Abstract
To identify circulating proteins influencing Coronavirus Disease 2019 (COVID-19) susceptibility and severity, we undertook a two-sample Mendelian randomization (MR) study, rapidly scanning hundreds of circulating proteins while reducing bias due to reverse causation and confounding. In up to 14,134 cases and 1.2 million controls, we found that an s.d. increase in OAS1 levels was associated with reduced COVID-19 death or ventilation (odds ratio (OR) = 0.54, P = 7 × 10−8), hospitalization (OR = 0.61, P = 8 × 10−8) and susceptibility (OR = 0.78, P = 8 × 10−6). Measuring OAS1 levels in 504 individuals, we found that higher plasma OAS1 levels in a non-infectious state were associated with reduced COVID-19 susceptibility and severity. Further analyses suggested that a Neanderthal isoform of OAS1 in individuals of European ancestry affords this protection. Thus, evidence from MR and a case–control study support a protective role for OAS1 in COVID-19 adverse outcomes. Available pharmacological agents that increase OAS1 levels could be prioritized for drug development.
Similar content being viewed by others
Main
To date, the COVID-19 pandemic has caused more than 2 million deaths worldwide and infected approximately 100 million individuals1. Despite the scale of the epidemic, there are, at present, few disease-specific therapies2 to reduce the morbidity and mortality of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. Apart from dexamethasone therapy in oxygen-dependent patients3, most clinical trials have shown, at most, mild or inconsistent benefits on disease outcomes4,5,6. Therefore, validated targets are needed for COVID-19 therapeutic development.
One source of such targets is circulating proteins. Recent advances in large-scale proteomics have enabled the measurement of thousands of circulating proteins—and when combined with evidence from human genetics, such targets greatly improve the probability of drug development success7,8,9. Although de novo drug development will take time, the repurposing of currently available molecules targeting those proteins could provide an accelerated opportunity to deliver new therapies to patients.
Nevertheless, because confounding and reverse causation often bias traditional circulating protein studies, methods are needed to dissect causal relationships. This is especially the case in COVID-19, where exposure to SARS-CoV-2 unleashes profound changes in circulating protein levels10. One way to address these limitations is by using MR, a genetic epidemiology method that uses genetic variants as instrumental variables to test the effect of an exposure (here, protein levels) on an outcome (here, COVID-19 outcomes). The process of random assignment of alleles at conception greatly reduces bias from confounding. Because genotypes are always assigned before disease onset, MR studies are not influenced by reverse causation. However, MR rests on several assumptions11, the most problematic being horizontal pleiotropy of the genetic instruments (wherein the genotype influences the outcome, independently of the exposure). One way to help avoid this bias is to use genetic variants that influence circulating protein levels that are adjacent to the gene that encodes the circulating protein through the use of cis-protein quantitative trait loci (cis-pQTLs)9. cis-pQTLs are likely to influence the level of the circulating protein by directly influencing its transcription or translation and, therefore, less likely to affect the outcome of interest through pleiotropic pathways. Nevertheless, a causal genetic association between the exposure and outcome might be confounded by linkage disequilibrium (LD)12, which can be detected through co-localization testing.
Understanding the etiologic role of circulating proteins in infectious diseases is challenging because the infection itself often leads to large changes in circulating protein levels10. Thus, it might appear that an increase in a circulating protein, such as a cytokine, is associated with a worsened outcome, when, in fact, the cytokine might be the host’s response to this infection and help to mitigate this outcome. It is, therefore, important to identify genetic determinants of the protein levels in the non-infected state, which would reflect a person’s baseline predisposition to the level of a protein.
MR studies can be complemented by traditional case–control studies, where the protein is longitudinally measured in patients with COVID-19 and controls, allowing for an estimation of the association between the protein level and COVID-19 outcomes. However, MR studies tend to predict the effect of the protein in the non-infectious state when the genetic determinants of such proteins are measured in the non-infected population. Because MR and case–control studies rely on different assumptions and might be influenced by different biases, concordant results between the two study designs can strengthen the cumulative evidence13.
In this study, we, therefore, undertook two-sample MR and co-localization analyses to combine results from large-scale genome-wide association studies (GWASs) of circulating protein levels and COVID-19 outcomes14. We began by identifying the genetic determinants of circulating protein levels in large-scale proteomic GWASs and then used MR to assess whether these cis-pQTLs were associated with COVID-19 outcomes in large COVID-19 GWASs. Next, we investigated expression QTL (eQTL) and splice QTL (sQTL) effects of lead proteins. We then measured the most promising protein, OAS1, in individuals ascertained for SARS-CoV-2 infection, followed for longitudinal sampling during and after their infection.
Results
MR using cis-pQTLs and pleiotropy assessment
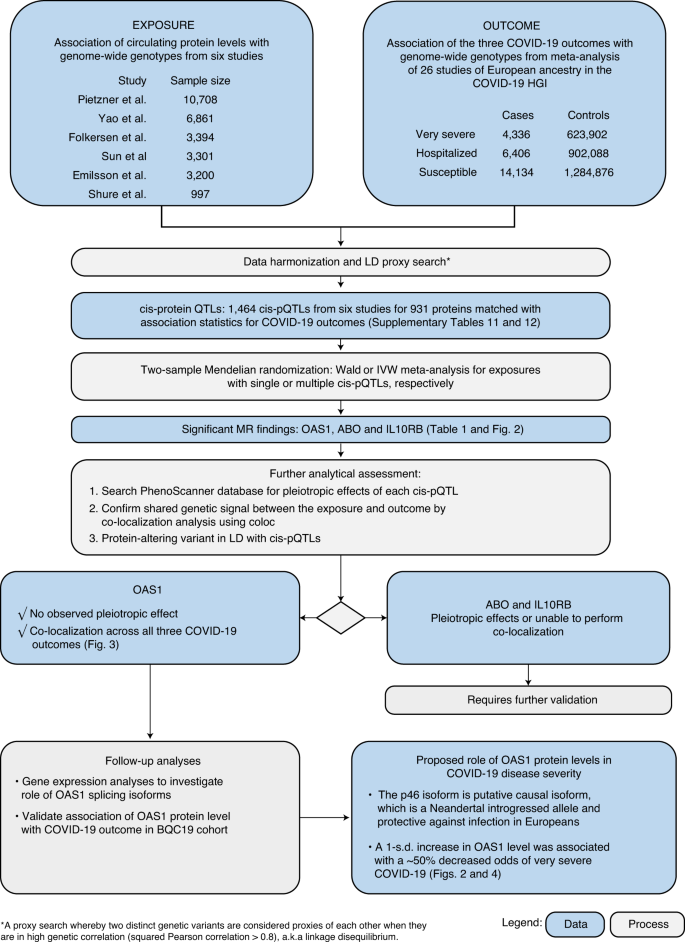
The study design is illustrated in Fig. 1. We began by obtaining the genetic determinants of circulating protein levels from six large proteomic GWASs of individuals of European ancestry (Sun et al.15 n = 3,301; Emilsson et al.16 n = 3,200; Pietzner et al.17 n = 10,708; Folkersen et al.18 n = 3,394; Yao et al.19 n = 6,861 and Suhre et al.20 n = 997). A total of 931 proteins from these six studies had genome-wide significant cis-pQTLs or highly correlated LD proxies (r2 > 0.8) in the meta-analyses of data from the COVID-19 Host Genetics Initiative21, which included results from the GenOMICC program22. We then undertook MR analyses using 1,425 cis-pQTLs and 39 LD proxies as genetic instruments for circulating proteins in three COVID-19 outcomes: 1) very severe COVID-19 disease (defined as individuals experiencing death, mechanical ventilation, non-invasive ventilation, high-flow oxygen or use of extra-corporeal membrane oxygenation; 99.7% of these individuals were of European ancestry) using 4,336 cases and 623,902 controls; 2) COVID-19 disease requiring hospitalization using 6,406 cases and 902,088 controls of European ancestry; and 3) COVID-19 susceptibility using 14,134 cases and 1,284,876 controls of European ancestry. In all outcomes, cases required evidence of SARS-CoV-2 infection. For the very severe COVID-19 and hospitalization outcomes, COVID-19 cases were defined as laboratory-confirmed SARS-CoV-2 infection based on nucleic acid amplification or serology tests. For the COVID-19 susceptibility outcome, cases were also identified by review of health records (using International Classification of Disease (ICD) codes or physician notes).
