Abstract
Brain tumors are among the most lethal and devastating cancers. Their study is limited by genetic heterogeneity and the incompleteness of available laboratory models. Three-dimensional organoid culture models offer innovative possibilities for the modeling of human disease. Here we establish a 3D in vitro model called a neoplastic cerebral organoid (neoCOR), in which we recapitulate brain tumorigenesis by introducing oncogenic mutations in cerebral organoids via transposon- and CRISPR–Cas9-mediated mutagenesis. By screening clinically relevant mutations identified in cancer genome projects, we defined mutation combinations that result in glioblastoma-like and central nervous system primitive neuroectodermal tumor (CNS-PNET)-like neoplasms. We demonstrate that neoCORs are suitable for use in investigations of aspects of tumor biology such as invasiveness, and for evaluation of drug effects in the context of specific DNA aberrations. NeoCORs will provide a valuable complement to the current basic and preclinical models used to study brain tumor biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
22 August 2018
In the originally published paper, the “before” image for the afatinib condition in Fig. 6c was incorrect. Instead of an image displaying a GBM-3 neoplastic organoid before afatinib treatment, this panel showed an image from the GBM-2 control (DMSO) group before treatment. This error has now been corrected in the HTML and PDF versions of the article; the “before, afatinib” panel in Fig. 6c now shows a representative image from the indicated experiment. The color of all error bars in Fig. 6 has also been changed to black, for consistency. All statistical analysis and all conclusions presented in the article are unaffected by this error. Nevertheless, we apologize for the mistake.
References
Ostrom, Q. T. et al. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2009–2013. Neuro Oncol. 18, v1–v75 (2016).
Lui, J. H., Hansen, D. V. & Kriegstein, A. R. Development and evolution of the human neocortex. Cell 146, 18–36 (2011).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Parsons, D. W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011).
Huszthy, P. C. et al. In vivo models of primary brain tumors: pitfalls and perspectives. Neuro Oncol. 14, 979–993 (2012).
Hu, B. et al. Epigenetic activation of WNT5A drives glioblastoma stem cell differentiation and invasive growth. Cell 167, 1281–1295 (2016).
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Hubert, C. G. et al. A three-dimensional organoid culture system derived from human glioblastomas recapitulates the hypoxic gradients and cancer stem cell heterogeneity of tumors found in vivo. Cancer Res. 76, 2465–2477 (2016).
Zhu, Z. et al. Zika virus has oncolytic activity against glioblastoma stem cells. J. Exp. Med. 214, 2843–2857 (2017).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Kelava, I. & Lancaster, M. A. Stem cell models of human brain development. Cell Stem Cell 18, 736–748 (2016).
Clevers, H., Loh, K. M. & Nusse, R. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012 (2014).
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Johnson, J. Z. & Hockemeyer, D. Human stem cell-based disease modeling: prospects and challenges. Curr. Opin. Cell Biol. 37, 84–90 (2015).
Neal, J. T. & Kuo, C. J. Organoids as models for neoplastic transformation. Annu. Rev. Pathol. 11, 199–220 (2016).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Bershteyn, M. et al. Human iPSC-derived cerebral organoids model cellular features of lissencephaly and reveal prolonged mitosis of outer radial glia. Cell Stem Cell 20, 435–449 (2017).
Mariani, J. et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162, 375–390 (2015).
Iefremova, V. et al. An organoid-based model of cortical development identifies non-cell-autonomous defects in Wnt signaling contributing to Miller-Dieker syndrome. Cell Rep. 19, 50–59 (2017).
Louis, D. N. et al. The2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 131, 803–820 (2016).
Zhu, Y. et al. Early inactivation of p53 tumor suppressor gene cooperating with NF1 loss induces malignant astrocytoma. Cancer Cell 8, 119–130 (2005).
Kwon, C.-H. et al. Pten haploinsufficiency accelerates formation of high-grade astrocytomas. Cancer Res. 68, 3286–3294 (2008).
Zheng, H. et al. p53 and Pten control neural and glioma stem/progenitor cell renewal and differentiation. Nature 455, 1129–1133 (2008).
