Abstract
Early recognition and treatment of sepsis are linked to improved patient outcomes. Machine learning-based early warning systems may reduce the time to recognition, but few systems have undergone clinical evaluation. In this prospective, multi-site cohort study, we examined the association between patient outcomes and provider interaction with a deployed sepsis alert system called the Targeted Real-time Early Warning System (TREWS). During the study, 590,736 patients were monitored by TREWS across five hospitals. We focused our analysis on 6,877 patients with sepsis who were identified by the alert before initiation of antibiotic therapy. Adjusting for patient presentation and severity, patients in this group whose alert was confirmed by a provider within 3 h of the alert had a reduced in-hospital mortality rate (3.3%, confidence interval (CI) 1.7, 5.1%, adjusted absolute reduction, and 18.7%, CI 9.4, 27.0%, adjusted relative reduction), organ failure and length of stay compared with patients whose alert was not confirmed by a provider within 3 h. Improvements in mortality rate (4.5%, CI 0.8, 8.3%, adjusted absolute reduction) and organ failure were larger among those patients who were additionally flagged as high risk. Our findings indicate that early warning systems have the potential to identify sepsis patients early and improve patient outcomes and that sepsis patients who would benefit the most from early treatment can be identified and prioritized at the time of the alert
Similar content being viewed by others
Main
Sepsis is a leading cause of in-hospital death in the United States, with a recent study finding sepsis as the immediate cause of nearly 35% of in-hospital deaths1. Effective intervention has been elusive; the current state has been referred to as a ‘treatment graveyard’2 because few effective sepsis treatments have been developed and there is persistent debate about best treatment practices2,3. There is agreement that early recognition and treatment are critical to successful outcomes4. Early administration of broad-spectrum intravenous antibiotics, in particular, is associated with decreased mortality and morbidity5,6,7,8,9,10. However, heterogeneity in the presentation of sepsis often makes early recognition challenging, and many patients receive delayed care1. This has spurred interest in the development of automated sepsis early warning systems to help clinicians recognize sepsis as early as possible.
Several retrospective studies have demonstrated that machine learning (ML)-based models can detect sepsis early11,12,13,14,15. However, few studies have reported on clinical implementations of these models11,16,17. Although the existing implementation studies of both general18 and sepsis-specific alert systems11,16,17,19,20 demonstrate the feasibility of deployment, several of these studies have relied on dedicated staff to respond to alerts and the reported clinical value has been mixed, with clinical suspicion of sepsis often noted before alerts19,20. Additional studies are needed to understand the impact of sepsis-specific early warning systems on patient outcomes and to demonstrate the potential of decentralized bedside alert systems.
After 3 years of development, the TREWS ML-based early warning system was deployed21, starting in 2018 as part of an electronic health record (EHR)-based sepsis alert system in two academic and three community hospitals in the Maryland and DC areas. Details of the model, deployment and workflow are described in a companion paper22. Of note, when a TREWS alert occurs, providers may open a dedicated TREWS page in the electronic medical record system (in this case, Epic) and, from that page, may enter an evaluation of the patient as having sepsis or not. In a companion paper, we analyzed the predictive performance of TREWS, the adoption of TREWS by providers and the association between interaction with TREWS and the time between the alert and antibiotic ordering22. Notably, we found that sepsis patients who had their alert evaluated and confirmed within 3 h had a 1.85-h lower median time from alert to first antibiotic order22.
In the present study, we analyzed the association between provider interaction with TREWS and patient outcomes for a target population of sepsis patients who had an alert before they received any antibiotics. Using EHR data collected from the five deployment sites, we conducted a prospective, multicenter, two-arm cohort study to evaluate the association between timely provider evaluation and confirmation of the TREWS alert and mortality for a target population of sepsis patients who had an alert before receiving any antibiotics. Additional outcomes of interest are progression in a patient’s total sequential organ failure assessment (SOFA) score, following the alert, and length of stay. In a secondary analysis, we further examined this association across patients identified as having increased risk of death in the absence of timely antibiotics, referred to henceforth as the high-risk cohort. This high-risk cohort was defined based on a secondary risk score that uses measurements available at or near the time of alert.
Results
Study population
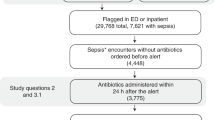
During the study period, 590,736 patient encounters involving patients over the age of 18 years who presented to an emergency department (ED) or were admitted to an inpatient unit at one of the five deployment hospitals were monitored by TREWS. Of these encounters, 42,089 (7.1%) triggered an alert and 13,680 (2.3%) met the retrospective criteria for sepsis23. Among encounters that triggered an alert, 24,799 (59%) had their first alert before admission to an inpatient unit, whereas 17,290 (41%) had their first alert after inpatient admission. A total of 6,877 patient encounters met the inclusion criteria for our target population: triggered an alert at most 1 h before ED triage or inpatient admission, met the retrospective definition for sepsis, received their first antibiotic after the alert and within 24 h of the alert and were not admitted directly to an intensive care unit (ICU). Of these, 2,366 were included in the high-risk cohort. In addition, 2,458 encounters involved patients who had sepsis but did not trigger an alert, and 3,006 encounters involved sepsis patients who received antibiotics before triggering an alert. These two off-target cohorts had lower general severity at admission than the primary analysis cohort (for example, the average Acute Physiology and Chronic Health Evaluation (APACHE II) and SOFA scores were lower in these groups), as well as having a lower in-hospital mortality rate. Figure 1 shows a waterfall diagram for the primary analysis cohort, and Table 1 shows summary statistics for all patients, sepsis patients with no alert, sepsis patients who received antibiotics before triggering an alert and our primary analysis cohort. A breakdown of the primary analysis cohort by hospital can be found in Extended Data Table 1.