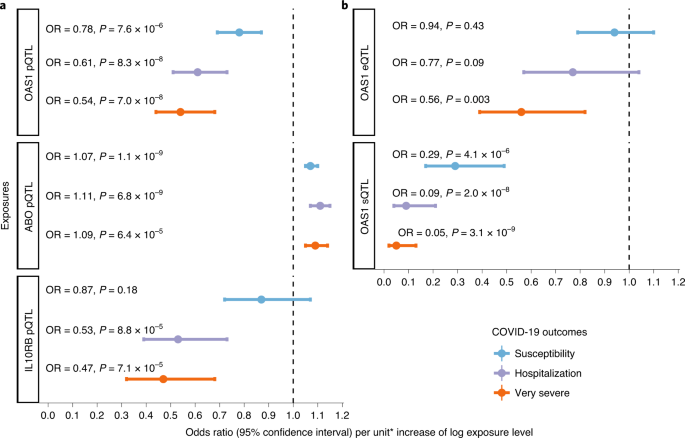
MR analyses revealed that the levels of three circulating proteins—2′–5′ oligoadenylate synthetase 1 (OAS1), interleukin-10 receptor beta subunit (IL10RB) and ABO—were associated with at least two COVID-19 outcomes after Benjamini–Hochberg false discovery rate correction (Table 1 and Supplementary Tables 1–6). Notably, increased OAS1 levels were strongly associated with protection from all three COVID-19 outcomes. Furthermore, these effect sizes were more pronounced with more severe outcomes, such that each s.d. increase in OAS1 levels was associated with decreased odds of very severe COVID-19 (OR = 0.54, 95% confidence interval (CI) 0.44–0.68, P = 7.0 × 10−8), hospitalization (OR = 0.61, 95% CI 0.51–0.73, P = 8.3 × 10−8) and susceptibility (OR = 0.78, 95% CI 0.69–0.87, P = 7.6 × 10−6) (Fig. 2a). We also identified OAS1 cis-pQTLs in Emilsson et al.16 and Pietzner et al.17, which were not included in the initial MR due to lack of genome-wide significance for their association with OAS1 levels16 or not included in their COVID-19 discovery panel17. MR analyses of using these additional cis-pQTLs yielded concordant results (Supplementary Table 7).
Forest plot showing OR and 95% CI from two sample MR analyses (two sided). P values are unadjusted. a, MR estimates of proteins influencing COVID-19 outcomes; unit: s.d. of log-normalized value. b, MR estimates of OAS1 messenger RNA influencing COVID-19 outcomes; unit: s.d. of normalized read counts.
We next assessed whether the cis-pQTL for OAS1 levels (rs4767027) was associated with over 5,000 other diseases, traits or protein levels, as catalogued in PhenoScanner23. rs4767027 was not associated with any other traits or protein levels (P < 5.0 × 10−5). These findings reduce the possibility that the MR estimate of the effect of OAS1 on COVID-19 outcomes is due to horizontal pleiotropy. Finally, except for COVID-19 susceptibility, the effect of rs4767027 did not demonstrate evidence of heterogeneity across COVID-19 Host Genetics Initiative GWAS meta-analyses (Table 1).
Using a cis-pQTL for IL10RB (rs2834167), we found that a 1-s.d. increase in circulating IL10RB level was associated with decreased odds of very severe COVID-19 (OR = 0.47, 95% CI 0.32–0.68, P = 7.1 × 10−5) and hospitalization (OR = 0.53, 95% CI 0.39–0.73, P = 8.8 × 10−5) but not susceptibility (Fig. 2a). Using PhenoScanner, we did not find evidence of pleiotropic effects of the cis-pQTL for IL10RB. A 1-s.d. increase in circulating ABO level was associated with increased odds of adverse COVID-19 outcomes (Table 1); however, we found that the cis-pQTL for ABO (rs505922) was strongly associated with the levels of several other proteins, suggesting potential horizontal pleiotropic effects (Supplementary Table 8). Given ABO’s known involvement in multiple physiological processes, these results were expected but highlight that MR analyses might suffer from significant bias from horizontal pleiotropy.
Co-localization studies
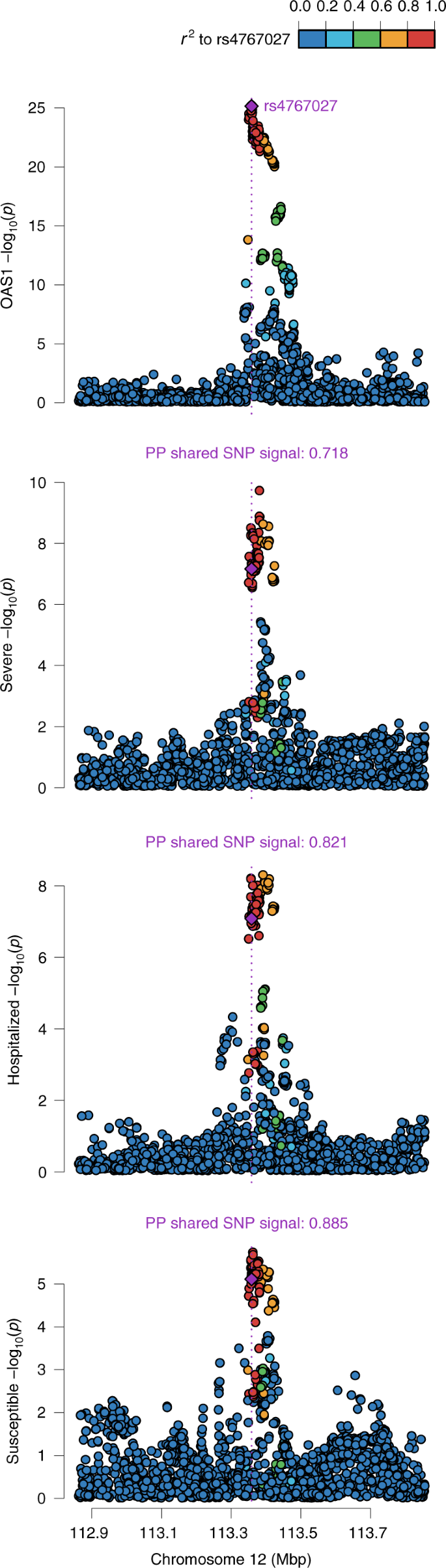
To test whether confounding due to LD might have influenced the estimated effect of circulating OAS1 on COVID-19 outcomes, we tested the probability that the genetic determinants of OAS1 circulating protein level were shared with the three COVID-19 outcomes using co-localization analyses, as implemented in coloc12. The posterior probability that OAS1 levels and COVID-19 outcomes shared a single causal signal in the 1-Mb locus around the cis-pQTL, rs4767027, was 0.72 for very severe COVID-19, 0.82 for hospitalization due to COVID-19 and 0.89 for COVID-19 susceptibility (Fig. 3). This co-localization result was also replicated using OAS1 cis-pQTL identified by Pietzner et al.17 (Supplementary Table 7). This suggests that there is likely a single shared causal signal for OAS1 circulating protein levels and COVID-19 outcomes.
Co-localization of genetic signal for OAS1 levels (top plot) and COVID-19 outcomes (three bottom plots) in the 1-Mb region around OAS1 pQTL rs4767027; color shows SNPs in the region in LD (r2) with rs4767027 (purple). The posterior probability (PP) of a shared single signal between OAS1 levels and the three COVID-19 outcomes was estimated by coloc.
Co-localization of ABO levels and different COVID-19 outcomes also showed co-localization between ABO level and different COVID-19 outcomes (posterior probability of single shared signal = 0.90, 0.98 and 1 for ABO level and very severe COVID-19, hospitalization due to COVID-19 and susceptibility, respectively) (Extended Data Fig. 1). We were unable to perform co-localization analyses for IL10RB due to a lack of genome-wide summary-level data from the original proteomic GWAS16.