Marino, S., Vooijs, M., van Der Gulden, H., Jonkers, J. & Berns, A. Induction of medulloblastomas inp53-null mutant mice by somatic inactivation of Rb in the external granular layer cells of the cerebellum. Genes Dev. 14, 994–1004 (2000).
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010).
Momota, H., Shih, A. H., Edgar, M. A. & Holland, E. C. c-Myc and β-catenin cooperate with loss of p53 to generate multiple members of the primitive neuroectodermal tumor family in mice. Oncogene 27, 4392–4401 (2008).
Han, Z.-Y. et al. The occurrence of intracranial rhabdoid tumours in mice depends on temporal control of Smarcb1 inactivation. Nat. Commun. 7, 10421 (2016).
Chen, J., McKay, R. M. & Parada, L. F. Malignant glioma: lessons from genomics, mouse models, and stem cells. Cell 149, 36–47 (2012).
Sturm, D. et al. New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 164, 1060–1072 (2016).
Biegel, J. A. et al. Germ-line and acquired mutations of INI1 in atypical teratoid and rhabdoid tumors. Cancer Res. 59, 74–79 (1999).
Rogers, H. A. et al. WNT/β-catenin pathway activation in Myc immortalised cerebellar progenitor cells inhibits neuronal differentiation and generates tumours resembling medulloblastoma. Br. J. Cancer 107, 1144–1152 (2012).
Hutter, S., Bolin, S., Weishaupt, H. & Swartling, F. J. Modeling and targeting MYC genes in childhood brain tumors. Genes (Basel) 8, 107–119 (2017).
Atkins, M. et al. An ectopic network of transcription factors regulated by Hippo signaling drives growth and invasion of a malignant tumor model. Curr. Biol. 26, 2101–2113 (2016).
Gutmann, D. H., Saporito-Irwin, S., DeClue, J. E., Wienecke, R. & Guha, A. Alterations in the rap1 signaling pathway are common in human gliomas. Oncogene 15, 1611–1616 (1997).
Clark, P. A. et al. Activation of multiple ERBB family receptors mediates glioblastoma cancer stem-like cell resistance to EGFR-targeted inhibition. Neoplasia 14, 420–428 (2012).
Mayer, A., Schneider, F., Vaupel, P., Sommer, C. & Schmidberger, H. Differential expression of HIF-1 in glioblastoma multiforme and anaplastic astrocytoma. Int. J. Oncol. 41, 1260–1270 (2012).
Puliyappadamba, V. T., Hatanpaa, K. J., Chakraborty, S. & Habib, A. A. The role of NF-κB in the pathogenesis of glioma. Mol. Cell. Oncol. 1, e963478 (2014).
Tavares, C. B. et al. Expression of estrogen and progesterone receptors in astrocytomas: a literature review. Clin. (São Paulo) 71, 481–486 (2016).
Ellison, D. et al. Neuropathology. 3rd edn (Mosby: Maryland Heights, MO, 2013).
Rocchi, A. et al. CD99 inhibits neural differentiation of human Ewing sarcoma cells and thereby contributes to oncogenesis. J. Clin. Invest. 120, 668–680 (2010).
Schmitz, M. et al. Identification of SOX2 as a novel glioma-associated antigen and potential target for T cell-based immunotherapy. Br. J. Cancer 96, 1293–1301 (2007).
Seol, H. J. et al. Overexpression of CD99 increases the migration and invasiveness of human malignant glioma cells. Genes Cancer 3, 535–549 (2012).
Iwadate, Y. Epithelial-mesenchymal transition in glioblastoma progression. Oncol. Lett. 11, 1615–1620 (2016).
Binder, D. K. & Berger, M. S. Proteases and the biology of glioma invasion. J. Neurooncol. 56, 149–158 (2002).
Paw, I., Carpenter, R. C., Watabe, K., Debinski, W. & Lo, H.-W. Mechanisms regulating glioma invasion. Cancer Lett. 362, 1–7 (2015).