Waterfall diagram describing the primary analysis and high-risk cohorts. A total of 590,736 unique adult patient encounters occurred during the study period. Of these, 6,877 patient encounters were included in our primary analysis, and 2,365 patient encounters were included in our analysis of high-risk patients.
Association between response to TREWS and patient outcomes
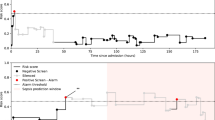
Table 1 shows select summary statistics for the treatment and comparison groups, and Extended Data Table 2 shows a comparison of all variables. Among included patients, 4,220 (61%) had TREWS used as intended with their alert evaluated and confirmed within 3 h of the alert (study arm) and 2,657 (39%) did not (comparison arm). The full distribution of time from alert occurrence to alert confirmation is shown in Fig. 2. Among other differences, patients in the present study arm were older (67 versus 65 years, P < 0.001), had lower SOFA scores at the time of the alert (4.0 versus 4.3, P < 0.001) and were more likely to have a lactate >2 mmol l−1 (75% versus 61%, P < 0.001).
Each bar has a width of 1 h, and the bar height represents the number of subjects who had their alert confirmed in that 1-h time bin. The dashed line is placed at 3 h, such that bars to the left and right of the line correspond to subjects in the study and comparison arms, respectively. Note that 1,840 (27%) patients in the target population either did not have an evaluation entered or had their alert dismissed by a provider and thus do not appear in this plot.
Table 2 shows the unadjusted outcomes and adjusted comparisons of outcomes between study and comparison arms. Patients in the treatment group had lower unadjusted mortality rates (14.6% versus 19.2%, P < 0.001), improved SOFA score progression (−0.8 versus −0.4, P < 0.001) and lower median length of stay among survivors (6.6 d versus 8.1 d, P < 0.001). After adjustment for patient demographics, medical history, laboratory measurements, vital signs, comorbidities and admitting hospital, timely alert confirmation by the provider was associated with lower mortality (adjusted risk difference (ARD) −3.34%, CI −5.10, −1.67%, and adjusted relative reduction (ARR) −18.18%, CI −26.31, −9.65%; P < 0.001), improved SOFA progression (ARD −0.26; CI −0.42, −0.11; P = 0.001) and lower median length of stay among survivors (ARD −11.58 h, CI −18.13, −5.03 h; P = 0.001). In sensitivity analyses, none of the following changed the direction or significance of these associations: changing the estimation method; changing the data source for comorbidities, including only the first encounter for each patient and only encounters before the COVID-19 pandemic (that is, before 31 March 2020); or restricting the list of antibiotics used (Extended Data Table 3).
Sepsis is highly heterogeneous24, and there is evidence that the importance of early antibiotic therapy is similarly heterogeneous7,25. Considering this, we sought to estimate the associations between provider interaction with TREWS and patient outcomes among those patients who were predicted, based on baseline variables, to be most sensitive to antibiotic timing. We predicted sensitivity to antibiotic timing as the increase in predicted mortality risk when using a model trained on patients who received timely antibiotics (within 3 h of their alert) versus a model trained on those who did not (see Methods for details). This approach was based in part on existing methods for predicting which patients are likely to respond most to a particular treatment26. We developed the two mortality risk models using retrospective, pre-deployment data and applied them prospectively to our study population to identify a high-risk cohort of patients. Among the 2,365 patients who fell into this high-risk cohort, alert evaluation and confirmation within 3 h was associated with a greater decrease in mortality (ARD −4.50%, CI −8.31, −0.78% and ARR −13.19%, CI −22.81, −2.45%; P = 0.012) and SOFA progression (ARD −0.38, CI −0.71, −0.05; P = 0.025). The association with length of stay was slightly greater in the high-risk cohort than in the full sample, although it was no longer statistically significant. Full results on the high-risk stratum are presented in Table 2.
Antibiotic timing relative to TREWS and patient outcomes
Previous work has established an association between antibiotic timing relative to ED admission and patient mortality5,6,7,8,25. In the present study, we instead measure the association between antibiotic timing relative to a TREWS alert and patient outcomes. Adjusting for patient demographics, medical history, labs, vital signs, comorbidities and admitting hospital, patients in the target population who received antibiotics within 3 h of their alert had a lower mortality rate (ARD −3.54%, CI −5.52, −1.66%, and ARR −18.69%, CI −27.03, −9.36%; P < 0.001), SOFA score progression (ARD −0.26, CI −0.42, −0.11; P = 0.001) and median length of stay (ARD −11.58 h, CI −18.13, −5.03 h; P = 0.001) compared with those who received antibiotics >3 h after their alert. Associations in the high-risk cohort are presented in Extended Data Table 4. In addition, for the purposes of comparison with previous retrospective studies, we estimated the adjusted association between in-hospital mortality and each hour of delay from the time of the alert to first antibiotics for patients receiving antibiotics within 6 and 12 h of their alert. The 6- and 12-h windows were chosen for comparison to previous literature. The adjusted odds ratios for in-hospital mortality per hour delay in antibiotics were 1.08 (CI 1.02, 1.15) and 1.05 (CI 1.02, 1.09) among patients who received antibiotics within 6 and 12 h, respectively. In the high-risk cohort, the adjusted odds ratios for in-hospital mortality per hour delay in antibiotics were 1.07 (CI 0.99, 1.17) and 1.08 (CI 1.03, 1.14) among patients who received antibiotics within 6 and 12 h, respectively.
Discussion
In the present study, we found that, after adjusting for a range of patient variables, timely provider evaluation and confirmation of the TREWS alert were associated with reduced mortality among target sepsis patients, as well as improved SOFA score progression and reduced median length of stay among survivors. Absolute associations were of greater magnitude among patients prospectively identified as being at high risk for death without timely treatment. In addition, we found that delay in antibiotics relative to the time of the alert was associated with increased patient mortality in our target population. These results complement our companion paper which found that TREWS had high adoption (89% of alerts received an evaluation) and timely provider confirmation of the TREWS alert was associated with a 1.85-h reduction in median time from alert to first antibiotic order22.
