Abstract
The treatment landscape of acute myeloid leukemia (AML) is evolving, with promising therapies entering clinical translation, yet patient responses remain heterogeneous, and biomarkers for tailoring treatment are lacking. To understand how disease heterogeneity links with therapy response, we determined the leukemia cell hierarchy makeup from bulk transcriptomes of more than 1,000 patients through deconvolution using single-cell reference profiles of leukemia stem, progenitor and mature cell types. Leukemia hierarchy composition was associated with functional, genomic and clinical properties and converged into four overall classes, spanning Primitive, Mature, GMP and Intermediate. Critically, variation in hierarchy composition along the Primitive versus GMP or Primitive versus Mature axes were associated with response to chemotherapy or drug sensitivity profiles of targeted therapies, respectively. A seven-gene biomarker derived from the Primitive versus Mature axis was associated with response to 105 investigational drugs. Cellular hierarchy composition constitutes a novel framework for understanding disease biology and advancing precision medicine in AML.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Processed and raw RNA-seq data generated in this study are available in the GEO under Superseries GSE199336. Citations and links of re-analyzed data from all clinical datasets analyzed in this study are provided in Supplementary Table 4. Citations and links of re-analyzed data from the literature screen are provided in Supplementary Table 13. Pathways and signatures used for relapse benchmarking were obtained from the MSigDB (https://www.gsea-msigdb.org/gsea/msigdb/). All deconvolution results are available with the analysis code on GitHub (https://github.com/andygxzeng/AMLHierarchies).
Code availability
Analysis notebooks for the main figures, as well as instructions for applying AML deconvolution, are available at: https://github.com/andygxzeng/AMLHierarchies.
References
Hungerford, D. A. & Nowell, P. C. A minute chromosome in human chronic granulocytic leukemia. Science 132, 1497–1499 (1960).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Klco, J. M. et al. Functional heterogeneity of genetically defined subclones in acute myeloid leukemia. Cancer Cell 25, 379–392 (2014).
Till, J. E. & McCULLOCH, E. A. A direct measurement of the radiation sensitivity of normal mouse bone marrow cells. Radiat. Res. 14, 213–222 (1961).
Clarkson, B., Ohkita, T., Ota, K. & Fried, J. Studies of cellular proliferation in human leukemia. I. Estimation of growth rates of leukemic and normal hematopoietic cells in two adults with acute leukemia given single injections of tritiated thymidine. J. Clin. Invest. 46, 506–529 (1967).
Minden, M. D., Till, J. E. & McCulloch, E. A. Proliferative state of blast cell progenitors in acute myeloblastic leukemia (AML). Blood 52, 592–600 (1978).
Griffin, J. D., Larcom, P. & Schlossman, S. F. Use of surface markers to identify a subset of acute myelomonocytic leukemia cells with progenitor cell properties. Blood 62, 1300–1303 (1983).
Wouters, R. & Löwenberg, B. On the maturation order of AML cells: a distinction on the basis of self-renewal properties and immunologic phenotypes. Blood 63, 684–689 (1984).
Buick, R. N., Minden, M. D. & McCulloch, E. A. Self-renewal in culture of proliferative blast progenitor cells in acute myeloblastic leukemia. Blood 54, 95–104 (1979).
Chang, L. J., Till, J. E. & McCulloch, E. A. The cellular basis of self renewal in culture by human acute myeloblastic leukemia blast cell progenitors. J. Cell. Physiol. 102, 217–222 (1980).
McCulloch, E. A. Stem cells in normal and leukemic hemopoiesis (Henry Stratton Lecture, 1982). Blood 62, 1–13 (1983).
Lapidot, T. et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367, 645–648 (1994).
Bonnet, D. & Dick, J. E. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat. Med. 3, 730–737 (1997).
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Gentles, A. J., Plevritis, S. K., Majeti, R. & Alizadeh, A. A. Association of a leukemic stem cell gene expression signature with clinical outcomes in acute myeloid leukemia. JAMA 304, 2706–2715 (2010).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Ng, S. W. K. et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016).
Elsayed, A. H. et al. A six-gene leukemic stem cell score identifies high risk pediatric acute myeloid leukemia. Leukemia 34, 735–745 (2020).