Aptamer-binding effects
Protein-altering variants (PAVs)15 might influence binding of affinity agents, such as aptamers or antibodies, that are used to quantify protein levels. We, thus, assessed if the cis-pQTLs for the MR-prioritized proteins were PAVs or in LD (r2 > 0.8) with PAVs. rs2834167 (IL10RB) is a nonsense variant and could, therefore, be subject to potential binding effects. rs505922 (ABO) is not in LD with known missense variants. rs4767027 (OAS1) is an intronic variant, which is in LD with a missense variant rs2660 (r2 = 1) in European ancestry. However, because expression studies derived from RNA sequencing are not subject to potential effects of missense variants that could influence aptamer binding, we next explored whether rs4767027 also influences OAS1 expression and/or splicing.
sQTL and eQTL studies for OAS genes
sQTLs are genetic variants that influence the transcription of different isoforms of a protein. The aptamer that targets OAS1 was developed against a synthetic protein comprising the amino acid sequence 1–364 of NP002525.2, which is common to the two major OAS1 isoforms: p46 and p42. Hence, the aptamer might identify both or either isoforms. rs10774671 is a known sQTL for OAS1 that induces alternate splicing and creates p46 and p42 isoforms. Most present-day individuals of European ancestry carry the alternative variant (rs10774671-A). The ancestral variant (rs10774671-G) is the major allele in African populations and became fixed in Neanderthal and Denisovan genomes24,25. However, the ancestral variant, with its increased expression of the p46 isoform, was reintroduced into the European population via gene flow from Neanderthals26. Previous analyses suggest that individuals with either the GG or GA genotype at rs10774671 express higher amounts of p46 (ref. 26), which is also the predominant isoform found in circulating blood27. Differences in antiviral activity have been observed between isoforms, with p46 being more active in certain viral infections28. Interestingly, the OAS1 pQTL rs4767027 is in high LD (r2 = 0.97) with rs10774671 (ref. 26) in European populations. Functional studies support that the G allele at rs10774671 increases expression of the p46 isoform but decreases expression of the p42 isoform27. This G allele at the sQTL rs10774671 reflects the T allele at pQTL rs4767027, which itself is associated with higher measured OAS1 levels and reduced odds of COVID-19 severity and susceptibility. These separate lines of evidence suggest that OAS1 levels, as measured by the SomaScan platform, predominantly identify the p46 isoform, which might protect against COVID-19 outcomes.
Undertaking MR studies of OAS1 splicing, we found that increased expression of the p46 isoform (as defined by normalized read counts of the intron cluster defined by LeafCutter29,30) was associated with reduced odds of COVID-19 outcomes (OR = 0.29, 95% CI 0.17–0.49, P = 4.1 × 10−6 for susceptibility, OR = 0.09, 95% CI 0.04–0.21, P = 2.0 × 10−8 for hospitalization and OR = 0.05, 95% CI 0.02–0.13, P = 3.1 × 10−9 for very severe COVID-19) (Fig. 2b). Co-localization analyses also supported a shared causal signal among the sQTL for OAS1, the pQTL and COVID-19 outcomes (Extended Data Fig. 2). Interestingly, the co-localization analyses supported a stronger probability of a shared signal with the sQTL than the pQTL, suggesting that the p46 isoform might be the driver of the association of OAS1 levels with COVID-19 outcomes.
Next, we tested, using eQTL MR analyses, whether increased expression of OAS1 levels, without respect to isoform, was associated with COVID-19 outcomes. We identified an eQTL for total OAS1, rs10744785, from GTEx v8 (ref. 31). Total OAS1 expression levels were not associated with COVID-19 susceptibility and hospitalization (Fig. 2b). We also found that increased OAS3 expression in whole blood was positively associated with COVID-19 outcomes in MR analyses and support for co-localization of their genetic signal (Extended Data Fig. 3 nd Supplementary Table 9).
Taken together, these pQTL, sQTL and eQTL studies suggest that increased levels of the p46 isoform of OAS1 seem to protect against COVID-19 adverse outcomes.
Association of measured OAS1 protein level with COVID-19 outcomes
Because MR studies were derived from protein levels measured in a non-infected state, we tested the hypothesis that increased OAS1 protein levels in a non-infected state would be associated with reduced odds of COVID-19 outcomes. To do so, we undertook a case–control study, measuring OAS1 protein levels using the SomaScan platform in 1,039 longitudinal samples from 399 patients who tested positive for SARS-CoV-2 by polymerase chain reaction (PCR) that were collected at multiple time points during their COVID-19 infection and 105 individuals who presented with COVID-19 symptoms but had negative SARS-CoV-2 PCR nasal swabs from the Biobanque Quebecoise de la COVID-19 cohort (www.BQC19.ca). Individuals who had undergone nasal swabs for SARS-CoV-2 infection were recruited prospectively (Table 2).
We defined non-infectious samples as those collected from convalescent patients with SARS-CoV-2 at least 31 d after onset of their symptoms (n = 115) or samples collected from patients negative for SARS-CoV-2 by PCR (n = 105). We also measured OAS1 levels in individuals with samples from patients positive for SARS-CoV-2 <14 d after symptom onset (n = 313), which showed increased OAS1 levels during infection (Extended Data Figs. 4–6). OAS1 levels are not associated with age and sex in non-infectious samples (Extended Data Fig. 7). After sample quality control (Methods), 308 patients with at least one sample collected during infection, 113 patients with at least one sample collected during a non-infectious state and 103 COVID-19-negative controls were included in the analyses (Extended Data Fig. 8).
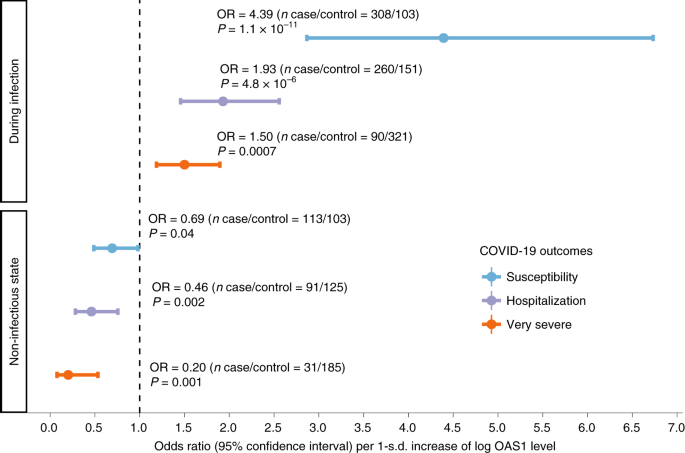
To test whether OAS1 levels in a non-infectious state were associated with COVID-19 outcomes, we undertook logistic regression controlling for age, sex, age*age, plate, recruitment center and sample processing time. OAS1 levels were log-transformed and standardized to match the transformation procedure of the MR study. We found that, in the non-infectious samples, each s.d. increase in OAS1 levels on the log-transformed scale was associated with reduced odds of COVID-19 outcomes (OR = 0.20, 95% CI 0.08–0.53, P = 0.001 for very severe COVID-19; OR = 0.46, 95% CI 0.28–0.76, P = 0.002 for hospitalization; and OR = 0.69, 95% CI 0.49–0.98, P = 0.04 for susceptibility) (Fig. 4, Extended Data Fig. 9 and Supplementary Table 10). These results are consistent with our findings from MR, where increased circulating OAS1 levels in a non-infectious state were associated with protection against all of these adverse COVID-19 outcomes.
Forest plot showing ORs and 95% CIs from logistic regression analyses (two sided). P values are unadjusted. During infection: patient samples that were collected within 14 d from the date of symptom onset. For individuals with two or more samples collected within 14 d of symptom onset, the earliest time point was used. Non-infectious state: patient samples that were collected at least 31 d from the date of symptom onset. For individuals with two or more samples collected at different time points at least 31 d from symptom onset, the latest time point was used. Additional information is also described in Supplementary Table 10.