Although studies of deployed sepsis early warning systems have been uncommon, a few studies have found positive benefits when using such systems16,27,28,29. Three independent pre-/post-deployment studies found that sepsis-related mortality dropped after system deployment27,28,29. One major limitation of pre-/post-studies of sepsis-related mortality is the potential for surveillance bias, which occurs when more patients are coded as septic in the post-period due to the alerting tool or concurrent changes in coding practices. Without appropriate adjustments, clinical outcomes appear to have improved because the group of patients coded as septic are less sick than before the alert was implemented. Our study avoided this issue by including only post-deployment data, making use instead of natural variability in provider practice to create a convenience control group30,31. The reasons behind this variability in provider practice are explored more deeply in our companion paper22.
One randomized trial by Shimabukuro et al. found that use of a sepsis alert system led to reductions in all-cause mortality and length of stay16. Although limited in size (42 septic patients and 142 total patients) and restricted to ICU patients, Shimabukuro et al. is, to our knowledge, the only randomized trial of a prospective sepsis alerting system. In contrast, our study does not make use of randomization. However, our findings are consistent with those of Shimabukuro et al. and complement their findings by including a larger sample of patients from diverse units across multiple sites.
Studies of two other systems did not demonstrate impact on patient outcomes11,19,20. One pre-/post-deployment study found that deployment led to modest but statistically significant increases in lactate ordering and intravenous fluid administration but did not impact antibiotic ordering or patient outcomes11. In follow-up interviews, providers reported that the alerts were unlikely to affect their perception of a patient20. A similar pre-/post-deployment study found that deployment of an early warning system did not lead to changes in clinical practice or patient outcomes19. A notable difference between these systems and TREWS is the accuracy of the alerts. For example, the latter study revealed that only 29% of sepsis cases were flagged by the alerts and all alerts on sepsis patients occurred after the administration of antibiotics, at which point the alerts had limited clinical utility19. A recent evaluation of the Epic Sepsis Model, one of the most widely deployed sepsis early warning systems, found that only 12% of alerts occurred on sepsis patients and only 33% of sepsis cases were flagged by the system32. In contrast to these three studies, our companion paper found that 82% of sepsis cases were flagged by the TREWS alerts and 38% of alerts with an evaluation were confirmed by a provider22.
Beyond sepsis, a recent large-scale study of the Advanced Alert Monitor—an alert system for general deterioration—found that the alert system improved patient outcomes18. Although an important demonstration of the potential of such systems, this study relied on dedicated staff to respond to each alert. In contrast, the system studied in the present study is a decentralized bedside system that does not trigger pop-ups or phone calls that interrupt the clinical workflow. There were no dedicated staff for reviewing and responding to alerts and providers chose if, when and how to respond to alerts. Having a decentralized bedside system may allow TREWS to better scale to hospital systems that lack the resources to allocate staff to review all alerts. On the other hand, systems without dedicated staff require buy-in and adoption from providers to be successful. In our companion paper, we used EHR data to examine various patient, provider and environmental factors associated with the adoption of TREWS22. In a further separate study, we analyzed qualitative impressions of TREWS and factors influencing its integration into workflow through semi-structured interviews with providers using the system33.
Much of the work on early detection and treatment for sepsis has been motivated by a collection of retrospective studies showing that delays in antibiotics are associated with worse patient outcomes5,6,7,8,25. In the absence of a viable alternative, these studies generally measured the time to antibiotics using ED presentation as time zero. Although informative, these results do not suggest a way to recognize sepsis earlier and, as a result, much of the subsequent work has focused on reducing the time from recognition to treatment, for example, the extensive work on sepsis treatment bundle guidelines9,10. A recent position paper from the Infectious Disease Society of America highlights the need for such a time zero, noting that existing proposals for time zero are overly complex and subjective24. In the current study, the per-hour associations we found between time-to-first antibiotics and mortality closely matched two of the largest such retrospective studies7,25. Among sepsis patients who received antibiotics within 6 h and 12 h, these studies found an adjusted per-hour odds ratios for mortality of 1.09 (CI 1.05, 1.13) and 1.04 (CI 1.03, 1.06), respectively (we found odds ratios of 1.08 (CI 1.02, 1.15) and 1.05 (CI 1.02, 1.09)). The implication of this result is that the potential benefits of early recognition implied by these retrospective studies could be realized using a high-precision bedside early warning system such as TREWS. Furthermore, such high-precision alerts may be used as a more objective time zero when measuring compliance with sepsis treatment guidelines.
Previous studies have reported different magnitudes of mortality reduction associated with early antibiotic administration, with the strongest benefits observed in patients with septic shock7,8. However, thus far, these retrospective studies have not outlined methods to prospectively identify patients at highest risk for developing septic shock. One important finding of the present study is that patients who benefit more from timely treatment, namely, the high-risk cohort, can be identified near the time of the alert, which may be used to prioritize alerts. Adding an indicator of alert priority, alerting a rapid response team to high-risk cases or limiting alerts to such cases could reduce alert burden and help providers allocate time and resources to the patients most likely to benefit from timely intervention.
Our study has several limitations. First, the present study reflects one set of prespecified alert settings; the behavior of the alert, and thus associations between alert use and clinical outcomes, may vary under different alert settings. For example, in this deployment, the timing of the alerts was optimized to issue alerts only when patients had notable findings such as the presence of deterioration in key markers. In future deployments, this restriction may be relaxed to provide earlier warning and more lead time.