Pierce, G. B. & Speers, W. C. Tumors as caricatures of the process of tissue renewal: prospects for therapy by directing differentiation. Cancer Res. 48, 1996–2004 (1988).
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
van Galen, P. et al. Single-cell RNA-seq reveals AML hierarchies relevant to disease progression and immunity. Cell 176, 1265–1281 (2019).
Wu, J. et al. A single-cell survey of cellular hierarchy in acute myeloid leukemia. J. Hematol. Oncol. 13, 128 (2020).
Tarashansky, A. J., Xue, Y., Li, P., Quake, S. R. & Wang, B. Self-assembling manifolds in single-cell RNA sequencing data. eLife 8, e48994 (2019).
Xie, S. Z. et al. Sphingolipid modulation activates proteostasis programs to govern human hematopoietic stem cell self-renewal. Cell Stem Cell 25, 639–653 (2019).
Xie, S. Z. et al. Sphingosine-1-phosphate receptor 3 potentiates inflammatory programs in normal and leukemia stem cells to promote differentiation. Blood Cancer Discov. 2, 32–53 (2021).
Takayama, N. et al. The transition from quiescent to activated states in human hematopoietic stem cells is governed by dynamic 3D genome reorganization. Cell Stem Cell 28, 488–501 (2021).
Hope, K. J., Jin, L. & Dick, J. E. Acute myeloid leukemia originates from a hierarchy of leukemic stem cell classes that differ in self-renewal capacity. Nat. Immunol. 5, 738–743 (2004).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Dick, J. E. Stem cell concepts renew cancer research. Blood 112, 4793–4807 (2008).
Quek, L. et al. Genetically distinct leukemic stem cells in human CD34− acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J. Exp. Med. 213, 1513–1535 (2016).
Pabst, C. et al. GPR56 identifies primary human acute myeloid leukemia cells with high repopulating potential in vivo. Blood 127, 2018–2027 (2016).
Cancer Genome Atlas Research Network et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Tyner, J. W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526–531 (2018).
Marquis, M. et al. High expression of HMGA2 independently predicts poor clinical outcomes in acute myeloid leukemia. Blood Cancer J. 8, 68 (2018).
Verhaak, R. G. W. et al. Prediction of molecular subtypes in acute myeloid leukemia based on gene expression profiling. Haematologica 94, 131–134 (2009).
Bolouri, H. et al. The molecular landscape of pediatric acute myeloid leukemia reveals recurrent structural alterations and age-specific mutational interactions. Nat. Med. 24, 103–112 (2017).
Docking, T. R. et al. A clinical transcriptome approach to patient stratification and therapy selection in acute myeloid leukemia. Nat. Commun. 12, 2474 (2021).
Wagner, S. et al. A parsimonious 3-gene signature predicts clinical outcomes in an acute myeloid leukemia multicohort study. Blood Adv. 3, 1330–1346 (2019).
Nehme, A. et al. Horizontal meta-analysis identifies common deregulated genes across AML subgroups providing a robust prognostic signature. Blood Adv. 4, 5322–5335 (2020).
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016).
Christopher, M. J. et al. Immune escape of relapsed AML cells after allogeneic transplant. N. Engl. J. Med. 379, 2330–2341 (2018).
Cocciardi, S. et al. Clonal evolution patterns in acute myeloid leukemia with NPM1 mutation. Nat. Commun. 10, 2031 (2019).
Abbas, H. A. et al. Single cell T cell landscape and T cell receptor repertoire profiling of AML in context of PD-1 blockade therapy. Nat. Commun. 12, 6071 (2021).
Vosberg, S. & Greif, P. A. Clonal evolution of acute myeloid leukemia from diagnosis to relapse. Genes Chromosomes Cancer 58, 839–849 (2019).
Lee, S.-I. et al. A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia. Nat. Commun. 9, 42 (2018).
Ng, S. W. K. et al. A clinical laboratory-developed LSC17 stemness score assay for rapid risk assessment of patients with acute myeloid leukemia. Blood Adv. 6, 1064–1073 (2022).
Castaigne, S. et al. Effect of gemtuzumab ozogamicin on survival of adult patients with de-novo acute myeloid leukaemia (ALFA-0701): a randomised, open-label, phase 3 study. Lancet 379, 1508–1516 (2012).
