Abstract
The implementation of recommendations for type 2 diabetes (T2D) screening and diagnosis focuses on the measurement of glycated hemoglobin (HbA1c) and fasting glucose. This approach leaves a large number of individuals with isolated impaired glucose tolerance (iIGT), who are only detectable through oral glucose tolerance tests (OGTTs), at risk of diabetes and its severe complications. We applied machine learning to the proteomic profiles of a single fasted sample from 11,546 participants of the Fenland study to test discrimination of iIGT defined using the gold-standard OGTTs. We observed significantly improved discriminative performance by adding only three proteins (RTN4R, CBPM and GHR) to the best clinical model (AUROC = 0.80 (95% confidence interval: 0.79–0.86), P = 0.004), which we validated in an external cohort. Increased plasma levels of these candidate proteins were associated with an increased risk for future T2D in an independent cohort and were also increased in individuals genetically susceptible to impaired glucose homeostasis and T2D. Assessment of a limited number of proteins can identify individuals likely to be missed by current diagnostic strategies and at high risk of T2D and its complications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data access for the Fenland and EPIC studies can be requested by bona fide researchers for specified scientific purposes through a simple application process via the study websites below. Data will either be shared through an institutional data sharing agreement, or arrangements will be made for analyses to be conducted remotely without the necessity for data transfer. Fenland: https://www.mrc-epid.cam.ac.uk/research/studies/fenland/information-for-researchers. EPIC-Norfolk: https://www.mrc-epid.cam.ac.uk/research/studies/epic-norfolk.Source data are provided with this paper.
Code availability
The code used for the machine learning developed framework has been deposited in the following repository: https://github.com/MRC-Epid/iigt_prediction_proteomics.
References
American Diabetes Association 2. Classification and diagnosis of diabetes: standards of medical care in diabetes—2018. Diabetes Care 41, S13–S27 (2018).
International Expert Committee. International Expert Committee report on the role of the A1C assay in the diagnosis of diabetes. Diabetes Care 32, 1327–1334 (2009).
Saeedi, P. et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas, 9th edition. Diabetes Res. Clin. Pract. 157, 107843 (2019).
Meisinger, C. et al. Prevalence of undiagnosed diabetes and impaired glucose regulation in 35–59-year-old individuals in Southern Germany: the KORA F4 Study. Diabet. Med. 27, 360–362 (2010).
Cheng, Y. J. et al. Prevalence of diabetes by race and ethnicity in the United States, 2011–2016. JAMA 322, 2389–2398 (2019).
Richter, B., Hemmingsen, B., Metzendorf, M. I. & Takwoingi, Y. Development of type 2 diabetes mellitus in people with intermediate hyperglycaemia. Cochrane Database Syst. Rev. 10, CD012661 (2018).
Yip, W. C. Y., Sequeira, I. R., Plank, L. D. & Poppit, S. D. Prevalence of pre-diabetes across ethnicities: a review of impaired fasting glucose (IFG) and impaired glucose tolerance (IGT) for classification of dysglycaemia. Nutrients 9, 1273 (2017).
Campbell, M. D. et al. Benefit of lifestyle-based T2DM prevention is influenced by prediabetes phenotype. Nat. Rev. Endocrinol. 16, 395–400 (2020).
Nichols, G. A., Arondekar, B. & Herman, W. H. Complications of dysglycemia and medical costs associated with nondiabetic hyperglycemia. Am. J. Manag Care 14, 791–798 (2008).
Cowie, C. C. et al. Prevalence of diabetes and high risk for diabetes using A1C criteria in the U.S. population in 1988–2006. Diabetes Care 33, 562–568 (2010).
Cederberg, H. et al. Postchallenge glucose, A1C, and fasting glucose as predictors of type 2 diabetes and cardiovascular disease: a 10-year prospective cohort study. Diabetes Care 33, 2077–2083 (2010).
Balkau, B. The DECODE study. Diabetes epidemiology: collaborative analysis of diagnostic criteria in Europe. Diabetes Metab. 26, 282–286 (2000).
Gerstein, H. C. et al. Annual incidence and relative risk of diabetes in people with various categories of dysglycemia: a systematic overview and meta-analysis of prospective studies. Diabetes Res. Clin. Pract. 78, 305–312 (2007).