Second, although we adjusted for a substantial list of potential confounding variables, we did not conduct a randomized trial, and therefore, we observed that associations may be subject to residual confounding; that is, it is possible that unobserved variables remained not included in our analysis, which cause both the provider response to TREWS and patient outcomes. Aspects of patient presentation that are not captured through clinical data in the EHR or billing codes are not represented in our patient variables. A recent paper raised the concern that observational studies on sepsis bundles do not account sufficiently for providers appropriately delaying care on medically complex patients24. We have attempted to address this concern by adjusting for the coding of comorbidities that may mimic sepsis, but it is possible that suspected comorbidities that are not documented as codes were not included. In addition, besides the hospital at which the encounter occurred, we have not adjusted for provider-specific variables. Thus, if alert confirmation was associated with patient care, for reasons that are not explained by patient variables, this may lead to additional confounding. Unfortunately, large-scale randomized trials of early warning systems are operationally difficult to carry out, requiring tight control over clinical response if randomizing at the patient level or deployments spanning many sites if randomizing at the unit or hospital level. The most common alternative, pre-/post-deployment studies, suffer from their own set of potential biases such as surveillance bias and changes in treatment standards over time. Although we used a prospective cohort design in the present study, it is likely that all three types of studies—randomized, pre/post and cohort—will be needed to improve our understanding of these systems.
Third, there is continued debate about how to reliably identify sepsis retrospectively. We identified sepsis cases using an EHR-based sepsis phenotype that accounts for comorbidities that mimic sepsis and has shown increased sensitivity and precision compared with the alternatives23,34, but we cannot exclude the possibility that some identified patients had noninfectious syndromes with symptoms that mimic the presentation of sepsis. Fourth, we did not have data on whether the antibiotics given to a patient were appropriate for their infection. Assessing the appropriateness of antimicrobial therapy, potentially via positive blood culture results when available, may give more accurate assessments of when effective treatment began and sepsis-related outcomes5,35. Fifth, our study was limited to a single hospital system and geographical region. Although studies including more diverse populations from other geographical regions are needed, the concordance of our results with several diverse studies on antibiotics timing suggests that our results may be applicable to a broader target population. Finally, the present study considers the associations between alert use and patient outcomes on a target population of sepsis patients who received an alert but had not received antibiotics at the time of the alert. It is also important to consider the impact of TREWS on other populations. For example, it is important to ensure that alerts on nonsepsis patients do not lead to over-prescription of antibiotics and that resources are not being drawn away from sepsis patients who do not receive an alert. These populations will be the subject of future studies.
Although large-scale randomized trials are needed, our findings indicate the potential for high-precision alert systems to identify sepsis patients early and improve patient outcomes. Furthermore, our findings among high-risk patients indicate that future alert systems may reduce overall alert burden by prioritizing alerts on certain patients. Work is under way to further improve on TREWS and to generalize TREWS to other acute conditions. Furthermore, work is needed to understand how the use and benefits of alert systems evolve over time, best practices for presenting information from ML-based alerts in critical care settings, and the tradeoffs between workflows with and without dedicated staff in terms of provider burden, benefits to patients and scalability.
Methods
The present study was approved by the Johns Hopkins University internal review board (IRB no. 00252594) and a waiver of consent was obtained.
TREWS
TREWS is an ML-based early warning system that continuously monitors patients for risk of sepsis using routinely collected EHR data. Its underlying model uses vital signs, laboratory data, clinician notes, medication history (excluding antibiotic orders), procedure history and clinical history to generate a sepsis risk score in real time as new information becomes available in the EHR. The score is based on an approach that allows the model36 to account for different sepsis presentations and to incorporate patient context such as confounding comorbidities, procedures and level of care. For a full description of the TREWS model and a retrospective performance evaluation, as well as the interface, deployment and integration into the clinical workflow, see our companion paper22. Importantly, when an alert occurs, providers can enter a dedicated TREWS page in the EHR that displays various pieces of information about the alert and the patient and allows the provider to enter a patient evaluation. An evaluation consists of entering a suspected source of infection (confirmation) or clicking a button to indicate that no new or worsening infection is present and then reviewing the list of organ dysfunction indicators and removing any that the provider believes should not be attributed to sepsis (dismissal). If a provider left the TREWS page without entering an evaluation, the interaction is not counted as an evaluation in the context of the present study. We used these evaluations as our primary measure of provider interaction with TREWS and to define study and comparison arms.
Population
We conducted a prospective, multi-site, two-arm cohort study using EHR data from two academic and three community hospitals at which the TREWS alert system was deployed. Patients were eligible for inclusion if they were aged ≥18 years and presented to the ED or were admitted to an inpatient unit at one of the five deployment hospitals between deployment of TREWS and 30 September 2020. The specific hospitals and deployment dates were Howard County General Hospital (HCGH, 1 April 2018), Suburban Hospital (10 October 2018), Bayview Medical Center (BMC, 27 February 2019), Johns Hopkins Hospital (JHH, 16 April 2019) and Sibley Memorial Hospital (15 May 2019). We treated each time a patient presented to the ED or was admitted as a unique patient encounter and included each encounter separately.
For our primary analysis of patient outcomes, we included all patients who triggered an alert at most 1 h before admission or ED triage and met the criteria for sepsis based on a strategy called EHR-based sepsis phenotyping23,34. Briefly, this approach is a refinement of the criteria used in the third sepsis consensus definition (Sepsis-3)37 and the CDC Adult Sepsis Event Toolkit sepsis38, in which patients with features consistent with sepsis, but explained by other conditions (for example, hemorrhage) are excluded. As the full description of this sepsis definition is involved, we refer interested readers to the original work in which it appeared23.
To ensure that the alerts considered were actionable and to exclude outliers, we excluded patients who did not receive antibiotics within 24 h of the alert. See Extended Data Table 5 for a list of the antibiotics considered. Patients who had antibiotics administered before the alert were excluded, because we are interested primarily in patients for whom the alert may impact antibiotic administration timing and antibiotics given before the alert could not have been influenced by the alert. Finally, patients admitted directly to the ICU who did not pass through the ED were excluded because such patients were likely to have received previous treatment not reflected in the health record. A waterfall diagram for this cohort is shown in Fig. 1.
