Abstract
Immune and targeted therapies achieve long-term survival in metastatic melanoma; however, new treatment strategies are needed to improve patients’ outcomes1,2. We report on the efficacy, safety and biomarker analysis from the single-arm safety run-in (part 1; n = 9) and biomarker (part 2; n = 27) cohorts of the randomized, placebo-controlled, phase 3 COMBI-i trial (NCT02967692) of the anti-PD-1 antibody spartalizumab, in combination with the BRAF inhibitor dabrafenib and MEK inhibitor trametinib. Patients (n = 36) had previously untreated BRAF V600-mutant unresectable or metastatic melanoma. In part 1, the recommended phase 3 regimen was identified based on the incidence of dose-limiting toxicities (DLTs; primary endpoint): 400 mg of spartalizumab every 4 weeks plus 150 mg of dabrafenib twice daily plus 2 mg of trametinib once daily. Part 2 characterized changes in PD-L1 levels and CD8+ cells following treatment (primary endpoint), and analyzed additional biomarkers. Assessments of efficacy and safety were key secondary endpoints (median follow-up, 24.3 months). Spartalizumab plus dabrafenib and trametinib led to an objective response rate (ORR) of 78%, including 44% complete responses (CRs). Grade ≥3 treatment-related adverse events (TRAEs) were experienced by 72% of patients. All patients had temporary dose modifications, and 17% permanently discontinued all three study drugs due to TRAEs. Early progression-free survival (PFS) events were associated with low tumor mutational burden/T cell–inflamed gene expression signature (GES) or high immunosuppressive tumor microenvironment (TME) GES levels at baseline; an immunosuppressive TME may also preclude CR. Overall, the efficacy, safety and on-treatment biomarker modulations associated with spartalizumab plus dabrafenib and trametinib are promising, and biomarkers that may predict long-term benefit were identified.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Novartis is committed to sharing, with qualified external researchers, access to patient-level data and supporting clinical documents from eligible studies. Requests are reviewed and approved by an independent review panel on the basis of scientific merit. All data provided are anonymized to respect the privacy of patients who have participated in the trial, in line with applicable laws and regulations. This trial data availability is in accordance with the criteria and process described on ClinicalStudyDataRequest.com. Publicly available databases utilized for the biomarker analyses in this study include RefSeq (https://www.ncbi.nlm.nih.gov/refseq/), dbSNP (https://www.ncbi.nlm.nih.gov/snp/), MSigDB C2 Canonical Pathways (https://www.gsea-msigdb.org/gsea/msigdb), the Exome Sequencing Project (http://evs.gs.washington.edu/EVS/), the Exome Aggregation Consortium (now part of gnomAD, https://gnomad.broadinstitute.org/) and COSMIC (https://cancer.sanger.ac.uk/cosmic).
References
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma (KEYNOTE-006): post-hoc 5-year results from an open-label, multicentre, randomised, controlled, phase 3 study. Lancet Oncol. 20, 1239–1251 (2019).
Robert, C. et al. Five-year outcomes with dabrafenib plus trametinib in metastatic melanoma. N. Engl. J. Med. 381, 626–636 (2019).
Larkin, J. et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 381, 1535–1546 (2019).
McArthur, G. et al. 5-Year survival update of cobimetinib plus vemurafenib in BRAF V600 mutation-positive advanced melanoma: final analysis of the coBRIM study. Pigment Cell Melanoma Res. 33, 148–255 (2020).
Ugurel, S. et al. Survival of patients with advanced metastatic melanoma: the impact of MAP kinase pathway inhibition and immune checkpoint inhibition – update 2019. Eur. J. Cancer 130, 126–138 (2020).
Ugurel, S. et al. Survival of patients with advanced metastatic melanoma: the impact of novel therapies – update 2017. Eur. J. Cancer 83, 247–257 (2017).
Hu-Lieskovan, S. et al. Improved antitumor activity of immunotherapy with BRAF and MEK inhibitors in BRAF(V600E) melanoma. Sci. Transl. Med. 7, 279ra241 (2015).
Kakavand, H. et al. PD-L1 expression and tumor-infiltrating lymphocytes define different subsets of MAPK inhibitor-treated melanoma patients. Clin. Cancer Res. 21, 3140–3148 (2015).
Wilmott, J. S. et al. Selective BRAF inhibitors induce marked T-cell infiltration into human metastatic melanoma. Clin. Cancer Res. 18, 1386–1394 (2012).
Wilmott, J. S. et al. Dynamics of chemokine, cytokine, and growth factor serum levels in BRAF-mutant melanoma patients during BRAF inhibitor treatment. J. Immunol. 192, 2505–2513 (2014).
