Abstract
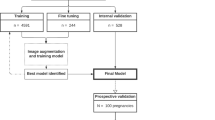
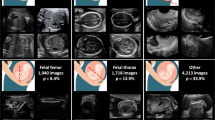
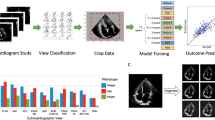
Congenital heart disease (CHD) is the most common birth defect. Fetal screening ultrasound provides five views of the heart that together can detect 90% of complex CHD, but in practice, sensitivity is as low as 30%. Here, using 107,823 images from 1,326 retrospective echocardiograms and screening ultrasounds from 18- to 24-week fetuses, we trained an ensemble of neural networks to identify recommended cardiac views and distinguish between normal hearts and complex CHD. We also used segmentation models to calculate standard fetal cardiothoracic measurements. In an internal test set of 4,108 fetal surveys (0.9% CHD, >4.4 million images), the model achieved an area under the curve (AUC) of 0.99, 95% sensitivity (95% confidence interval (CI), 84–99%), 96% specificity (95% CI, 95–97%) and 100% negative predictive value in distinguishing normal from abnormal hearts. Model sensitivity was comparable to that of clinicians and remained robust on outside-hospital and lower-quality images. The model’s decisions were based on clinically relevant features. Cardiac measurements correlated with reported measures for normal and abnormal hearts. Applied to guideline-recommended imaging, ensemble learning models could significantly improve detection of fetal CHD, a critical and global diagnostic challenge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Due to the sensitive nature of patient data, we are not able to make these data publicly available at this time. Source data are provided with this paper.
Code availability
ResNet and U-Net are publicly available and can be used with the settings described in the Methods and in Extended Data Fig. 1. The model weights that support this work are copyright of the Regents of the University of California and are available upon request. Additional code will be available upon publication at https://github.com/ArnaoutLabUCSF/cardioML.
References
Donofrio, M. T. et al. Diagnosis and treatment of fetal cardiac disease: a scientific statement from the American Heart Association. Circulation 129, 2183–2242 (2014).
Holland, B. J., Myers, J. A. & Woods, C. R. Jr. Prenatal diagnosis of critical congenital heart disease reduces risk of death from cardiovascular compromise prior to planned neonatal cardiac surgery: a meta-analysis. Ultrasound Obstet. Gynecol. 45, 631–638 (2015).
Wright, L. K. et al. Relation of prenatal diagnosis with one-year survival rate for infants with congenital heart disease. Am. J. Cardiol. 113, 1041–1044 (2014).
Bensemlali, M. et al. Neonatal management and outcomes of prenatally diagnosed CHDs. Cardiol. Young 27, 344–353 (2017).
Li, Y. F. et al. Efficacy of prenatal diagnosis of major congenital heart disease on perinatal management and perioperative mortality: a meta-analysis. World J. Pediatr. 12, 298–307 (2016).
Oster, M. E. et al. A population-based study of the association of prenatal diagnosis with survival rate for infants with congenital heart defects. Am. J. Cardiol. 113, 1036–1040 (2014).
Freud, L. R. et al. Fetal aortic valvuloplasty for evolving hypoplastic left heart syndrome: postnatal outcomes of the first 100 patients. Circulation 130, 638–645 (2014).
Sizarov, A. & Boudjemline, Y. Valve interventions in utero: understanding the timing, indications, and approaches. Can. J. Cardiol. 33, 1150–1158 (2017).
Committee on Practice, B.-O., the American Institute of Ultrasound in, M. Practice bulletin no. 175: ultrasound in pregnancy. Obstet. Gynecol. 128, e241–e256 (2016).
Tuncalp et al. WHO recommendations on antenatal care for a positive pregnancy experience—going beyond survival. BJOG 124, 860–862 (2017).
Carvalho, J. S. et al. ISUOG Practice Guidelines (updated): sonographic screening examination of the fetal heart. Ultrasound Obstet. Gynecol. 41, 348–359 (2013).
Bak, G. S. et al. Detection of fetal cardiac anomalies: is increasing the number of cardiac views cost-effective? Ultrasound Obstet. Gynecol. 55, 758–767 (2020).