Trojanowski, J. Q. et al. In vivo and in vitro models of medulloblastomas and other primitive neuroectodermal brain tumors of childhood. Mol. Chem. Neuropathol. 21, 219–239 (1994).
Grotzer, M. A., Neve, A. & Baumgartner, M. Dissecting brain tumor growth and metastasis in vitro and ex vivo. J. Cancer Metastasis Treat. 2, 149–162 (2016).
Muffat, J. et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nat. Med. 22, 1358–1367 (2016).
Cui, X. et al. Hacking macrophage-associated immunosuppression for regulating glioblastoma angiogenesis. Biomaterials 161, 164–178 (2018).
Bian, S. & Knoblich, J.A. Genetic engineering to initiate tumorigenesis in cerebral organoids. Protoc. Exch. https://doi.org/10.1038/protex.2018.071 (2018).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl. Acad. Sci. USA 101, 16–22 (2004).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Bagley, J. A., Reumann, D., Bian, S., Lévi-Strauss, J. & Knoblich, J. A. Fused cerebral organoids model interactions between brain regions. Nat. Methods 14, 743–751 (2017).
Acknowledgements
We are grateful to all members of the Knoblich laboratory for discussions, and H. Gustafson and S. Wolfinger for technical support. We thank O. Wüseke and F. Bonnay for comments on the manuscript. We thank T. Müller, P. Pasierbek, G. Petri, M. Weninger, G. Schmauss, and T. Lendl for FACS and help with imaging. We thank A. Piszczek, T. Engelmaier, J. Klughofer, and M. Zeba for tissue processing and immunohistochemical staining. We thank the Next Generation Sequencing Facility for next-generation sequencing. We thank Boehringer Ingelheim RCV GmbH & Co KG for providing EGFR inhibitors. We thank W. Hu for statistical advice. J.A. Bagley received funding from an EMBO postdoctoral fellowship, and from the European Union’s Horizon 2020 research and innovation program under Marie Skłodowska-Curie grant agreement no. 707109. Work in the Knoblich laboratory is supported by the Austrian Academy of Sciences, the Austrian Science Fund (Z_153_B09), and an advanced grant from the European Research Council (ERC).
Author information
Authors and Affiliations
Contributions
S.B. and J.A.K. conceived the project and experimental design and wrote the manuscript. S.B. performed experiments and analyzed data. M.R., Z.G., and C.K. performed experiments. A.K. contributed the histopathology data. T.B. performed bioinformatics analysis of RNA-seq data. J.A.B. contributed to RNA-seq analysis and quantification of immunostained tissues. J.A.K. directed and supervised the project.
Corresponding author
Ethics declarations
Competing interests
S.B. and J.A.K. have filed a patent application (EP 17190447.7) for use of this method in future disease modeling and preclinical investigation.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The strategy to introduce gene aberrations into neural stem/precursor cells in cerebral organoids.
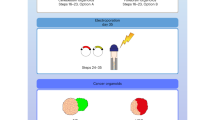
(a) Schematic of the strategy of genome-editing techniques to introduce oncogene amplification and/or tumour suppressor mutation/deletion. Sleeping Beauty transposon system was used to integrate oncogene-expression and GFP-expression elements into genome. CRISPR-Cas9 system was applied to introduce mutation/deletion of tumour suppressors. (b) Quantification of cellular identities of nucleofected cells in EBs one day after nucleofection by immunofluorescence staining on serial cryo-sections. The percentage of marker+ GFP+ cells in total GFP+ cells from different sections were presented. Cell number for different cellular markers analysed in this study was labelled under the graph. This experiment was performed twice independently with same results. (c, d) Immunofluorescence images (c) and quantification (d) of adherent cell culture of dissociated EBs one day after nucleofection stained with different cell markers. The percentage of marker+ GFP+ cells in total GFP+ cells from different EBs were presented. Cell number for different cellular markers analysed in this study was labelled under the graph. This experiment was performed once. All the detailed sample size and mean±SD were provided in Source Data.Scale bar: c, 50 μm.
Supplementary Figure 2 Verification of gene aberrations introduced by genome-editing techniques.