In samples drawn during active infection, we found that increased OAS1 levels were associated with increased odds of adverse COVID-19 outcomes (OR = 1.50, 95% CI 1.19–1.90, P = 0.0007 for very severe COVID-19; OR = 1.93, 95% CI 1.46–2.56, P = 4.8 × 10−6 for hospitalization; and OR = 4.39, 95% CI 2.87–6.73, P = 1.09 × 10−11 for susceptibility) (Fig. 4).
Taken together, these findings suggest that increased OAS1 levels in a non-infectious state are associated with better COVID-19 outcomes, and that, during infection, SARS-CoV-2 exposure likely causes OAS1 levels to increase, as interferon pathways are stimulated, which are known to increase OAS1 levels32.
Discussion
Disease-specific therapies are needed to reduce the morbidity and mortality associated with COVID-19 outcomes. In this large-scale, two-sample MR study of 931 proteins assessed for three COVID-19 outcomes in up to 14,134 cases and 1.2 million controls of European ancestry, we provide evidence that increased OAS1 levels in the non-infectious state are strongly associated with reduced risks of very severe COVID-19, hospitalization and susceptibility. The protective effect size was particularly large, such that a 50% decrease in the odds of very severe COVID-19 was observed per s.d. increase in OAS1 circulating levels. OAS proteins are part of the innate immune response against RNA viruses. They are induced by interferons and activate latent RNase L, resulting in direct viral and endogenous RNA destruction, as demonstrated in in vitro studies33. Thus, OAS1 has a plausible biological activity against SARS-CoV-2. Because therapies exist that activate OAS1, repositioning them as potential COVID-19 treatments should be prioritized.
In populations outside of Sub-Saharan Africa, the protective alleles at both rs4767027-T (the OAS1 pQTL) and rs10774671-G (the OAS1 sQTL) are found on a Neanderthal haplotype34, which was passed on to modern humans ~50,000–60,000 years ago35. The correspondence between the previously described gene flow35 from Neandertals at this locus and the haplotype associated with protection against COVID-19 in the GWAS22 was recently demonstrated34. Even though these two single-nucleotide polymorphisms (SNPs) share a haplotype, their evolutionary histories differ. The rs4767027-T allele is derived from the Neanderthal lineage, whereas, for the rs10774671-G allele, Neanderthals preserved the ancestral state. OAS1 alternative splicing regulated by the rs10774671-G allele increases the isoform p46, which has a higher enzymatic activity against viruses than the p42 isoform36 and is the only OAS1 isoform robustly upregulated during infection26. Although further studies are needed to fully elucidate the functional relevance of the pQTL and sQTL for OAS1, the antiviral activity of the gene products is higher for the Neandertal haplotype than the common haplotype in Europeans28. In Europeans, the Neandertal haplotype has undergone positive selection26, and the rs4767027-T allele reaches an allele frequency of 0.32. Using MR and measurements of circulating proteins, we demonstrated here that increased OAS1 levels of the Neandertal haplotype in modern-day individuals of European ancestry confer this protective effect.
Our MR evidence indicated that higher p46 isoform levels of OAS1 and higher OAS1 total protein levels, as measured by the SomaScan assay, had protective effects on COVID-19 outcomes. These results were strongly supported by co-localization analysis. Given the consistent co-localization between the sQTL and pQTL for OAS1, the lack of co-localization between the eQTL and pQTL for OAS1 and the evidence that the SomaScan assay likely measures p46 isoforms, it seems probable that the protective effect of OAS1 is derived from the p46 isoform. However, further investigations are required to specifically measure each isoform in circulation, and isoform activity assays will be required to better understand if the p46 isoform, rather than total OAS1 levels, is most protective against COVID-19 outcomes.
The ancestral OAS1 splice variant encoding the more active p46 isoform was lost in the modern human population that left Africa. Several scenarios might explain this loss of function—for example, loss of purifying selection during the out-of-Africa exodus, which might be due to changes in environmental pathogens or potential harm induced by OAS1 antiviral activity37. Unfortunately, we do not have sufficient data to test if the OAS1 p46 ancestral allele in Sub-Saharan Africans also offers protection against COVID-19. Nevertheless, these findings further emphasize the importance of the Neanderthal genome in COVID-19 risk modulation, because a risk locus on chromosome 3 has also been reported to be inherited from Neanderthals38.
OAS1, OAS2 and OAS3 share considerable homology. As an interferon-stimulated gene39, OAS1 polymorphisms have been associated with the host immune response to several classes of viral infection40,41,42,43,44. Given that OAS1 is an intracellular enzyme-activating RNase L leading to viral RNA degradation, it is probable that the circulating levels of this enzyme reflect intracellular levels of this protein. However, there is experimental evidence that extracellular OAS1 might also be important in the viral immune response33.
Molecules currently exist that can influence OAS1 expression. Interferon beta-1b, which activates a cytokine cascade leading to increased OAS1 expression45, is currently used to treat multiple sclerosis and has been shown to induce OAS1 expression in blood cells46. Interferon-based therapy has also been used in other viral infections47. However, recent randomized trials have shown inconsistent results. Although intravenous interferon beta-1b combined with lopinavir–ritonavir reduced mortality due to MERS-CoV infections48, in the unblinded SOLIDARITY trial49 there was no demonstrated benefit of intravenous interferon-beta-1b. On the other hand, a recent phase 2 trial testing the effect of inhaled nebulized interferon beta-1a (which is closely related to interferon beta-1b) showed improved COVID-19 symptoms in the treatment arm50. Although this study was not powered to show a difference in mortality, all deaths occurred in the placebo group. Inhaled nebulized interferon beta results in a much higher tissue availability in the lung and might result in improved antiviral activity. Moreover, timing of administration is likely to play a role, as the administration of a pro-inflammatory cytokine might not provide benefit during the inflammation-driven phase of the disease. However, data on timing of administration are currently unavailable in the SOLIDARITY trial, and conclusions cannot yet be drawn. Lastly, the effect of interferon supplement might vary across ancestral populations, as different ancestries have different amounts of the more active p46 isoform of OAS1. Our study was limited to individuals of European ancestry, a population with higher expression of the p46 isoform. Interestingly, the SOLIDARITY trial enrolled 78% of its patients in South Asia, the Middle East, North Africa and Latin America, populations that might have higher expression of the p42 OAS1 isoform, whereas the study on inhaled interferon beta comprised 80% White patients from the United Kingdom. It is possible that interferon beta-1b might have different effects in populations of different ancestry due to different frequency of genetic variants in different populations.
There is in vitro evidence that pharmacological inhibition of phosphodiesterase-12, which degrades 2′–5′ oligoadenylate synthesized by OAS1, potentiates OAS-mediated antiviral activity51,52. Interestingly, coronaviruses in the same family as SARS-CoV-2 have been shown to produce viral proteins that degrade 2′–5′ oligoadenylate and reduce RNase-L activity, leading to evasion of the host immune response53,54. Our findings are also consistent with recent experimental work55 showing that there are situations where SARS-CoV-2 is sensitive to OAS1-related antiviral defenses. Our findings motivate pharmacologic strategies to increase OAS1 levels or activity, as well as further evaluation of the possible antiviral activity of extracellular OAS1 (ref. 33). Thus, existing preclinical molecules that lead to increased OAS1 levels51 could be optimized and tested for their effect on COVID-19 outcomes.
Our MR analyses found that higher levels of OAS3 expression is associated with worse COVID-19 outcomes, which is an opposite direction of effect compared to OAS1. The discordant effects of the p46 isoform for OAS1 and OAS3 were also reported by a previous study26, which might reflect complex biology of OAS genes for innate immune response. In a recent transcription-wide association study from the GenOMICC program22, genetically predicted high expression of OAS3 in lungs and whole blood was associated with a higher risk of patients with COVID-19 becoming critically ill. Although further studies to assess the roles of OAS genes specific to SARS-CoV-2 are needed, it is likely that OAS1 is the main driver of the protective effect of the p46 isoform for COVID-19 outcomes given previous functional studies demonstrating the antiviral effect of OAS genes26.