Friedberg, M. K. et al. Prenatal detection of congenital heart disease. J. Pediatr. 155, 26–31 (2009).
Sekar, P. et al. Diagnosis of congenital heart disease in an era of universal prenatal ultrasound screening in southwest Ohio. Cardiol. Young 25, 35–41 (2015).
Sklansky, M. & DeVore, G. R. Fetal cardiac screening: what are we (and our guidelines) doing wrong? J. Ultrasound Med. 35, 679–681 (2016).
Sun, H. Y., Proudfoot, J. A. & McCandless, R. T. Prenatal detection of critical cardiac outflow tract anomalies remains suboptimal despite revised obstetrical imaging guidelines. Congenit. Heart Dis. 13, 748–756 (2018).
Corcoran, S. et al. Prenatal detection of major congenital heart disease—optimising resources to improve outcomes. Eur. J. Obstet. Gynecol. Reprod. Biol. 203, 260–263 (2016).
Letourneau, K. M. et al. Advancing prenatal detection of congenital heart disease: a novel screening protocol improves early diagnosis of complex congenital heart disease. J. Ultrasound Med. 37, 1073–1079 (2018).
AIUM practice parameter for the performance of fetal echocardiography. J. Ultrasound Med. 39, E5–E16 (2020).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Chilamkurthy, S. et al. Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study. Lancet 392, 2388–2396 (2018).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316, 2402–2410 (2016).
Baumgartner, C. F. et al. SonoNet: real-time detection and localisation of fetal standard scan planes in freehand ultrasound. IEEE Trans. Med. Imaging 36, 2204–2215 (2017).
Arnaout, R. Toward a clearer picture of health. Nat. Med. 25, 12 (2019).
Ouyang, D. et al. Video-based AI for beat-to-beat assessment of cardiac function. Nature 580, 252–256 (2020).
Madani, A., Arnaout, R., Mofrad, M. & Arnaout, R. Fast and accurate view classification of echocardiograms using deep learning. npj Digital Med. 1, 6 (2018).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. Preprint at https://arxiv.org/abs/1603.05027 (2016).
Lee, W. et al. AIUM practice guideline for the performance of fetal echocardiography. J. Ultrasound Med. 32, 1067–1082 (2013).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. Preprint at https://arxiv.org/abs/1610.02391 (2016).
Liu, H. et al. Fetal echocardiography for congenital heart disease diagnosis: a meta-analysis, power analysis and missing data analysis. Eur. J. Prev. Cardiol. 22, 1531–1547 (2015).
Pinheiro, D. O. et al. Accuracy of prenatal diagnosis of congenital cardiac malformations. Rev. Bras. Ginecol. Obstet. 41, 11–16 (2019).
Chu, C. et al. Prenatal diagnosis of congenital heart diseases by fetal echocardiography in second trimester: a Chinese multicenter study. Acta Obstet. Gynecol. Scand. 96, 454–463 (2017).
Zech, J. R. et al. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: a cross-sectional study. PLoS Med. 15, e1002683 (2018).
Miceli, F. A review of the diagnostic accuracy of fetal cardiac anomalies. Australas. J. Ultrasound Med. 18, 3–9 (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. Preprint at https://arxiv.org/abs/1505.04597 (2015).
Zhao, Y. et al. Fetal cardiac axis in tetralogy of Fallot: associations with prenatal findings, genetic anomalies and postnatal outcome. Ultrasound Obstet. Gynecol. 50, 58–62 (2017).
Goldinfeld, M. et al. Evaluation of fetal cardiac contractility by two-dimensional ultrasonography. Prenat. Diagn. 24, 799–803 (2004).
Best, K. E. & Rankin, J. Long-term survival of individuals born with congenital heart disease: a systematic review and meta-analysis. J. Am. Heart Assoc. 5, e002846 (2016).
Attia, Z. I. et al. An artificial intelligence-enabled ECG algorithm for the identification of patients with atrial fibrillation during sinus rhythm: a retrospective analysis of outcome prediction. Lancet 394, 861–867 (2019).
Peahl, A. F., Smith, R. D. & Moniz, M. H. Prenatal care redesign: creating flexible maternity care models through virtual care. Am. J. Obstet. Gynecol. https://doi.org/10.1016/j.ajog.2020.05.029 (2020).