Lambert, J. et al. Gemtuzumab ozogamicin for de novo acute myeloid leukemia: final efficacy and safety updates from the open-label, phase III ALFA-0701 trial. Haematologica 104, 113–119 (2019).
Brunetti, L. et al. Mutant NPM1 maintains the leukemic state through HOX expression. Cancer Cell 34, 499–512 (2018).
Etchin, J. et al. Activity of a selective inhibitor of nuclear export, selinexor (KPT-330), against AML-initiating cells engrafted into immunosuppressed NSG mice. Leukemia 30, 190–199 (2016).
Chen, W. C. et al. An integrated analysis of heterogeneous drug responses in acute myeloid leukemia that enables the discovery of predictive biomarkers. Cancer Res. 76, 1214–1224 (2016).
Surka, C. et al. CC-90009, a novel cereblon E3 ligase modulator, targets acute myeloid leukemia blasts and leukemia stem cells. Blood 137, 661–677 (2021).
Pei, S. et al. Monocytic subclones confer resistance to venetoclax-based therapy in patients with acute myeloid leukemia. Cancer Discov. 10, 536–551 (2020).
Lun, A. T. L., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Tan, Y. & Cahan, P. SingleCellNet: a computational tool to classify single cell RNA-seq data across platforms and across species. Cell Syst. 9, 207–213 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2006).
Piccolo, S. R. et al. A single-sample microarray normalization method to facilitate personalized-medicine workflows. Genomics 100, 337–344 (2012).
Xu, C. et al. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Mol. Syst. Biol. 17, e9620 (2021).
Lotfollahi, M. et al. Mapping single-cell data to reference atlases by transfer learning. Nat. Biotechnol. 40, 121–130 (2022).
Deshwar, A. G. et al. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 16, 35 (2015).
Miller, C. A. et al. Visualizing tumor evolution with the fishplot package for R. BMC Genomics 17, 880 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Galkin, O. et al. SIRPαFc treatment targets human acute myeloid leukemia stem cells. Haematologica 106, 279–283 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank C. Jones, A. Tikhonova, N. Iscove, S. Chan, S. Abelson, B. Haibe-Kains, M. Anders and all members of the Dick Laboratory for valuable feedback on the manuscript. We thank P. Valk and the HOVON-SAKK trial group for providing the clinical annotations associated with GSE6891. A.G.X.Z. is supported by a CIHR Vanier scholarship. J.E.D. is supported by funds from the Ontario Institute for Cancer Research through funding provided by the Government of Ontario; the Canadian Institutes for Health Research (RN380110 - 409786); the International Development Research Centre Ottawa Canada; the Canadian Cancer Society (grant 703212 (end date 2019) and grant 706662 (end date 2025)); the Terry Fox New Frontiers Program Project Grant (Project 1106); University of Toronto’s Medicine by Design initiative with funding from the Canada First Research Excellence Fund; a Canada Research Chair; Princess Margaret Cancer Centre; The Princess Margaret Cancer Foundation; and the Ontario Ministry of Health. The Leukemia Tissue Bank is supported by funds to M.D.M. from the Wendy Eisen Princess Margaret Hospital Foundation Account and the Philip Orsino Chair in Leukemia Research. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.G.X.Z. and J.E.D. conceived the project. A.G.X.Z. carried out the analysis. S.B. contributed to analyses of post-treatment changes in cell composition. L.J. performed fedratinib and CC-90009 combination treatments. A.M. and W.C.C. performed RNA-seq extraction and library preparation for LSC fractions and fedratinib samples, respectively. M.C.-S.-Y. and V.V. performed alignment and pre-processing for LSC fractions and fredratinib RNA-seq data. H.A.A., N.D. and A.F. provided scRNA-seq data and leukemic cell annotations for relapsed AMLs. P.v.G. provided bulk RNA-seq data for AML samples. A.T. analyzed clinical flow data from the PMH Leukemia cohort. M.C., C.P. and H.D. provided GE and clinical data for the ALFA-0701 trial cohort. M.D.M. provided PMH AML samples. J.A.K. and M.D.M. provided clinical annotations for the PMH AML cohort. A.G.X.Z., J.A.K., J.C.Y.W. and J.E.D. interpreted the data. A.G.X.Z. and J.E.D. wrote the paper. J.A.K. and J.C.Y.W. revised the paper.