Study outcomes
The primary outcome was all-cause in-hospital mortality which was measured as the patient’s status at discharge. Secondary outcomes were the difference between a patient’s SOFA score at the time of the alert and 72 h after the alert and hospital length of stay after the alert until discharge among survivors39. SOFA scores were calculated using the worst measurements taken between 48 h and 72 h after the alert. For patients who died or were discharged before 72 h after the alert, the SOFA score was based on the worst measurements from the 24 h preceding death or discharge.
Study design
To evaluate the association between provider response to TREWS and patient outcomes, we compared outcomes between two arms: patients who had their alert evaluated and confirmed by a provider within 3 h of the alert (study arm) and patients who did not (comparison arm). The comparison arm is made up of patients who never had an evaluation entered into the TREWS page, as well as patients whose alert was dismissed. To account for potential differences between the arms, we adjusted for a variety of patient variables described in detail below30,31. We made this comparison among all included patients and in the cohort of high-risk patients (described in detail below). Patients who died during their stay were excluded from the length-of-stay analysis. Patients who left against medical advice, were discharged to a hospice or were transferred to another acute care facility were excluded from the SOFA progression analysis (all these patients were included in the mortality analysis).
Adjustment variables
The relationship between antibiotics timing and patient outcomes may be confounded by several aspects of patient presentation. We adjusted for a range of patient variables to account for potential confounding. Similar to previous studies7,8,25, we adjusted for patient age, documented sex, comorbidity history and measurements taken during the current encounter. To account for differences due to pre-existing conditions and patient history, we adjusted for the patient’s Charlson comorbidity index (CCI) as well as individual indicators for a history of diabetes (with and without complications), dementia, malignant tumors or metastatic solid tumors. These conditions were identified from International Classification of Diseases, 10th revision (ICD-10) codes included in the patient’s medical history49. To account for differences in treatment arising from patient presentation at the current visit, we adjusted for several sepsis-relevant labs, vital signs and treatments, including systolic blood pressure, Glasgow Coma Score (GCS) <15 (indicating any abnormal mental activity)8, temperature, white blood cell count, lactate >2 mmol l−1 and indicators for vasopressors and mechanical ventilation. We additionally adjusted for composite measures of acute patient severity, including individual SOFA score components and the APACHE II score39,40. For labs and vital signs, we used the most recent measurement taken in the 24 h before the alert and imputed a normal value if no measurement was observed during this period. SOFA and APACHE II scores were calculated using the worst measurements from the 24 h before the alert. In cases where the alert occurred within 12 h of presentation to the ED or admission to an inpatient unit (whichever came first), we used the worst measurements recorded within 12 h of presentation/admission to allow for lab processing and recording delays.
In addition to the variables used in previous studies, we included several additional patient and hospital-related variables. In medically complex patients, clinicians may wish to obtain more information on the patients before ordering antibiotics. To account for this, we further adjusted for the presence of several comorbidities that may obscure a sepsis diagnosis, including metastatic cancer, end-stage renal disease, congestive heart failure, acute liver disease, gastrointestinal bleeding and chronic obstructive pulmonary disease. These comorbidities were identified from ICD-10 codes listed as part of the patient’s problem list and marked as present on arrival. In addition, for ED patients, we adjusted for whether or not the trauma team was activated on arrival. To account for differences in clinical practice between hospitals, we included an indicator for the admitting hospital. Finally, we included a binary indicator for presentation at the hospital after 1 April 2020, to account for differences in treatment patterns arising from the COVID-19 pandemic.
High-risk cohort definition
In addition to the TREWS sepsis detection model, a separate model was developed for forecasting how much a patient’s mortality risk would increase without timely antibiotics (that is, sensitivity to antibiotic timing). Identifying patients with large increases in mortality risk may allow providers to prioritize alerts on patients who will benefit the most from timely care and allows us to examine heterogeneity in the associations between alert use and patient outcomes, that is, to perform stratification. This approach was inspired by previous work by VanderWeele et al. on identifying subgroups of patients who respond most to treatment26. VanderWeele et al. performed effect stratification by first developing a model to predict, based on baseline variables, whether a patient would respond to treatment and then estimated treatment effects among patients predicted to have a strong response. We applied a similar approach to our study by first developing a model to predict how much a patient’s mortality risk would increase without timely antibiotics (analogous to predicting treatment response) and then using this prediction to stratify the associations between interaction with TREWS and patient outcomes.
To predict sensitivity to antibiotic timing, we forecast each patient’s probability of in-hospital mortality with and without administration of antibiotics within 3 h of their alert and then took the ratio between these two probabilities. A higher ratio indicates a higher predicted sensitivity to antibiotic timing. The probability of in-hospital mortality with and without 3-h antibiotics was forecast using two separate ridge logistic regression models trained on retrospective EHR data from before deployment of TREWS. These data included 4,860 adult patients from our development dataset who were admitted to three of the five deployment hospitals (HCGH, JHH and BMC) between 1 January 2016 and 31 March 2018, who would have triggered an alert during their stay had the alert been active and who met all the inclusion criteria for our primary analysis (described above). Both models included as predictors all baseline variables described in the previous section except for the admitting hospital. All-cause in-hospital mortality was the binary prediction target. To account for nonlinearities, continuous lab values and vital signs were included as piecewise linear terms according to the thresholds used in APACHE II (ref. 40). The time from alert to first antibiotic administration was calculated on this retrospective data by running the TREWS model retrospectively to calculate the approximate time at which an alert would have occurred had TREWS been active for these patients. The two logistic regression models were trained separately on patients who did and did not receive an antibiotic within 3 h of when their alert would have occurred. To define a discrete high-risk cohort, a threshold for the ratio of mortality probabilities was chosen such that approximately one-third of patients in the retrospective data were identified as high risk.
