Abstract
The pathophysiology of adverse events following programmed cell death protein 1 (PD-1) blockade, including tuberculosis (TB) and autoimmunity, remains poorly characterized. We studied a patient with inherited PD-1 deficiency and TB who died of pulmonary autoimmunity. The patient’s leukocytes did not express PD-1 or respond to PD-1-mediated suppression. The patient’s lymphocytes produced only small amounts of interferon (IFN)-γ upon mycobacterial stimuli, similarly to patients with inborn errors of IFN-γ production who are vulnerable to TB. This phenotype resulted from a combined depletion of Vδ2+ γδ T, mucosal-associated invariant T and CD56bright natural killer lymphocytes and dysfunction of other T lymphocyte subsets. Moreover, the patient displayed hepatosplenomegaly and an expansion of total, activated and RORγT+ CD4−CD8− double-negative αβ T cells, similar to patients with STAT3 gain-of-function mutations who display lymphoproliferative autoimmunity. This phenotype resulted from excessive amounts of STAT3-activating cytokines interleukin (IL)-6 and IL-23 produced by activated T lymphocytes and monocytes, and the STAT3-dependent expression of RORγT by activated T lymphocytes. Our work highlights the indispensable role of human PD-1 in governing both antimycobacterial immunity and self-tolerance, while identifying potentially actionable molecular targets for the diagnostic and therapeutic management of TB and autoimmunity in patients on PD-1 blockade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
MAFs of PDCD1 variants in the general population were retrieved from gnomAD r2.1.1 (https://gnomad.broadinstitute.org/). Public scRNA-seq datasets were downloaded from the 10X Genomics website (https://support.10xgenomics.com/single-cell-gene-expression/datasets/). For GSEA, gene sets were obtained from MSigDB Collections (http://www.gsea-msigdb.org/gsea/msigdb/index.jsp). For the construction of the VirScan phage library, the complete proteome of most human-tropic viruses plus peptides derived from various microbial antigens and allergens annotated in the IEDB (https://www.iedb.org/) were utilized.
Raw scRNA-seq data (for Fig. 4i,j and Extended Data Fig. 6 (in the Supplementary Information)) and RNA-seq data of sorted DN T cells and monocytes (for Fig. 4e–g and Extended Data Fig. 9f) can be found at the NCBI Sequence Read Archive (accession no. PRJNA723618). Processed data can be found on Mendeley Data for scRNA-seq (https://doi.org/10.17632/nb26v3mx3x.2) and RNA-seq (https://doi.org/10.17632/nkhbn88v7g.2). Source data are provided with this paper. All other raw and processed data are available upon request from the corresponding authors under a Data Transfer Agreement.
Code availability
Custom codes for the analysis of scRNA-seq data and RNA-seq data are available on Mendeley Data (https://doi.org/10.17632/nb26v3mx3x.2 and https://doi.org/10.17632/nkhbn88v7g.2, respectively). All other custom codes are available upon request from the corresponding authors.
References
Houben, R. M. G. J. & Dodd, P. J. The global burden of latent tuberculosis infection: a re-estimation using mathematical modelling. PLoS Med. 13, 1–13 (2016).
Vynnycky, E. & Fine, P. E. M. Lifetime risks, incubation period and serial interval of tuberculosis. Am. J. Epidemiol. 152, 247–263 (2000).
Boisson-Dupuis, S. The monogenic basis of human tuberculosis. Hum. Genet. 139, 1001–1009 (2020).
Altare, F. et al. Interleukin-12 receptor β1 deficiency in a patient with abdominal tuberculosis. J. Infect. Dis. 184, 231–236 (2001).
Kreins, A. Y. et al. Human TYK2 deficiency: mycobacterial and viral infections without hyper-IgE syndrome. J. Exp. Med. 212, 1641–1662 (2015).
Boisson-Dupuis, S. et al. Tuberculosis and impaired IL-23-dependent IFN-γ immunity in humans homozygous for a common TYK2 missense variant. Sci. Immunol. 3, eaau8714 (2018).
Kerner, G. et al. Homozygosity for TYK2 P1104A underlies tuberculosis in about 1% of patients in a cohort of European ancestry. Proc. Natl Acad. Sci. USA 116, 10430–10434 (2019).