This study had limitations. First, we used MR to test the effect of circulating protein levels measured in a non-infected state because the effect of the cis-pQTLs on circulating proteins was estimated in individuals who had not been exposed to SARS-CoV-2. Once a person contracts SARS-CoV-2 infection, levels of circulating proteins could be altered, and this might be especially relevant for cytokines such as IL10 (which binds to IL10RB) and OAS1. Thus, the MR results presented in this paper should be interpreted as an estimation of the effect of circulating protein levels when measured in the non-infected state. Ongoing studies will help to clarify if the same cis-pQTLs influence circulating protein levels during infection. Second, this type of study suffers a high false-negative rate. Our goal was not to identify every circulating protein influencing COVID-19 outcomes but, rather, to provide evidence for a few proteins with strong cis-pQTLs, because these proteins are more likely to be robust to the assumptions of MR studies. Future large-scale proteomic studies with more circulating proteins properly assayed should help to overcome these limitations. Third, most MR studies assume a linear relationship between the exposure and the outcome. Thus, our findings would not identify proteins whose effect on COVID-19 outcomes has a clear threshold effect. Fourth, the overall OAS1 levels measured by RNA sequencing (not only p46) might be biased by the effect of alternative splicing, and the role of overall OAS1 and OAS3 levels indicated by the association of the cis-pQTL of OAS1 in protection against COVID-19 are possible and not yet explored. We also could not completely exclude the possibility that measurement of OAS1 levels might be influenced by aptamer-binding effects. Last, all data presented in this paper pertain to individuals of European ancestry only—once again underlining the importance of genotyping efforts in other populations.
In conclusion, we used genetic determinants of circulating protein levels and COVID-19 outcomes obtained from large-scale studies and found compelling evidence that OAS1 has a protective effect on COVID-19 susceptibility and severity. Measuring plasma OAS1 levels in a case–control study demonstrated that higher circulating levels of this protein in a non-infectious state are strongly associated with reduced risk of adverse COVID-19 outcomes. Interestingly, the available evidence suggests that the protective effect from OAS1 in individuals of European ancestry is likely due to the Neanderthal-introgressed p46 OAS1 isoform. Known pharmacological agents that increase OAS1 levels51 could be explored for their effect on COVID-19 outcomes.
Methods
pQTL GWAS
We systematically identified pQTL associations from six large proteomic GWASs15,16,17,18,19,20. Each of these studies undertook proteomic profiling using either SomaLogic SomaScans or O-link proximal extension assays.
COVID GWAS and COVID-19 outcomes
To assess the association of cis-pQTLs with COVID-19 outcomes, we used COVID-19 meta-analytic GWASs (data freeze 4) from the COVID-19 Host Genetics Initiative21. For our study, we used three of these GWAS meta-analyses, which included 25 cohorts of European ancestry and one cohort of admixed American ancestry. The outcomes tested were very severe COVID-19, hospitalization due to COVID-19 and susceptibility to COVID-19 (named A2, B2 and C2, respectively, by the COVID-19 Host Genetics Initiative).
Very severe COVID-19 cases were defined as hospitalized individuals with COVID-19 as the primary reason for hospital admission with laboratory-confirmed SARS-CoV-2 infection (nucleic acid amplification tests or serology based) and death or respiratory support (invasive ventilation, continuous positive airway pressure, bilevel positive airway pressure or continuous external negative pressure, high-flow nasal or face mask oxygen). Simple supplementary oxygen (for example, 2 L min−1 via nasal cannula) did not qualify for case status. Controls were all individuals in the participating cohorts who did not meet this case definition.
Hospitalized COVID-19 cases were defined as individuals hospitalized with laboratory-confirmed SARS-CoV-2 infection (using the same microbiology methods as for the very severe phenotype), where hospitalization was due to COVID-19-related symptoms. Controls were all individuals in the participating cohorts who did not meet this case definition.
Susceptibility to COVID-19 cases was defined as individuals with laboratory-confirmed SARS-CoV-2 infection, health record evidence of COVID-19 (ICD coding or physician confirmation) or with self-reported infections (for example, by questionnaire). Controls were all individuals who did not meet this case definition.
Two-sample MR
We used two-sample MR analyses to screen and test potential circulating proteins for their role in influencing COVID-19 outcomes. In two-sample MR, the effect of SNPs on the exposure and outcome are taken from separate GWASs. This method often improves statistical power because it allows for larger sample sizes for the exposure and outcome GWAS56.
Exposure definitions: We conducted MR using six large proteomic GWAS studies15,16,17,18,19,20. Circulating proteins from Sun et al., Emilsson et al. and Pietzner et al. were measured on the SomaLogic platform; Suhre et al., Yao et al. and Folkersen et al. used protein measurements on the O-link platform. We selected proteins with only cis-pQTLs to test their effects on COVID-19 outcomes because they are less likely to be affected by potential horizontal pleiotropy. The cis-pQTLs were defined as the genome-wide significant SNPs (P < 5 × 10−8) with the lowest P value within 1 Mb of the transcription start site of the gene encoding the measured protein9. For proteins from Emilsson et al., Pietzner et al., Suhre et al., Yao et al. and Folkersen et al., we used the sentinel cis-pQTL per protein per study as these were the data available. For proteins from Sun et al., we used PLINK 1.9 (ref. 57) and the 1000 Genome58 European population reference panels to clump and select LD-independent cis-pQTL (r2 < 0.001, distance 1,000 kb) with the lowest P value from reported summary statistics for each SOMAmer-bound protein. We included the same proteins represented by different cis-pQTLs from different studies to cross-examine the findings. For cis-pQTLs that were not present in the COVID-19 GWAS, SNPs with LD r2 > 0.8 and with minor allele frequency (MAF) < 0.42 were selected as proxies; MAF > 0.3 was used for allelic alignment for proxy SNPs. cis-pQTLs with palindromic effects and with MAF > 0.42 were removed before MR to prevent allele mismatches. Benjamini–Hochberg correction was used to control for the total number of proteins tested using MR. MR analyses were performed using the TwoSampleMR package in R59. For proteins with a single (sentinel) cis-pQTL, we used the Wald ratio to estimate the effect of each circulating protein on each of the three COVID-19 outcomes. For any proteins/SOMAmer reagents with multiple independent cis-pQTL, an inverse variance-weighted method was used to meta-analyze their combined effects. After harmonizing the cis-pQTLs of proteins with COVID-19 GWAS, a total of 566 SOMAmer reagents (529 proteins, 565 directly matched cis-pQTL and 26 proxies) from Sun et al., 760 proteins (747 directly matched cis-pQTL and 11 proxies) from Emilsson et al., 91 proteins (90 directly matched cis-pQTLs and two proxies) from Pietzner et al., 74 proteins (72 directly matched cis-pQTL) from Suhre et al., 24 proteins (24 directly matched cis-pQTLs) from Yao et al. and 13 proteins (13 directly matched cis-pQTLs) from Folkersen et al. were used as instruments for the MR analyses across the three COVID-19 outcomes (Supplementary Tables 11 and 12)15,16,17,18,19,20.
Pleiotropy assessments
A common pitfall of MR is horizontal pleiotropy, which occurs when the genetic variant affects the outcome via pathways independent of the exposure. The use of circulating protein cis-pQTLs greatly reduces the possibility of pleiotropy, for reasons described above. We also searched in the PhenoScanner23 database, a large catalog of observed SNP–outcome relationships involving >5,000 GWASs done to date to assess potentially pleiotropic effects of the cis-pQTLs of MR-prioritized proteins by testing the association of cis-pQTLs with other circulating proteins (that is, if they were trans-pQTLs to other proteins or significantly associated with other unrelated diseases or traits). For cis-pQTLs of MR-prioritized proteins measured on the SomaLogic platform, we assessed the possibility of potential aptamer-binding effects (where the presence of PAVs might affect protein measurements). We also checked if cis-pQTLs of MR-prioritized proteins had significantly heterogeneous associations across COVID-19 populations in each COVID-19 outcome GWAS.