Chen, Y. et al. Associations of progression to diabetes and regression to normal glucose tolerance with development of cardiovascular and microvascular disease among people with impaired glucose tolerance: a secondary analysis of the 30 year Da Qing Diabetes Prevention Outcome Study. Diabetologia 64, 1279–1287 (2021).
Shaw, J. E., Hodge, A. M., de Courten, M., Chitson, P. & Zimmet, P. Z. Isolated post-challenge hyperglycaemia confirmed as a risk factor for mortality. Diabetologia 42, 1050–1054 (1999).
Silbernagel, G. et al. Isolated post-challenge hyperglycaemia predicts increased cardiovascular mortality. Atherosclerosis 225, 194–199 (2012).
Zhou, W. et al. Longitudinal multi-omics of host–microbe dynamics in prediabetes. Nature 569, 663–671 (2019).
Williams, S. A. et al. Plasma protein patterns as comprehensive indicators of health. Nat. Med. 25, 1851–1857 (2019).
Schussler-Fiorenza Rose, S. M. et al. A longitudinal big data approach for precision health. Nat. Med. 25, 792–804 (2019).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Lindsay, T. et al. Descriptive epidemiology of physical activity energy expenditure in UK adults (The Fenland study). Int. J. Behav. Nutr. Phys. Act. 16, 126 (2019).
Rahman, M., Simmons, R. K., Harding, A. H., Wareham, N. J. & Griffin, S. J. A simple risk score identifies individuals at high risk of developing type 2 diabetes: a prospective cohort study. Fam. Pract. 25, 191–196 (2008).
Deora, A. B., Kreitzer, G., Jacovina, A. T. & Hajjar, K. A. An annexin 2 phosphorylation switch mediates p11-dependent translocation of annexin 2 to the cell surface. J. Biol. Chem. 279, 43411–43418 (2004).
Guevara-Aguirre, J. et al. Growth hormone receptor deficiency is associated with a major reduction in pro-aging signaling, cancer, and diabetes in humans. Sci. Transl. Med. 3, 70ra13 (2011).
Tiaden, A. N. et al. Novel function of serine protease HTRA1 in inhibiting adipogenic differentiation of human mesenchymal stem cells via MAP kinase-mediated MMP upregulation. Stem Cells 34, 1601–1614 (2016).
Haddad, Y. & Couture, R. Kininase 1 as a preclinical therapeutic target for kinin B1 receptor in insulin resistance. Front. Pharmacol. 8, 509 (2017).
Klement, J. et al. Oxytocin improves beta-cell responsivity and glucose tolerance in healthy men. Diabetes 66, 264–271 (2017).
Zhong, C. et al. Cbln1 and Cbln4 are structurally similar but differ in GluD2 binding interactions. Cell Rep. 20, 2328–2340 (2017).
Weingarten, M. F. J. et al. Circulating oxytocin is genetically determined and associated with obesity and impaired glucose tolerance. J. Clin. Endocrinol. Metab. 104, 5621–5632 (2019).
Wu, T. et al. CILP-2 is a novel secreted protein and associated with insulin resistance. J. Mol. Cell Biol. 11, 1083–1094 (2019).
Slieker, R.C., et al. Novel biomarkers for glycaemic deterioration in type 2 diabetes: an IMI RHAPSODY study. Preprint at medRxiv https://doi.org/10.1101/2021.04.22.21255625 (2021).
Shen, Z., Gantcheva, S., Mansson, B., Heinegard, D. & Sommarin, Y. Chondroadherin expression changes in skeletal development. Biochem. J. 330, 549–557 (1998).
Hessle, L. et al. The skeletal phenotype of chondroadherin deficient mice. PLoS ONE 8, e63080 (2014).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Scott, R. A. et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat. Genet. 44, 991–1005 (2012).
Lotta, L. A. et al. Association of genetic variants related to gluteofemoral vs abdominal fat distribution with type 2 diabetes, coronary disease, and cardiovascular risk factors. JAMA 320, 2553–2563 (2018).
Day, N. et al. EPIC-Norfolk: study design and characteristics of the cohort. European Prospective Investigation of Cancer. Br. J. Cancer 80, 95–103 (1999).
Pietzner, M. et al. Plasma metabolites to profile pathways in noncommunicable disease multimorbidity. Nat. Med. 27, 471–479 (2021).