The regularization strength for both models was tuned using grid search and 20 random 50:50 splits of the development data. For each random split, the models were trained and a high-risk threshold was chosen on the training set, the high-risk patients were identified on the test set and the adjusted association between timely antibiotics and in-hospital mortality was estimated among high-risk patients in the test set. Then, these estimates were averaged across the 20 random test sets, the regularization strength was chosen that maximized this average and the models were retrained using all of the retrospective data and the selected regularization strength.
To determine whether a patient in our study population fell into the high-risk cohort, both logistic regression models were applied to the patient’s baseline variables, and the patient was identified as high risk if the ratio of the two predicted probabilities was above the high-risk threshold. Note that these models were used to identify high-risk patients only for analysis and were not presented to the frontline users as part of the TREWS system.
Statistical analyses
For in-hospital mortality, we used logistic regression to estimate the ARD and ARR between study and comparison arms41. We used linear regression to estimate the adjusted difference in mean SOFA progression and quantile regression to estimate the adjusted difference in median length of stay42. To account for nonlinearities, continuous lab values and vital signs were included as piecewise linear terms according to the thresholds used in APACHE II (ref. 40). For all analyses, heteroskedasticity-robust estimators were used to estimate standard errors and construct CIs. All reported CIs are 95% CIs, and all P values correspond to two-sided tests. All statistical models were implemented via the Python packages ‘scikit-learn’ v.0.24.2 and ‘statsmodels’ v.0.12.1 (refs. 43,44).
Secondary and sensitivity analyses
As a secondary analysis, we estimated the association between time from alert to first antibiotic administration and patient outcomes. Despite previous work demonstrating an association between timely antibiotics and mortality when time-to-first antibiotics is measured from admission, onset or recognition5,6,7,8,9,10, it is important to establish that there is still an association when the TREWS alert is used as time zero and validate the potential to use a TREWS alert to initiate treatments in future studies. For example, suppose that providers can recognize most sepsis patients who need immediate antibiotics before the alert occurs. In this case, we might expect to see little association between antibiotic timing and patient outcomes in the remaining sepsis patients who did not receive antibiotics before their alert (that is, our study population). Using the same methods described above, we compared adjusted outcomes between patients in the target population who received their first antibiotic within 3 h of their alert with those who did not. In addition, for the purposes of direct comparison with previous work on antibiotic timing, we emulated the analyses from two previous retrospective studies on antibiotic timing7,25. These studies were chosen for their size and replicability. We estimated the per-hour adjusted odds ratio between time from alert to first antibiotic administration and in-hospital mortality using logistic regression and including all variables described above. We repeated this estimate including patients who receive their first antibiotics within 6 and 12 h, corresponding to the two previous studies.
Last of all, we tested the robustness of our analyses by repeating the primary analysis (adjusted comparison of patient outcomes between arms) under several different modifications. First, to ensure that our conclusions were not dependent on our choice of statistical method, we repeated the analysis using an alternative method, inverse probability of treatment weighting45,46,47. We used logistic regression for the probability of treatment model and stabilized and truncated weights at the 1st and 99th weight percentiles to reduce estimation variance48. Second, because a patient’s problem list reflects real-time diagnoses made by providers, it is less reliable than final diagnoses that reflect all information from a patient’s stay. Similarly, clinicians may not always mark a present-on-arrival diagnosis as such. We tested the sensitivity to these issues by replacing ICD-10 codes in the patient’s problem list with final diagnosis ICD-10 codes. Final diagnosis codes are more comprehensive but may include information from well after the patient’s alert, leading to potential over-adjustment. Third, because we included each patient encounter separately in our data, we wished to ensure that our conclusions were not overly sensitive to a few patients with multiple encounters. We tested this sensitivity by repeating the analysis using only the first encounter for each patient. Fourth, the COVID-19 pandemic impacted both characteristics of patients in our data and the care those patients received. We tested the sensitivity to inclusion of data during the COVID-19 pandemic by repeating the analysis using only patients admitted before 1 April 2020. Finally, the list of antibiotics used to determine the time of first antibiotic administration was intentionally inclusive. To ensure that our results are not sensitive to the specific antibiotics included, we repeated the analysis using a restricted list of antibiotics which removes some antibiotics that are unlikely to be used either to treat sepsis or on adult patients. The restricted list removes amoxicillin, azithromycin, cefotaxime, dapsone, erythromycin, neomycin and rifampin from the antibiotics listed in Extended Data Table 5.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data are not publicly available because they are from EHRs approved for limited use by Johns Hopkins University investigators. Making the data publicly available without additional consent, ethical or legal approval might compromise patients’ privacy and the original ethical approval. To perform additional analyses using this data, researchers should contact A.W.W. or S.S. to apply for an IRB-approved research collaboration and obtain an appropriate data use agreement.
Code availability
The TREWS early warning system described in the present study is available from Bayesian Health, New York. The underlying source code is proprietary intellectual property and is not available. Code for the primary statistical analyses and development of the high-risk cohort can be found at https://github.com/royadams/adams_et_al_2022_code.
References
Rhee, C. et al. Prevalence, underlying causes, and preventability of sepsis-associated mortality in US acute care hospitals. JAMA Netw. Open 2, e187571–e187571 (2019).
Riedemann, N. C., Guo, R. F. & Ward, P. A. The enigma of sepsis. J. Clin. Invest. 112, 460–467 (2003).
Marshall, J. C. Why have clinical trials in sepsis failed? Trends Mol. Med. 20, 195–203 (2014).
Rhodes, A. et al. Surviving Sepsis Campaign: International Guidelines for Management of Sepsis and Septic Shock: 2016. Crit. Care Med. 43, 304–377 (2017).
Kumar, A. et al. Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit. Care Med. 34, 1589–1596 (2006).