Co-localization analysis
Next, we tested co-localization of the genetic signal for the circulating protein and each of the three COVID-19 outcomes using co-localization analyses, which assess potential confounding by LD. Specifically, for each of these MR-significant proteins with genome-wide summary data available, for the proteomic GWASs a stringent Bayesian analysis was implemented in coloc12 R package to analyze all variants in the 1-Mb genomic locus centered on the cis-pQTL. Co-localizations with posterior probability for hypothesis 4 (PP4, that there is an association for both protein level and COVID-19 outcomes, and they are driven by the same causal variant) > 0.5 were considered likely to co-localize (which means the highest posterior probability for all five coloc hypotheses), and PP4 > 0.8 was considered to be highly likely to co-localize.
sQTL and eQTL MR and co-localization studies for OAS genes
We performed MR and co-localization analysis using GTEx project v8 (ref. 31) GWAS summary data to understand the effects of expression and alternative splicing of OAS genes in whole blood. The genetic instruments were conditionally independent (r2 < 0.001) sQTLs and eQTLs for OAS1 and eQTLs for OAS2 and OAS3 identified by using stepwise regression in GTEx31. The sQTL SNP for OAS1 (rs10774671) was originally identified for the normalized read counts of LeafCutter29 cluster of the last intron of the p46 isoform (chr12:112,917,700–112,919,389, GRCh38) in GTEx30 and was used to estimate the effect of the p46 isoform. Co-localization analysis was performed using GWAS summary statistics from GTEx by restricting to the regions within 1 Mb of each QTL.
Measurement of plasma OAS1 protein levels associated with COVID-19 outcomes in BQC19
BQC19 is a Québec-wide initiative to enable research into the causes and consequences of COVID-19 disease. The patients included in this study were recruited at the Jewish General Hospital (JGH) and the Centre Hospitalier de l’Université de Montréal (CHUM) in Montréal, Québec, Canada.
COVID-19 case–control status was defined to be consistent with the GWAS study from the COVID-19 Host Genetics Initiative, from which the MR results were derived. Namely, we tested the association of OAS1 protein levels with the three different COVID-19 outcome definitions both in samples procured from non-infected stages and samples procured during the acute phase of the infection. The three outcomes were as follows. 1) Very severe COVID-19—defined as hospitalized individuals with laboratory-confirmed SARS-CoV-2 infection (nucleic acid amplification tests or serology based) and death or respiratory support (invasive ventilation, continuous positive airway pressure, bilevel positive airway pressure or continuous external negative pressure, high-flow nasal or face mask oxygen). Controls were all individuals who did not meet this case definition. 2) Hospitalized COVID-19 cases—defined as individuals hospitalized with laboratory-confirmed SARS-CoV-2 infection. Controls were all individuals who did not meet this case definition. 3) Susceptibility to COVID-19—cases were defined as individuals with laboratory-confirmed SARS-CoV-2 infection, and controls were all individuals who underwent PCR testing for SARS-CoV-2 but were negative. The date of symptom onset for patients with COVID-19 was collected from patient charts or estimated from their first positive COVID-19 tests if missing. Case inclusion criteria were not exclusive, which means that some individuals who were cases in the susceptibility analyses were also included in the hospitalization and very severe COVID-19 cohorts if they met case definitions.
A total of 125 individuals were recruited from CHUM, and 379 individuals were recruited from the JGH. Individuals had blood sampling done at up to five different time points (200 individuals had one measurement, 113 individuals had two measurements, 152 individuals had three measurements, 38 individuals had four measurements and one individual had five measurements). Days from symptom onset (T1) were calculated for each sample based on the date of symptom onset and blood draw date. For individuals who were negative for COVID-19, T1 was set to 0. Sample processing time (in hours) for each sample was also calculated to measure the duration of time from sample collection to processing to account for the increase in the amount of protein released from cell lysis due to extended sample handling time.
Protein levels in citrated (ACD) plasma samples were measured using the SomaScan assay. In total, 1,039 samples from 399 patients who were positive for SARS-CoV-2 and 105 patients who were negative for SARS-CoV-2 of mainly European descent underwent SomaScan assays, which included 5,284 SOMAmer reagents targeting 4,742 proteins. The SomaScan assay uses single-stranded DNA aptamers (‘SOMAmers’), which are designed to selectively bind to a particular protein target60. SOMAmer reagent binding is quantified by microarray, measuring abundance in relative fluorescent units (RFUs). The RFUs for each protein underwent four normalization processes, including hybridization control, intraplate median signal normalization, plate scaling and calibration and median signal normalization to a reference generated from internal data across all samples. All normalizations were conducted by SomaLogic and detailed in their Technical Note61.
Of particpants who were positive for SARS-CoV-2, we defined samples procured from patients during the infectious state as those sampled within 14 d (including the 14th day) from the first date of symptoms62. For patients with more than one sample within 14 d of symptom onset, the earliest sample was used. We defined samples procured from patients who were non-infectious as samples from SARS-CoV-2-positive patients taken at least 31 d after symptom onset. We selected 31 d, as this is the upper limit of the interquartile range of the duration of SARS-CoV-2 positivity in a recent systematic review and coincided with the first scheduled outpatient follow-up blood test in the BQC19 (ref. 63). For individuals with more than one sample at least 31 d after symptom onset, the latest sample was used.
OAS1 level was measured by one SOMAmer reagent (OAS1.10361.25). Within each group, median signal-normalized OAS1 levels were natural log-transformed and adjusted for sample processing time, and the residuals were further standardized. For each group, we removed samples that were outliers with long sample processing time (sample processing time > 50 h) or high OAS1 level (log OAS1 level > 8). Logistic regression was performed to test the association-standardized OAS1 level with the three COVID-19 outcomes including age, sex, age*age, center of recruitment and plates as covariates.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data from proteomics studies and GTEx consortium (GTEx project v8 (ref. 31)) are available from the referenced peer-reviewed studies15,16,17,18,19,20 or their corresponding authors, as applicable. The PhenoScanner online database is available at http://www.phenoscanner.medschl.cam.ac.uk/. Summary statistics for the COVID-19 outcomes are publicly available for download on the COVID-19 Host Genetics Initiative website (www.covid19hg.org). The BQC19 is an Open Science biobank. Instructions on how to access data for individuals from the BQC19 at the Jewish General Hospital site are available here: https://www.mcgill.ca/genepi/mcg-covid-19-biobank. Instructions on how to access data from other sites of the BQC19 are available here: https://www.bqc19.ca/en/access-data-samples.
References
Johns Hopkins University of Medicine. Coronavirus Resource Center. https://coronavirus.jhu.edu/ (2020).
Weinreich, D. M. et al. REGN-COV2, a neutralizing antibody cocktail, in outpatients with Covid-19. N. Engl. J. Med. 384, 238–251 (2020).
Horby, P. et al. Dexamethasone in hospitalized patients with Covid-19—preliminary report. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2021436 (2020).
Sterne, J. A. C. et al. Association between administration of systemic corticosteroids and mortality among critically ill patients with COVID-19: a meta-analysis. JAMA 324, 1330–1341 (2020).
Beigel, J. H., Tomashek, K. M. & Dodd, L. E. Remdesivir for the treatment of Covid-19—preliminary report. N. Engl. J. Med. 383, 994 (2020).
Cavalcanti, A. B. et al. Hydroxychloroquine with or without azithromycin in mild-to-moderate Covid-19. N. Engl. J. Med. 383, 2041–2052 (2020).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Cook, D. et al. Lessons learned from the fate of AstraZeneca’s drug pipeline: a five-dimensional framework. Nat. Rev. Drug Discov. 13, 419–431 (2014).
Zheng, J. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat. Genet. 52, 1122–1131 (2020).
Filbin, M. R. et al. Plasma proteomics reveals tissue-specific cell death and mediators of cell–cell interactions in severe COVID-19 patients. Preprint at bioRxiv https://doi.org/10.1101/2020.11.02.365536 (2020).
Davey Smith, G., Ebrahim, S., Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Lawlor, D. A., Tilling, K. & Smith, G. D. Triangulation in aetiological epidemiology. Int. J. Epidemiol. 45, 1866–1886 (2016).
COVID-19 Host Genetics Initiative. The COVID-19 Host Genetics Initiative, a global initiative to elucidate the role of host genetic factors in susceptibility and severity of the SARS-CoV-2 virus pandemic. Eur. J. Hum. Genet. 28, 715–718 (2020).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Emilsson, V. et al. Co-regulatory networks of human serum proteins link genetics to disease. Science 361, 769–773 (2018).