Kerner, G. et al. Human ancient DNA analyses reveal the high burden of tuberculosis in Europeans over the last 2,000 years. Am. J. Hum. Genet. 108, 517–524 (2021).
Toubiana, J. et al. Heterozygous STAT1 gain-of-function mutations underlie an unexpectedly broad clinical phenotype. Blood 127, 3154–3164 (2016).
Milner, J. D. et al. Early-onset lymphoproliferation and autoimmunity caused by germline STAT3 gain-of-function mutations. Blood 125, 591–599 (2015).
Haapaniemi, E. M. et al. Autoimmunity, hypogammaglobulinemia, lymphoproliferation and mycobacterial disease in patients with activating mutations in STAT3. Blood 125, 639–648 (2015).
Gonzalez-Mancera, M. S., Johnson, B. & Mirsaeidi, M. STAT3 gain-of-function mutation in a patient with pulmonary Mycobacterium abscessus infection. Respir. Med. Case Rep. https://doi.org/10.1016/j.rmcr.2020.101125 (2020).
Kalluri, R., Gattone, V. H., Noelken, M. E. & Hudson, B. G. The α3 chain of type IV collagen induces autoimmune Goodpasture syndrome. Proc. Natl Acad. Sci. USA 91, 6201–6205 (1994).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894 (2019).
Itan, Y. et al. The mutation significance cutoff: gene-level thresholds for variant predictions. Nat. Methods 13, 109–110 (2016).
Rapaport, F. et al. Negative selection on human genes underlying inborn errors depends on disease outcome and both the mode and mechanism of inheritance. Proc. Natl. Acad. Sci. USA 118, e2001248118 (2021).
Ishida, Y., Agata, Y., Shibahara, K. & Honjo, T. Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. EMBO J. 11, 3887–3895 (1992).
Wang, J. et al. PD-1 deficiency results in the development of fatal myocarditis in MRL mice. Int. Immunol. 22, 443–452 (2010).
Nishino, M., Sholl, L. M., Hatabu, H., Ramaiya, N. H. & Hodi, F. S. Anti–PD-1–related pneumonitis during cancer Immunotherapy. N. Engl. J. Med. 373, 288–290 (2015).
Zhang, B., Chikuma, S., Hori, S., Fagarasan, S. & Honjo, T. Nonoverlapping roles of PD-1 and FoxP3 in maintaining immune tolerance in a novel autoimmune pancreatitis mouse model. Proc. Natl Acad. Sci. USA 113, 8490–8495 (2016).
Picchi, H. et al. Infectious complications associated with the use of immune checkpoint inhibitors in oncology: reactivation of tuberculosis after anti PD-1 treatment. Clin. Microbiol. Infect. 24, 216–218 (2018).
Lázár-Molnár, E. et al. Programmed death-1 (PD-1)-deficient mice are extraordinarily sensitive to tuberculosis. Proc. Natl Acad. Sci. USA 107, 13402–13407 (2010).
Barber, D. L. et al. Tuberculosis following PD-1 blockade for cancer immunotherapy. Sci. Transl. Med. 11, eaat2702 (2019).
Wang, L., Das, H., Kamath, A. & Bukowski, J. F. Human Vγ2Vδ2 T cells produce IFN-γ and TNF-α with an on/off/on cycling pattern in response to live bacterial products. J. Immunol. 167, 6195–6201 (2001).
Le Bourhis, L. et al. Antimicrobial activity of mucosal-associated invariant T cells. Nat. Immunol. 11, 701–708 (2010).
Gold, M. C., Napier, R. J. & Lewinsohn, D. M. MR1-restricted mucosal associated invariant T (MAIT) cells in the immune response to Mycobacterium tuberculosis. Immunol. Rev. 264, 154–166 (2015).
Cooper, M. A. et al. Human natural killer cells: a unique innate immunoregulatory role for the CD56bright subset. Blood 97, 3146–3151 (2001).
Ndhlovu, Z. M. et al. Magnitude and kinetics of CD8+ T cell activation during hyperacute HIV infection impact viral set point. Immunity 43, 591–604 (2015).
Li, M. et al. T cell immunoglobulin and ITIM domain (TIGIT) receptor/poliovirus receptor (PVR) ligand engagement suppresses interferon-γ production of natural killer cells via β-arrestin 2-mediated negative signaling. J. Biol. Chem. 289, 17647–17657 (2014).