Corresponding author
Ethics declarations
Competing interests
J.E.D. reports research funding from Celgene/Bristol Myers Squibb and advisory board/royalities from Trillium Therapeutics. All other authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Craig Jordan, Simon Haas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Editor recognition statement: primary handling editors were Javier Carmona and Michael Basson, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Features of leukemia stem and progenitor cell populations from scRNA-seq.
A) UMAP and PCA embeddings of 4163 AML LSPCs after feature weight derivation with the Self-Assembling Manifolds (SAM) algorithm. B-F) Diffusion map of re-annotated LSPC populations using SAM-derived feature weights, depicting: B) patient identity, C) prior cell type annotation, D) enrichment of LSC-specific genes from Ng et al (2016) and Shannon Diversity Index, E) scaled CDK6 expression and enrichment of the E2F3 regulon, and F) enrichment of E2F1 and CTCF regulons. G) Cell cycle status of Quiescent (n = 1855), Primed (n = 1240), and Cycling LSPCs (n = 1068). H) Enrichment of inflammatory signaling pathways and regulons in LSPCs. I) Transcription factor regulon activity, inferred through pySCENIC, specific to each LSPC. J) Normalized confusion matrix depicting classifier accuracy of prior and new cell type annotations for primitive AML cells. The classifier was built using SingleCellNet, an Ensemble classifier for scRNA-seq data trained from the top pairs of genes unique to each cell type. 800 cells from each cell type were used for training and 250 were used for validation.
Extended Data Fig. 2 Benchmarking gene expression deconvolution approaches for AML.
A) Pearson correlation between observed relative abundance of 14 cell types from scRNA-seq and predicted abundance of each cell type from deconvolution of matched bulk RNA-seq data, analyzed by patient. Gene expression deconvolution using CIBERSORTx (S-mode or No Batch Correction), DWLS, Bisque, or MuSIC (direct or recursive) were benchmarked across these samples. B) scRNA-seq of 1389 ProMono-like cells across 10 patients, demonstrating separation between AML556 and other patients. C) Pearson correlation depicting deconvolution performance of CIBERSORTx for AML556’s bulk RNA-seq profile using patient-specific reference signatures derived from scRNA-seq data from AML556. D-E) Correlation between deconvolution and clinical flow cytometry for 7 AML patients from the Toronto PMH cohort. Deconvolution using scRNA-seq reference profiles was performed on RNA-seq data and matched with clinical flow cytometry data, both obtained from peripheral blood. D) Pearson correlation between total mature myeloid abundance (ProMono-like + Mono-like + cDC-like) from deconvolution with pan-myeloid surface marker CD64. E) Pearson correlation between mono-like abundance from deconvolution with monocyte-specific surface marker CD14. F) Dendrogram depicting associations between leukemic cell-types across scRNA-seq samples from 12 diagnostic AML patients. G) Observed associations between leukemic cell types from deconvolution analysis of 173 patients within the TCGA cohort, depicted for each deconvolution tool. MuSIC Direct was excluded due to multiple cell types having a detection rate of zero in bulk RNA-seq. H-I) Correlation between observed transcriptomic profiles and synthetic transcriptomic profiles reconstructed based on predicted cell-type abundance from CIBERSORTx. Higher correlation suggests greater deconvolution confidence. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range). Comparisons were performed through two-sided Wilcoxon signed-rank tests. These correlations are depicted for H) Deconvolution of 864 patients across three AML cohorts using reference signatures from leukemic populations compared to deconvolution with reference signatures from matched healthy populations, and I) RNA-seq compared to microarray from 158 matched TCGA patient samples. Prior to deconvolution, microarray data was normalized through either chip-based (RMA) or single-sample (SCAN) normalization approaches. J) Pearson correlation of estimated LSPC abundances between RNA-seq deconvolution and Microarray deconvolution, normalized with either RMA or SCAN, among 158 matched patient samples from the TCGA cohort.
Extended Data Fig. 3 Comparison of new and prior leukemia stem and progenitor cell annotations for discerning biological phenotypes.