Yeo, L., Markush, D. & Romero, R. Prenatal diagnosis of tetralogy of Fallot with pulmonary atresia using: Fetal Intelligent Navigation Echocardiography (FINE). J. Matern. Fetal Neonatal Med. 32, 3699–3702 (2019).
Cohen, L. et al. Three-dimensional fast acquisition with sonographically based volume computer-aided analysis for imaging of the fetal heart at 18 to 22 weeks’ gestation. J. Ultrasound Med. 29, 751–757 (2010).
World Health Organization. WHO Guideline: Recommendations on Digital Interventions for Health System Strengthening (2019).
Yagel, S., Cohen, S. M. & Achiron, R. Examination of the fetal heart by five short-axis views: a proposed screening method for comprehensive cardiac evaluation. Ultrasound Obstet. Gynecol. 17, 367–369 (2001).
Norgeot, B. et al. Minimum information about clinical artificial intelligence modeling: the MI-CLAIM checklist. Nat. Med. 26, 1320–1324 (2020).
Springenberg, J. T., Dosovitskiy, A., Brox, T. & Riedmiller, M. Striving for simplicity: the all convolutional net. Preprint at https://arxiv.org/abs/1412.6806 (2014).
Acknowledgements
We thank A. Butte and D. Srivastava for critical reading of the manuscript and M. Brook, M. Kohli, W. Tworetzky and K. Jenkins for facilitating data access. We thank all clinicians who served as human participants, including C. Springston, K. Kosiv, C. Tai and D. Abel; others wished to remain anonymous. The American Heart Association Precision Medicine Platform (https://precision.heart.org/) was used for data analysis. This project was also supported by the UCSF Academic Research Systems and the National Center for Advancing Translational Sciences, National Institutes of Health, through UCSF-CTSI grant UL1 TR991872. R.A., Y.Z., J.C.L., E.C. and A.J.M.-G. were supported by the National Institutes of Health (R01HL150394) and the American Heart Association (17IGMV33870001) and the Department of Defense (W81XWH-19-1-0294), all to R.A.
Author information
Authors and Affiliations
Contributions
R.A. and A.J.M.-G. conceived of the study. R.A. and E.C. designed and implemented all computational aspects of image processing, data labeling, pipeline design, neural network design, tuning and testing and data visualizations. R.A., L.C., Y.Z. and A.J.M.-G. labeled and validated images. J.C.L. curated and sent external data. R.A. wrote the manuscript with critical input from A.J.M.-G., E.C. and all authors.
Corresponding author
Ethics declarations
Competing interests
Some methods used in this work have been filed in a provisional patent application.
Additional information
Peer review information Nature Medicine thanks Zachi Attia, Declan O’Regan and Shaine Morris for their contribution to the peer review of this work. Editor recognition statement: Michael Basson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Neural network architectures and schematic of rules-based classifier.
a, Neural network architecture used for classification, based on ResNet (He et. al. 2015). Numbers indicate the number of filters in each layer, while the legend indicates the type of layer. For convolutional layers (grey), the size and stride of the convolutional filters is indicated in the legend. b, Neural network architecture used for segmentation, based on UNet (Ronneberger et. al. 2015). Numbers indicate the pixel dimensions at each layer. c, A schematic for the rules-based classifier (‘Composite dx classifier,’ Figure 1b) used to unite per-view, per-image predictions from neural network classifiers into a composite (per-heart) prediction of normal vs. CHD. Only views with AUC > 0.85 on validation data were used. For each view, there are various numbers of images k,l,m,n, each with a per-image prediction probability pCHD and pNL. For each view, per-image pCHD and pNL were summed and scaled (see Methods) into a pair of overall prediction values for each view (for example PCHD3VT and PNL3VT). These are in turn summed for a composite classification. Evaluating true positive, false positive, true negative, and false negative with different offset numbers allowed construction of an ROC curve for each test dataset (Figure 3e). 3VT, 3-vessel trachea. 3VV, 3-vessel view. LVOT, left ventricular outflow tract. A4C, axial 4-chamber.
Extended Data Fig. 2 Bland-Altman plots comparing cardiac measurements from labeled vs. predicted structures.