Jacquelot, N. et al. Sustained type I interferon signaling as a mechanism of resistance to PD-1 blockade. Cell Res. 29, 846–861 (2019).
Nguyen, K. B. et al. Interferon α/β-mediated inhibition and promotion of interferon γ: STAT1 resolves a paradox. Nat. Immunol. 1, 70–76 (2000).
Moreira-Teixeira, L. et al. Type I IFN exacerbates disease in tuberculosis-susceptible mice by inducing neutrophil-mediated lung inflammation and NETosis. Nat. Commun. 11, 5566 (2020).
Cantaert, T., Baeten, D., Tak, P. P. & van Baarsen, L. G. M. Type I IFN and TNFα cross-regulation in immune-mediated inflammatory disease: basic concepts and clinical relevance. Arthritis Res. Ther. 12, 219 (2010).
Flanagan, S. E. et al. Activating germline mutations in STAT3 cause early-onset multi-organ autoimmune disease. Nat. Genet. 46, 812–814 (2014).
Nabhani, S. et al. STAT3 gain-of-function mutations associated with autoimmune lymphoproliferative syndrome like disease deregulate lymphocyte apoptosis and can be targeted by BH3 mimetic compounds. Clin. Immunol. 181, 32–42 (2017).
Magerus-Chatinet, A. et al. FAS-L, IL-10, and double-negative CD4−CD8− TCR α/β+ T cells are reliable markers of autoimmune lymphoproliferative syndrome (ALPS) associated with FAS loss of function. Blood 113, 3027–3030 (2009).
Wei, S. C. et al. Negative co-stimulation constrains T cell differentiation by imposing boundaries on possible cell states. Immunity 50, 1084–1098 (2019).
Ma, C. S. et al. Deficiency of Th17 cells in hyper IgE syndrome due to mutations in STAT3. J. Exp. Med. 205, 1551–1557 (2008).
Okada, S. et al. Impairment of immunity to Candida and Mycobacterium in humans with bi-allelic RORC mutations. Science 349, 606–613 (2015).
Casanova, J.-L., Conley, M. E., Seligman, S. J., Abel, L. & Notarangelo, L. D. Guidelines for genetic studies in single patients: lessons from primary immunodeficiencies. J. Exp. Med. 211, 2137–2149 (2014).
Postow, M. A., Sidlow, R. & Hellmann, M. D. Immune-related adverse events associated with immune checkpoint blockade. N. Engl. J. Med. 378, 158–168 (2018).
Fabre, A. et al. Clinical aspects of STAT3 gain-of-function germline mutations: a systematic review. J. Allergy Clin. Immunol. Pract. 7, 1958–1969 (2019).
Forbes, L. R. et al. Jakinibs for the treatment of immune dysregulation in patients with gain-of-function signal transducer and activator of transcription 1 (STAT1) or STAT3 mutations. J. Allergy Clin. Immunol. 142, 1665–1669 (2018).
Li, H. & Tsokos, G. C. Double-negative T cells in autoimmune diseases. Curr. Opin. Rheumatol. 33, 163–172 (2021).
Crispín, J. C. et al. Expanded double-negative T cells in patients with systemic lupus erythematosus produce IL-17 and infiltrate the kidneys. J. Immunol. 181, 8761–8766 (2008).
Prokunina, L. et al. A regulatory polymorphism in PDCD1 is associated with susceptibility to systemic lupus erythematosus in humans. Nat. Genet. 32, 666–669 (2002).
Stroud, C. R. et al. Tocilizumab for the management of immune-mediated adverse events secondary to PD-1 blockade. J. Oncol. Pharm. Pract. 25, 551–557 (2019).
Martins, F. et al. New therapeutic perspectives to manage refractory immune checkpoint-related toxicities. Lancet Oncol. 20, e54–e64 (2019).
Kauffman, K. D. et al. PD-1 blockade exacerbates Mycobacterium tuberculosis infection in rhesus macaques. Sci. Immunol. 6, eabf3861 (2021).
Tousif, S. et al. T cells from programmed death-1-deficient mice respond poorly to Mycobacterium tuberculosis infection. PLoS ONE 6, e19864 (2011).