Marmot, M. & Brunner, E. Cohort profile: the Whitehall II study. Int. J. Epidemiol. 34, 251–256 (2005).
Zhong, W. et al. Next generation plasma proteome profiling to monitor health and disease. Nat. Commun. 12, 2493 (2021).
Gong, Q. et al. Morbidity and mortality after lifestyle intervention for people with impaired glucose tolerance: 30-year results of the Da Qing Diabetes Prevention Outcome Study. Lancet Diabetes Endocrinol. 7, 452–461 (2019).
Barron, E., Clark, R., Hewings, R., Smith, J. & Valabhji, J. Progress of the Healthier You: NHS Diabetes Prevention Programme: referrals, uptake and participant characteristics. Diabet. Med. 35, 513–518 (2018).
Gong, Q. et al. Efficacy of lifestyle intervention in adults with impaired glucose tolerance with and without impaired fasting plasma glucose: a post hoc analysis of Da Qing Diabetes Prevention Outcome Study. Diabetes Obes. Metab. 23, 2385–2394 (2021).
Knowler, W. C. et al. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N. Engl. J. Med. 346, 393–403 (2002).
Bergman, M. et al. Lessons learned from the 1-hour post-load glucose level during OGTT: current screening recommendations for dysglycaemia should be revised. Diabetes Metab. Res. Rev. 34, e2992 (2018).
Pham, C. T. Neutrophil serine proteases: specific regulators of inflammation. Nat. Rev. Immunol. 6, 541–550 (2006).
Wiedow, O. & Meyer-Hoffert, U. Neutrophil serine proteases: potential key regulators of cell signalling during inflammation. J. Intern. Med. 257, 319–328 (2005).
Donath, M. Y. & Shoelson, S. E. Type 2 diabetes as an inflammatory disease. Nat. Rev. Immunol. 11, 98–107 (2011).
de Vries, M. A. et al. Glucose-dependent leukocyte activation in patients with type 2 diabetes mellitus, familial combined hyperlipidemia and healthy controls. Metabolism 64, 213–217 (2015).
Pietzner, M. et al. Synergistic insights into human health from aptamer- and antibody-based proteomic profiling. Nat. Commun. 12, 6822 (2021).
Lee, C. M. Y. et al. Comparing different definitions of prediabetes with subsequent risk of diabetes: an individual participant data meta-analysis involving 76 513 individuals and 8208 cases of incident diabetes. BMJ Open Diabetes Res. Care 7, e000794 (2019).
Inker, L. A. et al. Estimating glomerular filtration rate from serum creatinine and cystatin C. N. Engl. J. Med. 367, 20–29 (2012).
Mehta, S. R., Thomas, E. L., Bell, J. D., Johnston, D. G. & Taylor-Robinson, S. D. Non-invasive means of measuring hepatic fat content. World J. Gastroenterol. 14, 3476–3483 (2008).
Pietzner, M. et al. Mapping the proteo-genomic convergence of human diseases. Science 374, eabj1541 (2021).
Rohloff, J. C. et al. Nucleic acid ligands with protein-like side chains: modified aptamers and their use as diagnostic and therapeutic agents. Mol. Ther. Nucleic Acids 3, e201 (2014).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Huang, J. et al. Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat. Commun. 6, 8111 (2015).
Lunardon, N., Menardi, G. & Torelli, N. ROSE: a package for binary imbalanced learning. R. J. 6, 79–89 (2014).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Kundu, S., Aulchenko, Y. S., van Duijn, C. M. & Janssens, A. C. PredictABEL: an R package for the assessment of risk prediction models. Eur. J. Epidemiol. 26, 261–264 (2011).
Harrell, F. E. Jr. rms: regression modeling strategies. R. package version 5 (2017).
Bonate, P. L. & Howard, D. R. (eds) Pharmacokinetics in Drug Development: Clinical Study Design and Analysis (AAPS Press, 2004).
Hoffman, G. E. & Schadt, E. E. variancePartition: interpreting drivers of variation in complex gene expression studies. BMC Bioinformatics 17, 483 (2016).
InterAct Cosortium. Design and cohort description of the InterAct Project: an examination of the interaction of genetic and lifestyle factors on the incidence of type 2 diabetes in the EPIC Study. Diabetologia 54, 2272–2282 (2011).