CTR, cardiothoracic ratio; CA, cardiac axis; LV, left ventricle; RV, right ventricle; LA, left atrium, RA, right atrium. Legend indicates measures for normal hearts (NL), hypoplastic left heart syndrome (HLHS), and tetralogy of Fallot (TOF).
Extended Data Fig. 3 Model confidence on sub-optimal images.
Examples of sub-optimal quality images (target views found by the model but deemed low-quality by human experts) are shown for each view, along with violin plots showing prediction probabilities assigned to the sub-optimal target images (White dots signify mean, thick black line signifies 1st to 3rd quartiles). Numbers in parentheses on top of violin plots indicate the number of independent images represented in each plot. For 3VT images, minimum, Q1, median, Q3, and maximum prediction probabilities are 0.27, 0.55, 0.74, 0.89, and 1.0, respectively. For 3VV images, minimum, Q1, median, Q3, and maximum prediction probabilities are 0.27, 0.73, 0.91, 0.99 and 1.0, respectively. For LVOT images, minimum, Q1, median, Q3, and maximum prediction probabilities are 0.31, 0.75, 0.92, 0.99, and 1.0, respectively. For A4C images, minimum, Q1, median, Q3, and maximum prediction probabilities are 0.28, 0.80, 0.95, 0.99, and 1.0, respectively. For ABDO images, minimum, Q1, median, Q3, and maximum prediction probabilities are 0.36, 0.83, 0.97, 1.0, and 1.0, respectively. Scale bars indicate 5mm. 3VT, 3-vessel trachea. 3VV, 3-vessel view. LVOT, left ventricular outflow tract. A4C, axial 4-chamber; ABDO, abdomen.
Extended Data Fig. 4 Misclassifications from per-view diagnostic classifiers.
Top row: Example images misclassified by the diagnostic classifiers, with probabilities for the predicted class. Relevant cardiac structures are labeled. Second row: corresponding saliency map. Third row: Grad-CAM. Fourth row: possible interpretation of model’s misclassifications. Importantly, this is only to provide some context for readers who are unfamiliar with fetal cardiac anatomy; formally, it is not possible to know the true reason behind model misclassification. Fifth row: Clinician’s classification (normal vs. CHD) on the isolated example image. Sixth row: Model’s composite prediction of normal vs. CHD using all available images for the given study. For several of these examples, the composite diagnosis per study is correct, even when a particular image-level classification was incorrect. Scale bars indicate 5 mm. 3VV, 3-vessel view. A4C, axial 4-chamber. SVC, superior vena cava. PA, pulmonary artery. RA, right atrium. RV, right ventricle. LA, left atrium. LV, left ventricle.
Extended Data Fig. 5 Inter-observer agreement on a subset of labeled data.
Inter-observer agreement on a sample of FETAL-125 is shown as Cohen’s Kappa statistic across different views, where poor agreement is 0–0.20; fair agreement is 0.21–0.40; moderate agreement is 0.41–0.60; good agreement is 0.61–0.80 and excellent agreement is 0.81–1.0. Of note, images where clinicians did not agree were not included in model training (see Methods). Most agreement is good or excellent, with moderate agreement on including 3VT and 3VV views as diagnostic-quality vs. non-target. 3VT, 3-vessel trachea. 3VV, 3-vessel view. LVOT, left ventricular outflow tract. A4C, axial 4-chamber, ABDO, abdomen, NT, non-target.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Arnaout, R., Curran, L., Zhao, Y. et al. An ensemble of neural networks provides expert-level prenatal detection of complex congenital heart disease. Nat Med 27, 882–891 (2021). https://doi.org/10.1038/s41591-021-01342-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01342-5
This article is cited by
-
Improving the diagnosis and treatment of congenital heart disease through the combination of three-dimensional echocardiography and image guided surgery
BMC Medical Imaging (2024)
-
Comprehensive evaluation and performance analysis of machine learning in heart disease prediction
Scientific Reports (2024)
-
AI supported fetal echocardiography with quality assessment
Scientific Reports (2024)
-
Machine Learning to Predict Outcomes of Fetal Cardiac Disease: A Pilot Study
Pediatric Cardiology (2024)
-
Artificial intelligence in cardiovascular diseases: diagnostic and therapeutic perspectives
European Journal of Medical Research (2023)