(a) RNA-seq and RT-PCR analysis showed that tumour cells from MYCOE neoCORs exhibit high MYC expression levels. The TPM value from RNA-seq analysis was analyzed using four organoids from three independent cultures (P<0.0001). The TPM values of three organoids from CTRL groups from three independent cultures were presented as control. (b) Three example sequences of CRISPR-Cas9 targeting CDKN2A and CDKN2B locus in tumour cells from GBM-1 neoCORs. RNA-seq and RT-PCR analysis showed that tumour cells from GBM-1 neoCORs exhibit high expression levels of both EGFR and EGFRvIII. The TPM value from RNA-seq analysis was analyzed using four organoids from three independent cultures (p = 0.0068). The TPM values of three organoids from CTRL groups from three independent cultures were presented as control. (c) Three example sequences of CRISPR-Cas9 targeting NF1, PTEN, and p53 locus in tumour cells from GBM-2 neoCORs. (d) Three example sequences of CRISPR-Cas9 targeting CDKN2A and PTEN locus in tumour cells from GBM-3 neoCORs. RNA-seq and RT-PCR analysis showed that tumour cells from GBM-3 neoCORs exhibit high expression level of EGFRvIII, but not EGFR. The TPM value from RNA-seq analysis was analyzed using three organoids from three independent cultures (P = 0.0124). The TPM values of three organoids from CTRL groups from three independent cultures were presented as control. Statistical analysis of quantification was performed using unpaired two-tailed Student’s t-test. Data were presented as mean±s.d. All the detailed sample size, mean±s.d., as well as P value were provided in the Source Data. The percentage of non-homologous end joining (NHEJ) for individual genes by sequencing TA-vector colonies carrying amplified targeting locus were also provided in Source Data. *, P<0.05; **, P<0.01; ***, P<0.001.
Supplementary Figure 3 Venn diagram hypergeometric and KEGG pathway analysis of tumor cells from the cluster 2 and cluster 3 neoCORs.
(a) Venn diagram hypergeometric test showed overlap of differentially expressed genes (DESeq, adjusted absolute log2fc value >0.5 and adjusted P value <0.05) in Cluster 2 (n = 3 organoids from one experiment) or Cluster 3 (n = 7 organoids from one experiment), in each case relative to CTRL organoids (n = 3 organoids from one experiment). P values for overlaps were calculated by hypergeometric test. (b) KEGG pathway enrichment analysis for differentially expressed genes (DESeq, adjusted absolute log2fc value >0.5 and adjusted P value <0.05) in tumour cells from Cluster 2 and Cluster 3 neoCORs.
Supplementary Figure 4 Low-magnification images revealed that 4-month-old neoCORs showed brain-tumor-subtype-specific cellular identities.
(a-f) Immunofluorescence images and quantification of neuronal marker HuC/D (a, magenta), precursor marker SOX2 (b, magenta), cell cycle marker Ki67 (c, magenta), glial marker S100β (d, magenta) and GFAP (e, magenta), as well as CNS-PNET marker CD99 (f, magenta). The staining was performed from six independent experiments with the similar results. Scale bar: a-f, 1000 μm.
Supplementary Figure 5 High-magnification images revealed that 4-month-old neoCORs showed brain-tumor-subtype-specific cellular identities.
Representative immunofluorescence images of four-month-old neoCORs from GBM-2 and GBM-3 groups. Neuronal marker HuC/D (magenta), precursor marker SOX2 (magenta), cell cycle marker Ki67 (magenta), glial marker S100β (magenta) and GFAP (magenta), as well as CNS-PNET marker CD99 (magenta) were presented. The staining was performed from six independent experiments with the similar results. Scale bar: 100 μm.
Supplementary Figure 6 High-magnification images revealed that 1-month-old neoCORs showed brain-tumor-subtype-specific cellular identities.