Nathan, C. F., Murray, H. W., Wiebe, M. E. & Rubin, B. Y. Identification of interferon-gamma as the lymphokine that activates human macrophage oxidative metabolism and antimicrobial activity. J. Exp. Med. 158, 670–689 (1983).
Jouanguy, E. et al. Partial interferon-gamma receptor 1 deficiency in a child with tuberculoid bacillus Calmette-Guérin infection and a sibling with clinical tuberculosis. J. Clin. Invest. 100, 2658–2664 (1997).
Flynn, J. L. et al. Tumor necrosis factor-alpha is required in the protective immune response against Mycobacterium tuberculosis in mice. Immunity 2, 561–572 (1995).
Harris, J. & Keane, J. How tumour necrosis factor blockers interfere with tuberculosis immunity. Clin. Exp. Immunol. 161, 1–9 (2010).
Schwartz, G. W. et al. TooManyCells identifies and visualizes relationships of single-cell clades. Nat. Methods 17, 405–413 (2020).
Xu, G. J. et al. Comprehensive serological profiling of human populations using a synthetic human virome. Science 348, aaa0698 (2015).
Kerner, G. et al. Inherited human IFN-γ deficiency underlies mycobacterial disease. J. Clin. Invest. 130, 3158–3171 (2020).
Mina, M. J. et al. Measles virus infection diminishes preexisting antibodies that offer protection from other pathogens. Science 366, 599–606 (2019).
Mohan, D. et al. PhIP–seq characterization of serum antibodies using oligonucleotide-encoded peptidomes. Nat. Protoc. 13, 1958–1978 (2018).
Belkadi, A. et al. Whole-exome sequencing to analyze population structure, parental inbreeding and familial linkage. Proc. Natl Acad. Sci. USA 113, 6713–6718 (2016).
Maffucci, P. et al. Blacklisting variants common in private cohorts but not in public databases optimizes human exome analysis. Proc. Natl Acad. Sci. USA 116, 950–959 (2019).
Gentili, M. et al. Transmission of innate immune signaling by packaging of cGAMP in viral particles. Science 349, 1232–1236 (2015).
Patsoukis, N. et al. Selective effects of PD-1 on Akt and Ras pathways regulate molecular components of the cell cycle and inhibit T cell proliferation. Sci. Signal. 5, ra46 (2012).
Yang, R. et al. Human T-bet governs innate and innate-like adaptive IFN-γ immunity against mycobacteria. Cell 183, 1826–1847 (2020).
Ma, C. S. et al. Unique and shared signaling pathways cooperate to regulate the differentiation of human CD4+ T cells into distinct effector subsets. J. Exp. Med. 213, 1589–1608 (2016).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Crowell, H. L. et al. muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data. Nat. Commun. 11, 6077 (2020).
Eisenberg, E. & Levanon, E. Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Tarazona, S., Garcia-Alcalde, F., Dopazo, J., Ferrer, A. & Conesa, A. Differential expression in RNA-seq: a matter of depth. Genome Res. 21, 2213–2223 (2011).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing. https://www.Rproject.org/ (2018).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Royal Stat. Soc. Ser. B 57, 289–300 (1995).