A) Workflow to compare prior (HSC-like and Prog-like) annotations and new (Quiescent LSPC, Primed LSPC, and Cycling LSPC) annotations with regard to their utility in predicting important biological phenotypes in AML. This was measured through the performance of logistic regression and random forest models trained on the relative abundance of these populations. Models were trained using nested cross-validation wherein samples were subject to a 5-fold split, in which 80% of samples (white boxes) were used to train a model and 20% of samples (orange boxes) were used to evaluate the model. Within each training set, hyperparameter optimization was performed by grid search with 5-fold internal cross validation. The model AUC from each outer cross-validation split was averaged to estimate overall classifier performance. This nested cross-validation process was repeated over 1000 iterations, with samples being shuffled between each iteration, to generate a distribution of AUC metrics. B-F) Model performance for predicting key biological and clinical phenotypes from either HSC-like and Prog-like abundance or Quiescent, Primed, and Cycling LSPC abundance. Performance metrics are paired by iteration, wherein sample order and cross validation splits were identical for each model. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range). Statistical significance was evaluated through a two-sided Wilcoxon signed-rank test. B) Prediction of functional LSC activity measured by leukemic engraftment from 72 LSC+ fractions and 38 LSC- fractions. C) Prediction of overall survival in the TCGA and BEAT-AML cohorts, evaluated through the likelihood ratio statistic from stratified cox regression (combined n = 454). In this case LASSO and Ridge regression was performed and these models were trained on splits of the TCGA and BEAT-AML cohorts, stratified by cohort. The repeated nested cross validation approach remained the same. D) Prediction of diagnosis vs relapse status from 44 relapsed and 44 diagnostic samples. E) Prediction of Adverse cytogenetic status in TCGA from 37 patients with Adverse cytogenetics and 131 patients with Intermediate or Favorable cytogenetics. F) Prediction of Adverse cytogenetic status in BEAT-AML from 53 patients with Adverse cytogenetics and 175 patients with Intermediate or Favorable cytogenetics.
Extended Data Fig. 4 Biological and genomic correlates of AML hierarchies.
A) Correlation between deconvoluted abundance of leukemic and immune cell types with clinical features in TCGA (n = 173). Only correlations with P < 0.05 are depicted, and correlations with FDR < 0.05 are noted with an asterisk. B) Cellular hierarchy projections of patient samples classified as FAB M0 (n = 30), M1 (n = 122), M2 (n = 77), M3 (n = 30), M4 (n = 58), M4Eo (n = 22), M5A (n = 28), or M5B (n = 11) C) Density plots depicting all mutation combinations along the Primitive versus Mature axis (PC2). D) Density plots depicting all mutation combinations along the Primitive versus GMP axis (PC1). E) Density plots depicting all cytogenetic alterations along the Primitive versus Mature axis (PC2). F-G) Impact of DNMT3A R882 mutations compared to other DNMT3A mutations on leukemic hierarchy organization along the Primitive versus Mature axis (PC2). F) Boxplot comparing PC2 of AMLs with DNMT3A R882 mutations (n = 84) compared to other DNMT3A (n = 96) mutations, split by mutational partner. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range). Statistical significance was evaluated through a two-sided Wilcoxon rank sum test. G) Density plot depicting PC2 of mutational combinations with either DNMT3A R882 or other DNMT3A mutations.
Extended Data Fig. 5 The Primitive versus GMP axis governs AML prognosis.