Ferrer, R. et al. Empiric antibiotic treatment reduces mortality in severe sepsis and septic shock from the first hour: results from a guideline-based performance improvement program. Crit. Care Med. 42, 1749–1755 (2014).
Liu, V. X. et al. The timing of early antibiotics and hospital mortality in sepsis. Am. J. Respir. Crit. Care Med. 196, 856–863 (2017).
Peltan, I. D. et al. ED door-to-antibiotic time and long-term mortality in sepsis. Chest 155, 938–946 (2019).
Chamberlain, D. J., Willis, E. M. & Bersten, A. B. The severe sepsis bundles as processes of care: a meta-analysis. Aust. Crit. Care 24, 229–243 (2011).
Damiani, E. et al. Effect of performance improvement programs on compliance with sepsis bundles and mortality: a systematic review and meta-analysis of observational studies. PLoS ONE 10, e0125827 (2015).
Giannini, H. M. et al. A machine learning algorithm to predict severe sepsis and septic shock: development, implementation, and impact on clinical practice. Crit. Care Med. 47, 1485–1492 (2019).
Desautels, T. et al. Prediction of sepsis in the intensive care unit with minimal electronic health record data: a machine learning approach. JMIR Med. Inform. 4, 1–15 (2016).
Shashikumar, S. P., Josef, C. S., Sharma, A. & Nemati, S. DeepAISE—an interpretable and recurrent neural survival model for early prediction of sepsis. Artif. Intell. Med. 113, 102036 (2021).
Horng, S. et al. Creating an automated trigger for sepsis clinical decision support at emergency department triage using machine learning. PLoS ONE 12, e0174708 (2017).
Bedoya, A. D. et al. Machine learning for early detection of sepsis: an internal and temporal validation study. JAMIA Open 3, 252–260 (2020).
Shimabukuro, D. W., Barton, C. W., Feldman, M. D., Mataraso, S. J. & Das, R. Effect of a machine learning-based severe sepsis prediction algorithm on patient survival and hospital length of stay: a randomised clinical trial. BMJ Open Respir. Res. 4, e000234 (2017).
McCoy, A. & Das, R. Reducing patient mortality, length of stay and readmissions through machine learning-based sepsis prediction in the emergency department, intensive care unit and hospital floor units. BMJ Open Qual. 6, e000158 (2017).
Escobar, G. J. et al. Automated identification of adults at risk for in-hospital clinical deterioration. N. Engl. J. Med. 383, 1951–1960 (2020).
Topiwala, R., Patel, K., Twigg, J., Rhule, J. & Meisenberg, B. Retrospective observational study of the clinical performance characteristics of a machine learning approach to early sepsis identification. Crit. Care Explor. 1, e0046 (2019).
Ginestra, J. C. et al. Clinician perception of a machine learning-based early warning system designed to predict severe sepsis and septic shock. Crit. Care Med. 47, 1477 (2019).
Henry, K. E., Hager, D. N., Pronovost, P. J. & Saria, S. A targeted real-time early warning score (TREWScore) for septic shock. Sci. Transl. Med. 7, 299ra122–299ra122 (2015).
Henry, K. E. et al. Factors driving provider adoption of the TREWS machine-learning-based early warning system and its effects on sepsis treatment timing. Nat. Med. https://doi.org/10.1038/s41591-022-01895-z (2022).
Henry, K. E., Hager, D. N., Osborn, T. M., Wu, A. W. & Saria, S. Comparison of automated sepsis identification methods and electronic health record-based sepsis phenotyping: improving case identification accuracy by accounting for confounding comorbid conditions. Crit. Care Explor. 1, e0053 (2019).
Rhee, C. et al. Infectious diseases society of america position paper: recommended revisions to the national severe sepsis and septic shock early management bundle (SEP-1) sepsis quality measure. Clin. Infect. Dis. 72, 541–552 (2021).
Seymour, C. W. et al. Time to treatment and mortality during mandated emergency care for sepsis. N. Engl. J. Med. 376, 2235–2244 (2017).
Vanderweele, T. J., Luedtke, A. R., Van Der Laan, M. J. & Kessler, R. C. Selecting optimal subgroups for treatment using many covariates. Epidemiology 30, 334–341 (2019).
Manaktala, S. & Claypool, S. R. Evaluating the impact of a computerized surveillance algorithm and decision support system on sepsis mortality. J. Am. Med. inform. Assoc. 24, 88–95 (2017).
Burdick, H. et al. Effect of a sepsis prediction algorithm on patient mortality, length of stay and readmission: a prospective multicentre clinical outcomes evaluation of real-world patient data from US hospitals. BMJ Health Care Inform. 27, e100109 (2020).
Guy, J. S., Jackson, E. & Perlin, J. B. Accelerating the clinical workflow using the sepsis prediction and optimization of therapy (SPOT) tool for real-time clinical monitoring. NEJM Catal. Innov. Care Deliv. https://doi.org/10.1056/CAT.19.1036 (2020).
Rosenbaum, P. R. & Briskman. Design of Observational Studies Vol. 10 (Springer, 2010).
Hernán, M. A. & Robins, J. M. Using big data to emulate a target trial when a randomized trial is not available. Am. J. Epidemiol. 183, 758–764 (2016).
Wong, A. et al. External validation of a widely implemented proprietary sepsis prediction model in hospitalized patients. JAMA Intern. Med. 48109, 1–6 (2021).
Henry, K. E. et al. Human-machine teaming is key to AI adoption: clinicians’ experiences with a deployed machine learning system. NPJ Digit. Med. https://doi.org/10.1038/s41746-022-00597-7 (2022).
Saria, S. & Henry, K. E. Too many definitions of sepsis: Can machine learning leverage the electronic health record to increase accuracy and bring consensus? Crit. Care Med. 48, 137–141 (2020). https://doi.org/10.1097/CCM.0000000000004144
Rhee, C. et al. Prevalence of antibiotic-resistant pathogens in culture-proven sepsis and outcomes associated with inadequate and broad-spectrum empiric antibiotic use. JAMA Netw. Open 3, e202899 (2020).