Acknowledgements
We thank the patients, their relatives and their physicians for participating in this study; D. Papandrea, Y. Nemirovskaya, M. Woollett, L. Lorenzo-Diaz and C. Patissier for administrative assistance; T. Kochetkov and J. Eun Han for technical assistance; and members of the laboratory for helpful discussions. We thank the Flow Cytometry Resource Center at the Rockefeller University and the Empire State Stem Cell Fund for providing support through NYSDOH contract no. C023046. We thank the Human Immune Monitoring Core at the Icahn School of Medicine at Mount Sinai for technical assistance with mass cytometry. We thank the NIH Tetramer Core Facility for providing the MR1 tetramer, which was developed jointly with J. McCluskey, J. Rossjohn and D. Fairlie. We thank Cycle for Survival, the Integrated Genomics Operation Core of the MSKCC, funded by the NIH/National Cancer Institute (NCI) Cancer Center Support Grant (CCSG, P30 CA008748), the Marie-Josée and Henry R. Kravis Center for Molecular Oncology, the Ludwig Center for Cancer Immunotherapy and the Parker Institute for Cancer Immunotherapy, for technical assistance with RNA-seq. The study was supported in part by grants from the St. Giles Foundation, The Rockefeller University, Institut National de la Santé et de la Recherche Médicale (INSERM), University of Paris, Sidra Medicine, the National Institute of Allergy and Infectious Diseases (R37AI095983 to J.-L.C., U19AI142737 to S.B.-D. and U19AI111143 and to M.S.G.), the National Institute of Diabetes and Digestive and Kidney Diseases (K08 DK113109 to S.V.), the National Center for Research Resources, the National Center for Advancing Sciences of the NIH (UL1TR001866), NIH/NCI Cancer Center Support Grant (CCSG, P30 CA008748), NIH/NCI R01 CA056821, the Swim Across America, the Ludwig Institute for Cancer Research, the Ludwig Center for Cancer Immunotherapy, the Cancer Research Institute, the Parker Institute for Cancer Immunotherapy, the French National Research Agency (ANR) under the ‘Investments for the Future’ program (ANR-10-IAHU-01), the Integrative Biology of Emerging Infectious Diseases Laboratory of Excellence (ANR-10-LABX-62-IBEID), GENMSMD (ANR-16-CE17.0005-01 to J.B.), the French Foundation for Medical Research (EQU201903007798), and the SCOR Corporate Foundation for Science. M.O. was supported by the David Rockefeller Graduate Program, the Funai Foundation for Information Technology, the Honjo International Scholarship Foundation, the New York Hideyo Noguchi Memorial Society and the Cooperative Center on Human Immunology at The Rockefeller University. R.Y. was supported by the Immune Deficiency Foundation and The Stony Wold-Herbert Fund. A.N.S. was supported by the European Commission (EC Horizon 2020 Marie Skłodowska-Curie Individual Fellowship no. 789645), the Dutch Research Council (NWO, Rubicon grant no. 019.171LW.015) and the European Molecular Biology Organization (Long-Term Fellowship no. ALTF 84-2017, non-stipendiary). J.R. was supported by the INSERM PhD program (poste d’accueil INSERM). S.C.W. was supported by a Medical Scientist Training Program grant from the National Institute of General Medical Sciences of the NIH to the Weill Cornell/Rockefeller/Sloan Kettering Tri-Institutional MD-PhD Program (T32GM007739). C.S.M. is supported by an Early-Mid Career Research Fellowship from the Department of Health of the New South Wales Government of Australia. S.G.T. is supported by an NHMRC Leadership 3 Investigator grant (1176665) and NHMRC program grant (1113904).
Author information
Authors and Affiliations
Contributions
M.O., S.B.-D. and J.-L.C. designed the study. M.O., R.Y., D.L., M.B., T.K., F.A.A., M.R., O.M.D., M.C., C.G., S.J.P., A.N.S., J.R., W.-T.L., S.D., G.R., C.S.M., Y.N., T.Y., K.C., S.C.W., J.-F.E., F. Rozenberg, G.A., M.S.G., D.B., N.M., L.D.N., S.G.T., T.H. and P.G. performed experiments. M.O. and P.Z. analyzed the single-cell and bulk RNA-seq data. P.Z., F. Rapaport, G.K. and L.A. assisted in the analysis of genetic data. M.D.H., M.K.C., M.A., P.W. and J.D.W. collected data on patients treated with anti-PD-1 monoclonal antibody. C.A., I.T., D.C., F.O.H., F.D., A.I., V.K.R., L.K., V.B. and J.B. collected clinical data and biological materials of patients. S.D. assisted with the experiments and the writing of the case report. M.O., S.B.-D. and J.-L.C. interpreted the data and wrote the manuscript with the help of all co-authors. All authors reviewed the manuscript and approved its submission. S.V., R.P.L., B.B., L.A., D.B., N.M., L.D.N., S.G.T., T.H. and P.G. are co-second-to-last authors contributing equally. S.B.-D. and J.-L.C. are co-last authors who jointly supervised the study.