A) 495 adult AML patients from GSE6891, projected by hierarchy composition and classified based on the reference cohorts. B) Overall survival outcomes of 495 AML patients from GSE6891, stratified by hierarchy subtype. C) 287 pediatric AML patients from the TARGET-AML cohort, projected by hierarchy composition and classified based on the reference cohorts. D) Overall survival outcomes 287 pediatric AML patients from TARGET-AML, stratified by hierarchy subtype. E) Correlation between a prognostic score trained by regularized cox regression using leukemic cell type abundances with the Primitive versus GMP axis (PC1) within the TCGA and BEAT-AML cohorts (combined n = 454). F) Overall survival outcomes of patients stratified by PC1 within the TCGA (n = 173), BEAT-AML (n=281), and GSE6891 (n = 495) cohorts. G) Association between cell-type abundance and induction failure in four independent studies: Bolouri et al 2018 (pediatric AML; n = 257 remission, n = 30 induction failure), Chiu et al 2019 (adult AML; n = 18 remission, n = 18 induction failure), Herold et al 2018 (adult AML; n = 164 remission, n = 86 induction failure), and Tyner et al 2018 (adult AML; n = 140 remission, n = 63 induction failure), represented through the test statistic from a two-sided Wilcoxon rank sum test. Green denotes higher relative abundance in induction failure patients compare to patients who achieved complete remission, while purple denotes lower relative abundance in induction failure patients. Differences with an uncorrected P < 0.10 are noted with an asterisk. H) Correlation between four prognostic AML scores with the relative abundance of each leukemic cell type across the TCGA, BEAT-AML, and Leucegene cohorts (combined n = 864). I) Relative abundance of Quiescent LSPC and GMP-like blasts among 864 AML patients split into high and low risk categories by median split for four prognostic scores. Significance was evaluated through two-sided Wilcoxon rank-sum tests. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range). J) GSEA analysis with gene signatures derived from the Primitive versus GMP axis in normal and malignant hematopoiesis, performed on genes ranked by univariate associations with overall survival within the TCGA and BEAT-AML cohorts.
Extended Data Fig. 6 Changes in hierarchy composition at AML relapse.
A) Hierarchy composition of 44 matched diagnosis and relapse pairs. Top row depicts hierarchy composition at diagnosis while the bottom row depicts hierarchy composition at relapse. Samples from the same patient are aligned vertically. B) Relative abundance of each leukemic cell population from scRNA-seq of 12 diagnostic AMLs (van Galen et al, 2019) compared with 8 relapsed AMLs (Abbas et al, 2021). Statistical significance was evaluated through two-sided Wilcoxon rank-sum tests. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range). C-F) Evolution of paired diagnosis and relapse AML samples depicted through shifts in cellular hierarchies, evolution of genetic subclones, and changes in cell-type composition. C) Patient 303642, in which significant genetic evolution is accompanied by a dramatic shift in cellular hierarchy from GMP to primitive. D) Patient 1019, in which replacement of an NRAS and IDH2 positive clone with an IDH1 positive clone is associated with a modest shift in cellular hierarchy. E) Patient 4, in which a loss of monocytic blasts is accompanied by a modest decrease in the the size of an NRAS bearing clone. F) Patient 150288, in which extensive linear genetic evolution is not associated with any appreciable change in cell type composition.
Extended Data Fig. 7 The Primitive versus Mature axis governs ex vivo drug sensitivity.
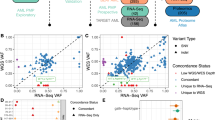
A) Volcano plot depicting associations between Primitive versus GMP axis (PC1) and ex vivo drug sensitivity from the BEAT-AML (Tyner et al 2018) drug screen (n = 202 patients), and between PC1 and PC2 and ex vivo drug sensitivity from Lee et al 2018 (n = 30 patients). B) Unsupervised clustering of 30 primary AML patients from Lee et al on the basis of ex vivo sensitivity to 159 drugs. Drug sensitivity values (scaled negative AUC) are depicted for the top drugs corresponding to each patient cluster. Red denotes sensitivity while blue denotes lower sensitivity. C) GSEA analysis with gene signatures derived from the Primitive vs Mature axis in normal and malignant hematopoiesis, performed on genes ranked by differential expression between the two drug response clusters from (B). D) Correlations of AML gene expression scores with ex vivo drug sensitivities from two drug screens (Tyner et al, n = 202; Lee et al, n = 30) performed on primary AML samples. E) Ex vivo drug sensitivity to Venetoclax (n = 114 from Tyner et al) and Azacytidine (n = 30 from Lee et al) of primary patient samples grouped into ‘High’ or ‘Low’ based on median splits of patient scores for each AML gene expression score. Significance was evaluated through two-sided Wilcoxon rank-sum tests. Box plots indicate the range of the central 50% of the data, with the central line marking the median. Whiskers extend from each box to 1.5*(interquartile range).
