows the number of antibodies in each category. Source data: Supplementary Table 2.
Supplementary Figure 2 PAGE-MAP yields reproducible data for antibody sensitivity.
The scatter plot to the left shows signal-to-noise ratios measured for 3,672 antibodies in two biological replicates. The plot to the right shows ratios measured for 1,712 antibodies in two technical replicates. R values indicate Pearson correlations. Source data: Supplementary Table 5.
Supplementary Figure 3 Shotgun MS signals in the cell type used as the source for IP and signal intensity in single-plex PAGE-IP-FCM are predictive for target identification by PAGE-IP-MS.
(a) A subset of beads from each single-plex IP were labeled with streptavidin–phycoerythrin and analyzed by FCM. The box plots show streptavidin fluorescence intensity of beads in IPs where the intended target was identified (blue) or not (red). (b) Shotgun MS signals measured in cell types used as sources for IPs. Blue and red shading indicates results for antibody targets that were detected and not detected by PAGE-IP-MS, respectively. Numbers in parentheses indicate the number of antibodies in each category. Whiskers indicate lower and upper quartiles. Source data: Supplementary Table 7.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Data
Supplementary Table 1
Overview of all supplementary material
Supplementary Table 2
Antibody-array and MS data
Supplementary Table 3
MS source data, MS metadata and assessment of P values for correlations
Supplementary Table 4
PAGE-MAP technical replicates
Supplementary Table 5
Reproducibility of signal-to-noise ratios in PAGE-MAP
Supplementary Table 6
Shotgun MS reproducibility
Supplementary Table 7
PAGE–IP-MS results
Supplementary Table 8
PAGE-MAP versus DigiWest and WB
Supplementary Table 9
List of antibodies that passed validation in PAGE-MAP
Rights and permissions
About this article
Cite this article
Sikorski, K., Mehta, A., Inngjerdingen, M. et al. A high-throughput pipeline for validation of antibodies. Nat Methods 15, 909–912 (2018). https://doi.org/10.1038/s41592-018-0179-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0179-8
This article is cited by
-
A T cell receptor targeting a recurrent driver mutation in FLT3 mediates elimination of primary human acute myeloid leukemia in vivo
Nature Cancer (2023)
-
Immune responses in Omicron SARS-CoV-2 breakthrough infection in vaccinated adults
Nature Communications (2022)
-
Using proteins to test for COVID antibodies
Nature (2022)
-
Titers of antibodies against ancestral SARS-CoV-2 correlate with levels of neutralizing antibodies to multiple variants
npj Vaccines (2022)
-
Towards reproducibility in large-scale analysis of protein–protein interactions
Nature Methods (2021)