Corresponding authors
Ethics declarations
Competing interests
M.S.G. has received consulting fees from Takeda, serves on the Scientific Advisory Board (SAB) of Vedanta Biosciences and receives consulting fees and equity and serves on the SAB of the Pandemic Response Lab of New York City. J.D.W. is a consultant for: Adaptive Biotech; Amgen; Apricity; Ascentage Pharma; Arsenal IO; Astellas; AstraZeneca; Bayer; Beigene; Boehringer Ingelheim; Bristol Myers Squibb; Celgene; Chugai; Daiichi Sankyo; Dragonfly; Eli Lilly; Elucida; F Star; Georgiamune; Idera; Imvaq; Kyowa Hakko Kirin; Linneaus; Maverick Therapeutics; Merck; Neon Therapeutics; Polynoma; Psioxus; Recepta; Takara Bio; Trieza; Truvax; Trishula; Sellas; Serametrix; Surface Oncology; Syndax; Syntalogic; and Werewolf Therapeutics. J.D.W. receives research support grants from: Bristol Myers Squibb and Sephora. J.D.W. has equity in: Tizona Pharmaceuticals; Adaptive Biotechnologies; Imvaq; Beigene; Linneaus; Apricity; Arsenal IO; and Georgiamune. M.D.H. reports grants from BMS; personal fees from Achilles, Arcus, AstraZeneca, Blueprint, BMS, Eli Lilly, Genentech/Roche, Genzyme/Sanofi, Janssen, Immunai, Instil Bio, Mana Therapeutics, Merck, Mirati, Natera, Pact Pharma, Shattuck Labs and Regeneron; and equity options from Factorial, Immunai, Shattuck Labs and Arcus. A patent filed by Memorial Sloan Kettering related to the use of tumor mutational burden to predict response to immunotherapy (PCT/US2015/062208) is pending and licensed by PGDx. S.D. became employed at Regeneron Pharmaceuticals after his contributions to this work. All other authors declare no competing interests.
Additional information
Peer review information Nature Medicine thanks Hakon Hakonarson, Weiping Zou, Caroline Robert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 3 Impaired IFN-γ and TNF production by PD-1-deficient leukocytes in response to mycobacterial and T-cell stimuli.
(a-c) Secreted cytokine levels in a whole-blood BCG assay. a, Secreted IFN-γ levels, as determined by ELISA after BCG stimulation of the whole-blood sample drawn from the patient two weeks before his death, while on immunosuppression therapy, including abatacept, for aggressive pulmonary autoimmunity. Samples from one control and the father (grouped as controls) were tested simultaneously. b, Secreted IL-12p40 levels, as determined by ELISA after BCG stimulation of the whole blood sample of the patient drawn after complete remission of abdominal TB. Samples from one control, the brother, and the parents (grouped as controls) were tested simultaneously. c, Secreted TNF levels, as determined with a LEGENDplex assay after BCG stimulation of the whole-blood sample from the patient drawn two weeks before his death, while on immunosuppression therapy. d and e, PBMC BCG assay. The patient’s PBMCs were obtained after complete remission of abdominal TB. Freshly thawed PBMCs from controls (N = 9, two BCG-naïve and seven BCG-vaccinated), the brother, the parents, and the patient, and one IL-12Rβ1-/- patient were dispended into a U-bottomed 96-well plate at a density of 2×105 cells/well and were stimulated. After 40 hours, a cytokine secretion inhibitor was added to the culture. After an additional eight hours of incubation, the supernatant and the cells were collected. d, Secreted cytokine levels, as determined with a LEGENDplex assay. e, Frequency of total IFN-γ+IL-10− and IFN-γ-IL-10+ cells, as determined by flow cytometry. f, PBMC stimulation assay. The patient’s cells were obtained one month after the initiation of anti-TB treatment. Secreted TNF levels were determined with a LEGENDplex assay after 24 hours of stimulation. Technical duplicates were prepared for the patient and his healthy brother and parents. Bars represent the mean and SEM. Statistical significance was determined via two-tailed non-paired Wilcoxon’s rank-sum tests with FDR adjustment for (e) the patient vs. local controls plus his brother and (f) the patient vs. local plus all family controls. P/I, PMA/ionomycin. (Extended Data Figs. 1, 2, 5 and 6 can be found in the Supplementary Information.).