Pietzner, M. et al. Genetic architecture of host proteins involved in SARS-CoV-2 infection. Nat. Commun. 11, 6397 (2020).
Folkersen, L. et al. Mapping of 79 loci for 83 plasma protein biomarkers in cardiovascular disease. PLoS Genet. 13, e1006706 (2017).
Yao, C. et al. Genome-wide mapping of plasma protein QTLs identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
COVID-19 Host Genetics Initiative. https://www.covid19hg.org/results/ (2021).
Pairo-Castineira, E. et al. Genetic mechanisms of critical illness in Covid-19. Nature https://doi.org/10.1038/s41586-020-03065-y (2020).
Staley, J. R. et al. PhenoScanner: a database of human genotype–phenotype associations. Bioinformatics 32, 3207–3209 (2016).
Prüfer, K. et al. A high-coverage Neandertal genome from Vindija Cave in Croatia. Science 358, 655–658 (2017).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Sams, A. J. et al. Adaptively introgressed Neandertal haplotype at the OAS locus functionally impacts innate immune responses in humans. Genome Biol. 17, 246 (2016).
Li, H. et al. Identification of a Sjögren’s syndrome susceptibility locus at OAS1 that influences isoform switching, protein expression, and responsiveness to type I interferons. PLoS Genet. 13, e1006820 (2017).
Liu, X. et al. A functional variant in the OAS1 gene is associated with Sjögren’s syndrome complicated with HBV infection. Sci. Rep. 7, 17571 (2017).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using LeafCutter. Nat. Genet. 50, 151–158 (2018).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Hornung, V., Hartmann, R., Ablasser, A. & Hopfner, K. P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat. Rev. Immunol. 14, 521–528 (2014).
Kristiansen, H. et al. Extracellular 2′–5′ oligoadenylate synthetase stimulates RNase L-independent antiviral activity: a novel mechanism of virus-induced innate immunity. J. Virol. 84, 11898–11904 (2010).
Zeberg, H. & Pääbo, S. A genomic region associated with protection against severe COVID-19 is inherited from Neandertals. Proc. Natl Acad. Sci. USA 118, e2026309118 (2021).
Mendez, F. L., Watkins, J. C. & Hammer, M. F. Neandertal origin of genetic variation at the cluster of OAS immunity genes. Mol. Biol. Evol. 30, 798–801 (2013).
Bonnevie-Nielsen, V. et al. Variation in antiviral 2′–5′-oligoadenylate synthetase (2′5′AS) enzyme activity is controlled by a single-nucleotide polymorphism at a splice-acceptor site in the OAS1 gene. Am. J. Hum. Genet. 76, 623–633 (2005).
Carey, C. M. et al. Recurrent loss-of-function mutations reveal costs to OAS1 antiviral activity in primates. Cell Host Microbe 25, 336–343 (2019).
Zeberg, H. & Pääbo, S. The major genetic risk factor for severe COVID-19 is inherited from Neanderthals. Nature 587, 610–612 (2020).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Min, J.-Y. & Krug, R. M. The primary function of RNA binding by the influenza A virus NS1 protein in infected cells: inhibiting the 2′–5′ oligo (A) synthetase/RNase L pathway. Proc. Natl Acad. Sci. USA 103, 7100–7105 (2006).
Hu, B. et al. Cellular responses to HSV-1 infection are linked to specific types of alterations in the host transcriptome. Sci. Rep. 6, 28075 (2016).
Lim, J. K. et al. Genetic variation in OAS1 is a risk factor for initial infection with West Nile virus in man. PLoS Pathog. 5, e1000321 (2009).
Simon-Loriere, E. et al. High anti-dengue virus activity of the OAS gene family is associated with increased severity of dengue. J. Infect. Dis. 212, 2011–2020 (2015).
Hamano, E. et al. Polymorphisms of interferon-inducible genes OAS-1 and MxA associated with SARS in the Vietnamese population. Biochem. Biophys. Res. Commun. 329, 1234–1239 (2005).
Cheng, G. et al. Pharmacologic activation of the innate immune system to prevent respiratory viral infections. Am. J. Respir. Cell Mol. Biol. 45, 480–488 (2011).
Harari, D., Orr, I., Rotkopf, R., Baranzini, S. E. & Schreiber, G. A robust type I interferon gene signature from blood RNA defines quantitative but not qualitative differences between three major IFNβ drugs in the treatment of multiple sclerosis. Hum. Mol. Genet. 24, 3192–3205 (2014).
Lin, F. & Young, H. A. Interferons: success in anti-viral immunotherapy. Cytokine Growth Factor Rev. 25, 369–376 (2014).
Arabi, Y. M. et al. Interferon beta-1b and lopinavir–ritonavir for Middle East respiratory syndrome. N. Engl. J. Med. 383, 1645–1656 (2020).
WHO Solidarity Trial Consortium. Repurposed antiviral drugs for COVID-19—interim WHO SOLIDARITY trial results. N. Engl. J. Med. 384, 497–511 (2021).
Monk, P. D. et al. Safety and efficacy of inhaled nebulised interferon beta-1a (SNG001) for treatment of SARS-CoV-2 infection: a randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Respir. Med. 9, 196–206 (2020).
Wood, E. R. et al. The role of phosphodiesterase 12 (PDE12) as a negative regulator of the innate immune response and the discovery of antiviral inhibitors. J. Biol. Chem. 290, 19681–19696 (2015).
Silverman, R. H. & Weiss, S. R. Viral phosphodiesterases that antagonize double-stranded RNA signaling to RNase L by degrading 2-5A. J. Interferon Cytokine Res. 34, 455–463 (2014).
Zhao, L. et al. Antagonism of the interferon-induced OAS-RNase L pathway by murine coronavirus ns2 protein is required for virus replication and liver pathology. Cell Host Microbe 11, 607–616 (2012).
Zhang, R. et al. Homologous 2′,5′-phosphodiesterases from disparate RNA viruses antagonize antiviral innate immunity. Proc. Natl Acad. Sci. USA 110, 13114–13119 (2013).
Li, Y. et al. SARS-CoV-2 induces double-stranded RNA-mediated innate immune responses in respiratory epithelial derived cells and cardiomyocytes. Preprint at bioRxiv https://doi.org/10.1101/2020.09.24.312553 (2020).
Burgess, S., Davies, N. M. & Thompson, S. G. Bias due to participant overlap in two-sample Mendelian randomization. Genet. Epidemiol. 40, 597–608 (2016).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Candia, J. et al. Assessment of variability in the SOMAscan assay. Sci. Rep. 7, 14248 (2017).
SomaLogic. Short Technical Note. https://somalogic.com/wp-content/uploads/2019/07/Short-Technical-Note-SOMAmer-specificity.pdf (2019).
Centers for Disease Control and Prevention. Duration of isolation and precautions for adults with COVID-19. https://www.cdc.gov/coronavirus/2019-ncov/hcp/duration-isolation.html (2021).
Cevik, M. et al. SARS-CoV-2, SARS-CoV, and MERS-CoV viral load dynamics, duration of viral shedding, and infectiousness: a systematic review and meta-analysis. Lancet Microbe 2, e13–e22 (2020).