Jordan, M. I. & Jacobs, R. A. Hierarchical mixtures of experts and the EM algorithm. Proceedings of International Conference on Neural Networks 2, 1339–1344 (1993).
Seymour, C. W. et al. Assessment of clinical criteria for sepsis for the third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA 315, 762–774 (2016).
Rhee, C., Dantes, R. B., Epstein, L. & Klompas, M. Using objective clinical data to track progress on preventing and treating sepsis: CDC’s new adult sepsis event surveillance strategy. BMJ Qual. Saf. 28, 305–309 (2019).
Vincent, J. L. et al. The SOFA (Sepsis-related Organ Failure Assessment) score to describe organ dysfunction/failure. Intens. Care Med. 22, 707–710 (1996).
Knaus, W. A., Draper, E. A., Wagner, D. P. & Zimmerman, J. E. APACHE II: a severity of disease classification system. Crit. Care Med. 13, 818–829 (1985).
Norton, E. C., Miller, M. M. & Kleinman, L. C. Computing adjusted risk ratios and risk differences in Stata. Stata J. 13, 492–509 (2013).
Peng, L. Quantile regression for survival data. Annu. Rev. Stat. Its Appl. 8, 413–437 (2021).
Seabold, S. & Perktold, J. statsmodels: econometric and statistical modeling with python. In van der Walt, S. & Millman, J. (Eds.) Proc. 9th Python in Science Conference 92–96 (2010).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Horvitz, D. G. & Thompson, D. J. A generalization of sampling without replacement from a finite universe. J. Am. Stat. Assoc. 47, 663–685 (1952).
Robins, J. M. Marginal structural models versus structural nested models as tools for causal inference. In Halloran, M. E. & Berry, D. (Eds.) Statistical Models in Epidemiology, the Environment, and Clinical Trials 95–133 (Springer, 2000).
Hernán, M. A. & Robins, J. M. Causal Inference: What If (Chapman & Hall/CRC, 2020).
Lee, B. K., Lessler, J. & Stuart, E. A. Weight trimming and propensity score weighting. PLoS ONE 6, e18174 (2011).
World Health Organization. ICD-10 : international statistical classification of diseases and related health problems : tenth revision (World Health Organization, 2004).
Acknowledgements
We thank Y. Ahmad, M. Yeo and Y. Karklin whose work significantly contributed to early iterations of the development of the deployed system. Further, we thank R. Demski, K. D’Souza, A. Kachalia, A. Chen and clinical and quality stakeholders who contributed to tool deployment, education and championing the work. We gratefully acknowledge the following sources of funding: the Gordon and Betty Moore Foundation (award no. 3926), the National Science Foundation (NSF) Future of Work at the Human-technology Frontier (award no. 1840088) and the Alfred P. Sloan Foundation research fellowship (2018). This information or content and conclusions are those of the authors and should not be construed as the official position or policy of, nor should any endorsements be inferred by, the NSF of the US Government.
Author information
Authors and Affiliations
Contributions
K.E.H., R.A., A.W.W. and S.S. contributed to the initial study design and preliminary analysis plan. S.S. led the development and deployment efforts for the TREWS software. K.E.H., H.S., N.R., A.Z., A.S., R.C.L., L.J., M.H., S.M., D.N.H., A.R.A., A.W.W. and S.S. contributed to the system deployment. K.E.H., R.A., E.Y.K., S.E.C., E.S.C., D.N.H., A.W.W. and S.S. contributed to the review and analysis of the results. All authors contributed to the final preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Under a license agreement between Bayesian Health and the Johns Hopkins University, K.E.H., S.S. and Johns Hopkins University are entitled to revenue distributions. In addition, the university owns equity in Bayesian Health. This arrangement has been reviewed and approved by the Johns Hopkins University in accordance with its conflict-of-interest policies. S.S. also has grants from the Gordon and Betty Moore Foundation, the NSF, the National Institutes of Health, Defense Advanced Research Projects Agency, the Food and Drug Administration and the American Heart Association; she is a founder of and holds equity in Bayesian Health; she is the scientific advisory board member for PatientPing; and she has received honoraria for talks from a number of biotechnology, research and health-tech companies. This arrangement has been reviewed and approved by the Johns Hopkins University in accordance with its conflict-of-interest policies. D.N.H. discloses salary support and funding to his institution from the Marcus Foundation for the conduct of the vitamin C, thiamine and steroids in sepsis trial. S.E.C. received consulting fees from Basilea for work on an infection adjudication committee for a Staphylococcus aureus bacteremia trial. The remaining authors declare no disclosures of conflicts of interest.
Peer review
Peer review information
Nature Medicine thanks Derek Angus, Melanie Wright and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling editor: Michael Basson in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Supplementary information
Rights and permissions
About this article
Cite this article
Adams, R., Henry, K.E., Sridharan, A. et al. Prospective, multi-site study of patient outcomes after implementation of the TREWS machine learning-based early warning system for sepsis. Nat Med 28, 1455–1460 (2022). https://doi.org/10.1038/s41591-022-01894-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01894-0
This article is cited by
-
Use of artificial intelligence in critical care: opportunities and obstacles
Critical Care (2024)
-
Artificial intelligence applications in histopathology
Nature Reviews Electrical Engineering (2024)
-
Measuring algorithmic bias to analyze the reliability of AI tools that predict depression risk using smartphone sensed-behavioral data
npj Mental Health Research (2024)
-
Impact of a deep learning sepsis prediction model on quality of care and survival
npj Digital Medicine (2024)
-
Real-time machine learning-assisted sepsis alert enhances the timeliness of antibiotic administration and diagnostic accuracy in emergency department patients with sepsis: a cluster-randomized trial
Internal and Emergency Medicine (2024)