Acknowledgements
The Fenland study (10.22025/2017.10.101.00001) is funded by the Medical Research Council (MC_UU_12015/1). We are grateful to all the volunteers and to the general practitioners and practice staff for assistance with recruitment. We thank the Fenland study investigators, Fenland study coordination team and epidemiology field, data and laboratory teams. We further acknowledge support for genomics from the Medical Research Council (MC_PC_13046). Proteomic measurements were supported and governed by a collaboration agreement between the University of Cambridge and SomaLogic. We thank I. von Carlowitz and K. Soucie for their contributions to the fasting proteome analysis.The EPIC-Norfolk study (10.22025/2019.10.105.00004) has received funding from the Medical Research Council (MR/N003284/1 MC-UU_12015/1 and MC_UU_00006/1) and Cancer Research UK (C864/A14136). We are grateful to all the participants who have been part of the project and to the many members of the study teams at the University of Cambridge who have enabled this research. We thank all participants in the Whitehall II Study, Whitehall II researchers and support staff who make the study possible. The UK Medical Research Council (MR/K013351/1; G0902037), British Heart Foundation (RG/13/2/30098) and the US National Institutes of Health (R01HL36310, R01AG013196) have supported collection of data in the Whitehall II Study. J.C.Z.S. is supported by a 4-year Wellcome Trust PhD Studentship and the Cambridge Trust, and C.L., E.W. and N.J.W. are funded by the Medical Research Council (MC_UU_12015/1). N.J.W. is an NIHR Senior Investigator. The WHII study and M.K. are supported by grants from the Wellcome Trust (221854/Z/20/Z), UK Medical Research Council (R024227) and NIA, NIH (R01AG056477). J.V.L. was supported by the Academy of Finland (311492 and 339568) and Helsinki Institute of Life Science (H970) grants paid to employer and by the Päivikki and Sakari Sohlberg foundation. The funders had no role in the study design, data collection and analysis and the decision to publish or in the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.C.Z.S., M.P., N.J.W. and C.L. designed the analysis and drafted the manuscript. J.C.Z.S. analyzed the data. J.V.L. performed the replication analyses in the WHII study. M.S. and M.W. performed the analysis for assessing the effect of fasting status on protein levels. N.J.W. is the principal investigator of the Fenland study cohort, and M.K. is the principal investigator of the WHII study. All authors contributed to the interpretation of the results and critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.S., M.W., D.D., R.O. and S.A.W. are employees of SomaLogic. E.W. and E.O. are now employees at AstraZeneca. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Peter Rossing, Jesse Meyer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Jennifer Sargent, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Feature selection ranking of proteins for postprandial glycaemia prediction.
Protein ranking based on the number of times selected over bootstrap resampling during feature selection for impaired glucose tolerance (IGT) (a) and isolated impaired glucose tolerance (iIGT) (b). Dashed lines represent thresholds for proteins selected in more than 80%, 90% or 95% of bootstrap samples to be taken forward to parameter optimization step.
Extended Data Fig. 2 Performance of LASSO trained protein-only models for IGT (a) and iIGT (b) discrimination in the internal validation test set.
a, Impaired glucose tolerance (IGT) discrimination was evaluated in the independent internal validation test set (N = 2881, 192 IGT individuals) for models based on proteins selected in more than 80% (65 proteins), 90% (18 proteins) or 95% (8 proteins) and kept after model optimization step or based on all proteins (4979 proteins). b, isolated impaired glucose tolerance (iIGT) discrimination was evaluated in the independent internal validation test set (N = 2819, 135 iIGT individuals) for models based on proteins selected in more than 80% (73 proteins), 90% (17 proteins) or 95% (3 proteins) and kept after model optimization step or based on all proteins (4979 proteins).
Extended Data Fig. 3
Performance of the T2D genetic risk score (T2D-GRS) for IGT (a) and iIGT (b) discrimination in the internal validation test set.
Extended Data Fig. 4 Validation of the clinical and clinical + protein models for IGT (a) and iIGT (b) in the independent WHII study.
The clinical + protein model significantly outperformed the clinical model (p-valueIGT = 5.26 × 10−5; p-valueiIGT = 1.5 × 10−17). The improvement was of similar magnitude than that observed in the Fenland study, although with overall lower AUROCs (clinical models: AUROCIGT = 0.66 (0.64–0.69), and AUROCiIGT = 0.60 (0.57–0.62); clinical + protein models: AUROCIGT = 0.70 (0.68–0.72) and AUROCiIGT = 0.69 (0.67–0.71)). Significant differences between the AUROCs were asses by the Delong method. This might be best explained by differences in the characteristics of the study population, the design and the lack of HbA1c to define iIGT (see Methods).