(a) Immunofluorescence images of control and neoplastic groups one day and one month after nucleofection confirmed the tumour-initiation capability of genetic disruptions. (b) Immunofluorescence images of DAPI (blue) and GFP (green) of control and tumour groups one month after nucleofection. (c-e) Immunofluorescence images and quantification of neuronal marker HuC/D (c, magenta), precursor marker SOX2 (c, blue), cell cycle marker Ki67 (d, magenta), as well as glial marker S100β (e, magenta). The staining was performed from three independent experiments with the similar results. Scale bar: a, upper panel: 200 μm, lower panel: 1000 μm; b, 1000 μm; c-e, 100 μm.
Supplementary Figure 7 Low-magnification images revealed that 1-month-old neoCORs showed brain-tumor-subtype-specific cellular identities.
(a) Immunofluorescence images of control and neoplastic groups one day and one month after nucleofection confirmed the tumour-initiation capability of genetic disruptions. (b) Immunofluorescence images of DAPI (blue) and GFP (green) staining of control and neoplastic groups one month after nucleofection. (c-e) Immunofluorescence images and quantification of neuronal marker HuC/D (c, magenta), precursor marker SOX2 (c, blue), cell cycle marker Ki67 (d, magenta), as well as glial marker S100β (e, magenta). The staining was performed from three independent experiments with the similar results. Scale bar: a, upper panel: 200 μm, lower panel: 1000 μm; b-e, 1000 μm.
Supplementary Figure 8 In vivo expansion of neoCORs after renal subcapsular implantation.
(a) Schematic of renal subcapsular xenograft procedure. (b) NeoCORs from MYCOE group and GBM-1 group were implanted into kidney capsule. Engrafted kidneys were analysed at one week and one and half months after xenograft to evaluate the in vivo expansion of neoCORs from MYCOE and GBM-1 groups. (c) Immunohistochemical staining of neuronal marker MAP2 in the MYCOE implant. Arrowhead shows a neuron. (d-f) H&E staining of implanted MYCOE organoids showing cell sheet (e) and rosette (f) structures. The implantation experiments were performed three times independently with similar results. Scale bar: b, 500 mm; d, 1000 μm; e,f, 50 μm.
Supplementary Figure 9 Drug testing assay showed the drug-screening potential of neoCORs.
(a) Schematic of luciferase assay-based drug testing strategy on neoCORs from GBM-1 group. (b) Quantification of relative luciferase activity revealed that EGFR inhibitors Afatinib (725.57±253.71; P = 0.0076) and Erlotinib (716.10±424.94; P = 0.0074) significantly reduced luciferase activity in GBM-1 (CDKN2A-/CDKN2B-/EGFROE/EGFRvIIIOE) neoCORs (DMSO: n = 9; Canertinib: n = 9; Pelitinib: n = 8; Afatinib: n = 9; Gefitinib: n = 9; Erlotinib: n = 9). Normalized luciferase activity was presented. This experiment was performed once. Statistical analysis of quantifications was performed using one-way ANOVA with Dunnett’s test. Data were presented as mean±SD. All the detailed sample size, mean±SD, as well as p value were provided in Source Data. **, P<0.01.
Supplementary Figure 10 An example of the gating strategy for FACS analysis.
(a) Gating for live cells. (b) Gating to exclude doublets and cell aggregates. (c) Gating for GFP+ cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–7, Supplementary Protocol
Rights and permissions
About this article
Cite this article
Bian, S., Repic, M., Guo, Z. et al. Genetically engineered cerebral organoids model brain tumor formation. Nat Methods 15, 631–639 (2018). https://doi.org/10.1038/s41592-018-0070-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0070-7
This article is cited by
-
Integration of pan-omics technologies and three-dimensional in vitro tumor models: an approach toward drug discovery and precision medicine
Molecular Cancer (2024)
-
Gliomas: a reflection of temporal gliogenic principles
Communications Biology (2024)
-
A multidimensional atlas of human glioblastoma-like organoids reveals highly coordinated molecular networks and effective drugs
npj Precision Oncology (2024)
-
Harnessing 3D in vitro systems to model immune responses to solid tumours: a step towards improving and creating personalized immunotherapies
Nature Reviews Immunology (2024)
-
Insights on Three Dimensional Organoid Studies for Stem Cell Therapy in Regenerative Medicine
Stem Cell Reviews and Reports (2024)