Acknowledgements
We thank N. Chomont for the discussion on the clinical phenotype of BQC19. We would like to also thank the MI4 and the MUHC Foundation that contributed for the SomaLogic panel. The Richards research group is supported by the Canadian Institutes of Health Research (CIHR) (365825 and 409511), the Lady Davis Institute of the Jewish General Hospital, the Canadian Foundation for Innovation (CFI), the NIH Foundation, Cancer Research UK, Genome Québec, the Public Health Agency of Canada, the McGill Interdisciplinary Initiative in Infection and Immunity and the Fonds de Recherche Québec Santé (FRQS). S.Z. is supported by a CIHR fellowship and a FRQS postdoctoral scholarship. G.B.L. is supported by a CIHR scholarship and a joint FRQS and Québec Ministry of Health and Social Services scholarship. T.N. is supported by research fellowships of the Japan Society for the Promotion of Science for Young Scientists. J.B.R. is supported by an FRQS Clinical Research Scholarship. Support from Calcul Québec and Compute Canada is acknowledged. TwinsUK is funded by the Welcome Trust, the Medical Research Council, the European Union, the National Institute for Health Research-funded BioResource and the Clinical Research Facility and Biomedical Research Centre based at Guy’s and St. Thomas’ NHS Foundation Trust in partnership with King’s College London. BQC19 is funded by FRQS, Genome Québec and the Public Health Agency of Canada. These funding agencies had no role in the design, implementation or interpretation of this study. V.M. is supported by a Canada Excellence Research Chair. M.D. is supported by an FRQS Clinical Research Scholarship. R.F. is funded by the Swedish Research Council (2014-02569 and 2014-07606). The Kaufmann lab’s COVID-19 work is supported by the CIHR/CITF (VR2-173203), AmFAR (110068-68-RGCV), the CFI and FRQS. M.H. is supported by a SciLifeLab/KAW national COVID-19 research program project grant (KAW 2020.0182).
Author information
Authors and Affiliations
Contributions
Conception and design: S.Z., G.B.L. and J.B.R. Data analyses: S.Z. and T.N. Data acquisition: T.N., G.B.L., D.M., D.E.K., J.A., M.A., L.L., E.B.R., D.H., N.K., Z.A., N.R., M.B., L.P., C.G., X.X., C.T., B.V., O.A., T.A., N.A., M.C., M.D., V.F., D.E.K. and J.B.R. Interpretation of data: S.Z., G.B.L., T.N., M.P., Y.C., D.E.K., V.F. and J.B.R. Funding acquisition: D.M., V.M., V.F. and J.B.R. Methodology: S.Z., K.Z., C.M.T.G. and J.B.R. Project administration: D.M., V.F. and J.B.R. Validation: S.Z., T.N., M.P., N.K., M.P., J.N., E.T., C.L., D.E.K. and J.B.R. Visualization: S.Z., T.N. and V.F. Writing—original draft: S.Z., G.B.L., T.N. and J.B.R. Writing—review and editing: S.Z., G.B.L., T.N., M.P., H.Z., V.M., M.P., R.F., M.L., M.H., C.P., D.E.K. and J.B.R. All authors were involved in further drafts of the manuscript and revised it critically for content. All authors gave final approval of the version to be published. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Corresponding author
Ethics declarations
Ethics declarations
All institutions contributing cohorts to the COVID-19 Host Genetics Initiative received ethics approval from their respective research ethics review boards. This study was approved by the research ethics boards of the Jewish General Hospital and the Centre Hospitalier du Université de Montréal. Informed consent was obtained from the participants of BQC19.
Competing interests
J.B.R. has served as an advisor to GlaxoSmithKline and Deerfield Capital. C.P. is an employee of SomaLogic. M.P. has consulted for Astellas Pharma, Ansh Labs, Immunomet, Zymeworks and Pfizer on unrelated projects. All other authors declare that there are no conflicts of interest.
Additional information
Peer review information Nature Medicine thanks Luis Barriero, Andre Franke, and Adam Butterworth for their contribution to the peer review of this work. Joao Monteiro was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Colocalization of the Genetic Determinants of ABO Plasma Protein Levels and COVID-19 Outcomes.
Colored circles represent the LD of SNPs within 1 Mb region around the leading cis-pQTL of ABO rs505922. PP: posterior probability of sharing the same genetic signal with ABO level, estimated by coloc.
Extended Data Fig. 2 Colocalization of the expression QTL (eQTL), splicing QTL (sQTL) and protein QTL (pQTL) of OAS1 with COVID-19 outcomes.
a, R2 was calculated using the rs4767027 SNP in each COVID-19 GWAS for 503 European individuals in the 1000 Genomes Project. b, A table illustrating the pairwise colocalization results between OAS1 expression (eQTL), splicing (sQTL), protein (pQTL) and COVID-19 outcomes. Colocalization analyses were performed using GWAS summary from GTEx by restricting to the regions within 1 Mb of rs4767027.
Extended Data Fig. 3 Colocalization of the expression QTL (eQTL) of OAS3 and COVID-19 outcomes.
R2 was calculated using the SNP with the lowest p-value in each COVID-19 GWAS for 503 European individuals in 1000 Genomes Project. Colocalization analysis was performed using GWAS summary from GTEx by restricting to the region within 1 Mb of rs4767027.
Extended Data Fig. 4 OAS1 Level Trajectory with Days Since Symptom Onset in different COVID-19 Outcomes.
Trajectory plot showing OAS1 level (in normalized RFUs) of 38 COVID-19 patients with their blood samples collected at four different time points, represented by days since symptom onset. Each line represents one patient. Blue shows COVID-19 patient without very severe outcomes and red shows COVID-19 patient with very severe outcome.
Extended Data Fig. 5 Correlation of log OAS1 level with Processing Time in Samples from a Non-infectious State and During Infection.
Pearson correlation (two-sided) of sample processing time and log OAS1 level of the two groups (During infection and Non-infectious state) after QC. The regression line was added for each of the two groups separately, with shaded parts showing the 95% confidence interval.
Extended Data Fig. 6 Distribution of log transformed of OAS1 in samples during non-infectious state and during infection.
Figure includes samples in each group before QC. X-axis is the natural log transformed OAS1 level. Showing 10 Quantiles of the log OAS1 level of each group.
Extended Data Fig. 7 OAS1 level association with Age and Sex in samples during non-infectious state.
Pearson correlation (two-sided) of age and log OAS1 level after the removal of outliers of OAS1 level and extended sample processing time, stratified by sex. The regression line was added for males and females separately, with shaded parts showing the 95% confidence interval.
Extended Data Fig. 8 Distribution of Days from Symptom Onset and Sample Processing Time in BQC-19 Samples.
a, Distribution of Days from Symptom Onset in all 934 samples from COVID-19 patients with SomaScan® measurement. b, Distribution of Sample Processing Time in all 1039 samples with SomaScan® measurement. c, Distribution of Days from Symptom Onset in 421 samples from COVID-19 patients included in the analyses after QC. d, Distribution of Sample Processing Time in 524 samples included in the analyses after QC.
Extended Data Fig. 9 Violin plot of OAS1 Level Measured by SomaScan® Technology in BQC19 cohort in Groups of Different COVID-19 Outcomes, in samples taken from a Non-infectious State and During Infection.
A violin plot with box plot showing the distribution and median value of natural log transformed OAS1 level (Y-axis) in each group (During infection and Non-infectious state) with each outcome (Very severe COVID-19, Hospitalization Due to COVID-19 and COVID-19 Susceptibility). The lower and upper hinges of the boxplot correspond to the first and third quartiles. The lower and upper whiskers of the boxplot extend from the hinge to the smallest and largest value at most 1.5 * IQR of the hinge. The sample size included in each group can be found in Fig. 4 and Supplementary Table 10. Controls of each group include COVID negative individuals.
Supplementary information
Supplementary Tables
Supplementary Tables 1–12.
Rights and permissions
About this article
Cite this article
Zhou, S., Butler-Laporte, G., Nakanishi, T. et al. A Neanderthal OAS1 isoform protects individuals of European ancestry against COVID-19 susceptibility and severity. Nat Med 27, 659–667 (2021). https://doi.org/10.1038/s41591-021-01281-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01281-1
This article is cited by
-
Polymorphisms of IFN signaling genes and FOXP4 influence the severity of COVID-19
BMC Infectious Diseases (2024)
-
Mendelian randomization and colocalization analysis reveal novel drug targets for myasthenia gravis
Human Genomics (2024)
-
Role of the Neanderthal Genome in Genetic Susceptibility to COVID-19: 3p21.31 Locus in the Spotlight
Biochemical Genetics (2024)
-
Time-sensitive testing pressures and COVID-19 outcomes: are socioeconomic inequalities over the first year of the pandemic explained by selection bias?
BMC Public Health (2023)
-
Integration of risk variants from GWAS with SARS-CoV-2 RNA interactome prioritizes FUBP1 and RAB2A as risk genes for COVID-19
Scientific Reports (2023)