Extended Data Fig. 4 Impaired IFN-γ production by splenocytes from PD-1-deficient mice repetitively stimulated in vitro.
a, Experimental design. Splenocytes were isolated from six PD-1-deficient mice and six age-matched wild-type (WT) controls and continuously cultured. b, Secreted IFN-γ levels, as determined by ELISA. For WT mice, N = 6 for all time points. For PD-1-deficient mice, N = 6 for days 4 and 8, and N = 5 for days 2 and 6. c, Absolute counts of IFN-γ-producing cells on day 8, as determined by flow cytometry. N = 6 and 5 for WT and PD-1-deficient mice, respectively. Bars represent the mean and SEM. Statistical significance was determined with a two-tailed Student’s t-test with FDR adjustment.
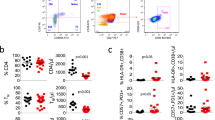
Extended Data Fig. 7 Expansion, activation, and proliferation of CD4−CD8− double-negative (DN) T cells in inherited and acquired PD-1 deficiency.
a and b, Effector DN αβ T cells. Previously published CyTOF data for peripheral lymph node-derived CD19−CD3ε+ T cells from adult PD-1-deficient and CTLA-4−deficient mice were downloaded from the FlowRepository (accessions FR-FCM-ZYFS and FR-FCM-ZYFQ, respectively)37. The control wild-type (WT) and heterozygous (Het) mice are the littermates of knockout (KO) mice. PD-1-deficient, N = 5, 2, and 7 for WT, Het, and KO, respectively. CTLA-4-deficient mice, N = 5, 3, and 7 for WT, Het, and KO, respectively. Bars represent the mean and SEM. Statistical significances were determined via two-tailed non-paired Wilcoxon’s rank-sum tests with FDR adjustment. c-f, Expansion and proliferation of DN T cells in patients on PD-1 blockade. A cohort of patients with advanced bladder cancer or melanoma treated with nivolumab monotherapy (N = 37) were immunophenotyped by flow cytometry. c and d, Total DN T cells. e and f, Ki67+ DN T cells. In (d and f), values before immunotherapy (Pre) and the highest values within 30 days after the initiation of immunotherapy (Post) were compared through two-tailed paired Wilcoxon’s rank-sum tests with FDR adjustment. The overlayed boxplot shows median (the horizontal bar), 25% and 75% quantiles (the lower and upper hinge), with the whiskers show the interquartile range multiplied by 1.5.
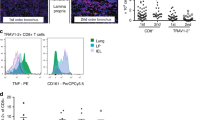
Extended Data Fig. 8 Cellular responses of PD-1-deficient CD4-CD8− double-negative (DN) T cells.
PBMCs from the patient (obtained one month after anti-TB treatment) and his healthy brother were stimulated for 24 hours with indicated stimuli, and DN T cells were analyzed by flow cytometry. Bars represent the mean and SEM.
Extended Data Fig. 9 Inherited PD-1 deficiency phenocopies STAT3 gain-of-function through excessive production of STAT3-activating cytokines by dysregulated T lymphocytes and myeloid cells.
a, Autoimmunity-associated gene expression levels in sorted CD4+ αβ T cells, as determined by RT-qPCR. To identify phenotypic similarities between the PD-1-deficient patient and other patients with monogenic forms of lymphoproliferative autoimmunity, PBMCs from the patient (obtained two weeks before his death), one patient with a heterozygous STAT3 gain-of-function (GOF) mutation, and two patients with FAS deficiency [that is, autoimmune lymphoproliferative syndrome (ALPS)] were studied. Selected autoimmunity-associated genes significantly upregulated in the patient’s effector T cells relative to his healthy brother’s effector T cells, as determined by the pseudobulk differential expression analysis of scRNASeq data, were tested by RT-qPCR with the patient’s PBMCs obtained at a different time point (that is, biological replicates). In addition, the gene expression levels were compared between cells with PD-1 deficiency, STAT3 GOF, and FAS deficiency to identify phenotypic similarities. b, Induction of STAT3 and STAT1 phosphorylation, as determined by flow cytometry. EBV-B cells were either left unstimulated or were stimulated with IL-6 (100 ng/mL) for 30 minutes. Technical triplicates were prepared only for the patient. c, Levels of IL-6 secreted by expanded PHA blasts, as determined in a LEGENDPlex assay. PHA blasts were either left unstimulated or were stimulated with anti-CD3/CD28 antibody-conjugated beads for 24 hours. Technical duplicates were prepared only for the patient. d and e, Cytokine levels in the bloodstream. Heparinized plasma samples from the patient, his father, and one local control were subjected to a custom LEGENDplex assay. d, STAT3-activating cytokines. e, Serological markers of FAS deficiency. sFAS, soluble FAS; sFASL, soluble FAS ligand. f, Aberrant expression of type I and type III IFN-encoding genes in the non-stimulated monocytes (CD3-CD19−CD56−CD14+/dim) sorted from PBMCs from the patient (obtained two weeks before his death), as determined by RNA sequencing. Log2-transformed size-factor-normalized read counts are shown. g, Immunophenotyping of myeloid cell subsets. PBMCs from the patient (obtained one month after anti-TB treatment) and his healthy brother were analyzed by flow cytometry. Bars represent the mean and SEM.