Extended Data Fig. 8 The Primitive versus Mature axis predicts clinical benefit from Gemtuzumab-Ozogamicin in both adult and pediatric AML.
A) Subgroup analysis of randomized clinical trial ALFA-0701 (n = 192) after stratification by PC2-34. Event-free survival (EFS) and Relapse-free survival (RFS) curves comparing chemotherapy only (Control arm) against chemotherapy + Gemtuzumab-Ozogamicin (GO arm). B) Lack of correlation between CD33 levels by flow cytometry and Primitive versus Mature axis (PC2 and PC2-34 score), evaluated across 23 Toronto PMH AML patients for which both RNA-seq and clinical flow information was available. C) Stratification of ALFA-0701 patients on the basis of both LSC17 and LinClass-7. EFS and RFS for the LinClass-7 Low (Mature > Primitive) and LSC17 Low subgroup (n = 56) is depicted as this was the only group to derive significant benefit from GO treatment. D) Stratification of ALFA-0701 patients on the basis of both LSC17 and PC2-34. EFS and RFS for the PC2-34 High (Mature > Primitive) and LSC17 Low subgroup (n = 54) is depicted as this was the only group to derive significant benefit from GO treatment. E-F) Subgroup analysis of a retrospective cohort of pediatric AML patients treated with either GO (n = 154) or Chemo (n = 91), stratified by the PC2 Primitive versus Mature axis and related gene expression scores. Outcomes are depicted for both overall survival (E) and event-free survival (F).
Extended Data Fig. 9 LinClass-7 as a companion score for LSC17.
A) LSC17 and LinClass-7 scores of 864 AML patients by RNA-seq. Patients belonging to each hierarchy subtype (Primitive, Intermediate, GMP, Mature) are also depicted. (B) LSC17 and LinClass-7 scores measured through a 17-gene NanoString assay. Normalized NanoString-derived LSC17 and LinClass-7 scores from 306 Toronto PMH patients from Ng et al (2016) are depicted.
Extended Data Fig. 10 Literature screen to identify treatment-induced changes in cellular composition.
A) ComplexHeatmap depicting changes in cell type composition following drug treatment from 43 preclinical studies in human AML, represented as the absolute log(P value) from a two-sided Wilcoxon rank sum test in the direction of the change. Green depicts an increase in cell type abundance and purple depicts a decrease in cell type abundance. Each treatment is labeled with its target(s) in parentheses. Changes in PC2 (Primitive versus Mature) are depicted above the heatmap, and candidate differentiation drugs (increase in PC2 with uncorrected P < 0.05) are denoted with an asterisk. AML sample type and key genomic characteristics are also depicted for each treatment condition. B) UMAP coordinates for each drug treatment condition depicting changes in each cell type after drug treatment compared to control, represented as the absolute log(P value) from a two-sided Wilcoxon rank sum test in the direction of the change. Tissue source (Primary vs Cell Line), MLL translocation status, and drug target are also depicted for each treatment condition.
Supplementary information
Supplementary Information
Supplementary table captions and Supplementary Notes 1 and 2
Supplementary Table
Supplemntary Tables 1–14
Rights and permissions
About this article
Cite this article
Zeng, A.G.X., Bansal, S., Jin, L. et al. A cellular hierarchy framework for understanding heterogeneity and predicting drug response in acute myeloid leukemia. Nat Med 28, 1212–1223 (2022). https://doi.org/10.1038/s41591-022-01819-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01819-x
This article is cited by
-
Decoding leukemia at the single-cell level: clonal architecture, classification, microenvironment, and drug resistance
Experimental Hematology & Oncology (2024)
-
Revealing the role of SPP1+ macrophages in glioma prognosis and therapeutic targeting by investigating tumor-associated macrophage landscape in grade 2 and 3 gliomas
Cell & Bioscience (2024)
-
Mapping AML heterogeneity - multi-cohort transcriptomic analysis identifies novel clusters and divergent ex-vivo drug responses
Leukemia (2024)
-
Cellular hierarchy insights reveal leukemic stem-like cells and early death risk in acute promyelocytic leukemia
Nature Communications (2024)
-
Heterogeneity of hepatocellular carcinoma: from mechanisms to clinical implications
Cancer Gene Therapy (2024)