Extended Data Fig. 5 Performance of LASSO trained models for isolated impaired glucose tolerance discrimination in the internal validation test set having excluded the top 3 selected proteins.
Isolated impaired glucose tolerance (iIGT) discrimination performance in the independent internal validation test set (N = 2795, 111 iIGT individuals) for the standard clinical model, a 68-protein model (selected in >80% of bootstrap samples and kept during optimization), and a clinical + 7 protein model (selected in >95% of bootstrap samples).
Extended Data Fig. 6 Internal validation of proposed 3-stage screening strategy in the test set only.
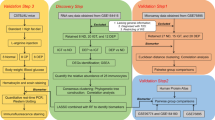
In the first stage, individuals in the Fenland test set were divided into low and high risk according to the Cambridge T2D risk score. The high risk group would undergo a second stage involving measurement of HbA1c and of the 3 iIGT proteins. Individuals with HbA1c levels within the T2D or prediabetic range would be referred for intervention and lifestyle modifications. Individuals with HbA1c below the prediabetic range, would further stratified using the final clinical + 3 iIGT protein model to identify a high risk group, which on a third stage would be taken forward for OGTT testing to identify iIGT cases that would have been otherwise by current screening guidelines. The NNS in the strata of individuals at high predicted risk based on the patient-derived information model, but HbA1c levels below cut-offs for prediabetes (N = 1043) was 14, while by additionally applying the clinical + 3-protein iIGT model the NNS was of only 5 (N = 88 at high-risk). Figure was designed with biorender.com.
Extended Data Fig. 7 Comparison of protein ranking during feature selection over bootstrap resampling for isolated impaired glucose tolerance (iIGT) and impaired glucose tolerance (IGT).
Comparison is shown for proteins that were selected in more 80% of bootstrap samples (shown by the red line) for either IGT (N = 2881, 192 IGT individuals) or iIGT (N = 2795, 111 iIGT individuals).
Extended Data Fig. 8 Percentage of variance explained in impaired glucose tolerance and isolated impaired glucose tolerance top discriminatory protein levels by clinical, biochemical, anthropometric and lifestyle risk factors.
Linear mixed models were fitted for each of the 24 clinical, biochemical, anthropometric, genetic and lifestyle risk factor variables adjusting by age and sex to estimate the percentage of explained variance in plasma abundances of discriminatory proteins as well as for the principal component of the 65-IGT and 68-iIGT protein signatures. Cis and trans scores with missing values represent proteins for which no protein quantitative trait loci could be identified.
Extended Data Fig. 9 Association of iIGT protein scores using Olink explore proteomics measures with incident cardiometabolic diseases.
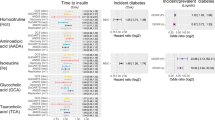
Association of iIGT prediction scores (left panel; red: Cambridge T2D risk score, orange: Cambridge T2D risk score variable + fasting glucose + 3 protein iIGT prediction model, darkblue: 3-protein iIGT prediction model) and individual top iIGT proteins (right panel) with 7 cardiometabolic disease outcomes in a sub-cohort the EPIC-Norfolk study (N = 602 individuals). 95% confidence intervals of hazard ratios (HR) are shown.
Supplementary information
Supplementary Tables
Supplementary Tables 1–19.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Carrasco-Zanini, J., Pietzner, M., Lindbohm, J.V. et al. Proteomic signatures for identification of impaired glucose tolerance. Nat Med 28, 2293–2300 (2022). https://doi.org/10.1038/s41591-022-02055-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-02055-z
This article is cited by
-
Inter-organ crosstalk during development and progression of type 2 diabetes mellitus
Nature Reviews Endocrinology (2024)
-
Lessons and Applications of Omics Research in Diabetes Epidemiology
Current Diabetes Reports (2024)
-
Harnessing the power of proteomics in precision diabetes medicine
Diabetologia (2024)
-
Multi-omic prediction of incident type 2 diabetes
Diabetologia (2024)
-
Genetic determinants of micronucleus formation in vivo
Nature (2024)