Extended Data Fig. 10 Activation-induced upregulation of RORγT in PD-1-deficient T lymphocytes.
a, c, and f, PBMCs from the patient (obtained one month after the initiation of anti-TB treatment) were studied. a, RORγT expression in αβ T cell subsets, as determined by flow cytometry. b, Expansion of RORγT-expressing DN αβ T cells in PD-1-deficient, but not in CTLA-4-deficient, mice. Previously published CyTOF data for peripheral lymph node-derived CD19−CD3ε+ T cells from adult PD-1-deficient and CTLA-4−deficient mice were downloaded from the FlowRepository (accessions FR-FCM-ZYFS and FR-FCM-ZYFQ, respectively)37. The control wild-type (WT) and heterozygous (Het) mice are the littermates of knockout (KO) mice. PD-1-deficient mice, N = 5, 2, and 7 for WT, Het, and KO, respectively. CTLA-4-deficient mice, N = 5, 3, and 7 for WT, Het, and KO, respectively. c, RORC mRNA levels in PBMCs stimulated with anti-CD3/CD28 antibody-conjugated Dynabeads for 24 hours, as determined by RT-qPCR with two different TaqMan probes. GUSB was used as an endogenous control. d, RORC mRNA levels in PHA-activated T cell blasts, as determined by RT-qPCR. Results from three consecutive experiments with technical duplicates are compiled. e, Induction of RORγT in HVS-T cells upon activation. HVS-T cells were incubated for 24 hours with mock Dynabeads, Dynabeads coated with anti-CD3 and anti-CD28 antibodies, Raji cells, or Raji cells plus blinatumomab (anti-CD3-CD19 bispecific engager). RORγT expression was measured by flow cytometry. f, IL-17A production. CD4+ αβ memory T cells were sorted from cryopreserved PBMCs and activated with beads conjugated with anti-CD2/CD3/CD28 antibodies. Supernatants were harvested on day 5, and the levels of secreted cytokines were determined in a cytometric bead array. Bars represent the mean and SEM. Statistical significances were determined via two-tailed non-paired Wilcoxon’s rank-sum tests with FDR adjustment.
Supplementary information
Supplementary Information
Supplementary Fig. 1, Supplementary Tables 1–4 and Extended Data Figs. 1, 2, 5 and 6.
Source data
Source Data Fig. 2
Unprocessed western blots for Fig. 2a.
Rights and permissions
About this article
Cite this article
Ogishi, M., Yang, R., Aytekin, C. et al. Inherited PD-1 deficiency underlies tuberculosis and autoimmunity in a child. Nat Med 27, 1646–1654 (2021). https://doi.org/10.1038/s41591-021-01388-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01388-5
This article is cited by
-
Clinical and translational attributes of immune-related adverse events
Nature Cancer (2024)
-
γδ T cells: origin and fate, subsets, diseases and immunotherapy
Signal Transduction and Targeted Therapy (2023)
-
Local γδ T cells: translating promise to practice in cancer immunotherapy
British Journal of Cancer (2023)
-
Transcriptional and post-transcriptional regulation of checkpoint genes on the tumour side of the immunological synapse
Heredity (2022)
-
Serial interferon-gamma release assay in lung cancer patients receiving immune checkpoint inhibitors: a prospective cohort study
Cancer Immunology, Immunotherapy (2022)