Ascierto, P. A. & Dummer, R. Immunological effects of BRAF + MEK inhibition. Oncoimmunology 7, e1468955 (2018).
Mandalà, M. et al. Immunomodulating property of MAPK inhibitors: from translational knowledge to clinical implementation. Lab. Invest. 97, 166–175 (2017).
Yu, C. et al. Combination of immunotherapy with targeted therapy: theory and practice in metastatic melanoma. Front. Immunol. 10, 990 (2019).
Kelderman, S., Schumacher, T. N. & Haanen, J. B. Acquired and intrinsic resistance in cancer immunotherapy. Mol. Oncol. 8, 1132–1139 (2014).
Ascierto, P. A. et al. Update on tolerability and overall survival in COLUMBUS: landmark analysis of a randomised phase 3 trial of encorafenib plus binimetinib vs vemurafenib or encorafenib in patients with BRAF V600-mutant melanoma. Eur. J. Cancer 126, 33–44 (2020).
Ascierto, P. A. et al. Dabrafenib, trametinib and pembrolizumab or placebo in BRAF-mutant melanoma. Nat. Med. 25, 941–946 (2019).
Ribas, A. et al. Combined BRAF and MEK inhibition with PD-1 blockade immunotherapy in BRAF-mutant melanoma. Nat. Med. 25, 936–940 (2019).
Sullivan, R. J. et al. Atezolizumab plus cobimetinib and vemurafenib in BRAF-mutated melanoma patients. Nat. Med. 25, 929–935 (2019).
Ferrucci, P. F. et al. Updated survival in patients with BRAF-mutant melanoma administered pemobrlizumab, dabrafenib, and trametinib. Pigment Cell Melanoma Res. 33, 148–255 (2020).
McArthur, G. A. et al. Evaluation of atezolizumab (A), cobimetinib (C), and vemurafenib (V) in previously untreated patients with BRAFV600 mutation-positive advanced melanoma: primary results from the phase 3 IMspire150 trial. In American Association for Cancer Research Virtual Annual Meeting I, abstract CT012 (2020).
Lin, C.-C. et al. Phase I/II study of spartalizumab (PDR001), an anti-PD1 mAb, in patients with advanced melanoma or non-small cell lung cancer. Ann. Oncol. 29, VIII413 (2018).
Larkin, J. et al. Overall survival in patients with advanced melanoma who received nivolumab versus investigator’s choice chemotherapy in CheckMate 037: a randomized, controlled, open-label phase III trial. J. Clin. Oncol. 36, 383–390 (2018).
Hamid, O. et al. Final analysis of a randomised trial comparing pembrolizumab versus investigator-choice chemotherapy for ipilimumab-refractory advanced melanoma. Eur. J. Cancer 86, 37–45 (2017).
Ribas, A. et al. Pembrolizumab versus investigator-choice chemotherapy for ipilimumab-refractory melanoma (KEYNOTE-002): a randomised, controlled, phase 2 trial. Lancet Oncol. 16, 908–918 (2015).
Wagle, M. C. et al. A transcriptional MAPK Pathway Activity Score (MPAS) is a clinically relevant biomarker in multiple cancer types. NPJ Precis. Oncol. 2, 7 (2018).
Long, G. V. et al. Characteristics of long-term survivors and subgroup analyses with combination nivolumab plus ipilimumab for advanced melanoma (CheckMate 067). Pigment Cell Melanoma Res. 33, 148–255 (2020).
Osgood, C. et al. FDA analysis of depth of response (DpR) and survival across 10 randomized controlled trials in patients with previously untreated unresectable metastatic melanoma (UMM) by therapy type. J. Clin. Oncol. 37, 9508 (2019).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Hodi, F. S. et al. Genomic analyses and immunotherapy in advanced melanoma. Cancer Res. 79, CT037 (2019).
Ribas, A. et al. Tumor mutational burden (TMB), T cell-inflamed gene expression profile (GEP) and PD-L1 are independently associated with response to pembrolizumab (pembro) in patients with advanced melanoma in the KEYNOTE (KN)-006 study. Cancer Res. 79, 4217 (2019).
Weber, J. S. et al. Adjuvant nivolumab (NIVO) versus ipilimumab (IPI) in resected stage III/IV melanoma: 3-year efficacy and biomarker results from the phase III CheckMate 238 trial. Ann. Oncol. 30, V533–V534 (2019).
Dummer, R. et al. Adjuvant dabrafenib plus trametinib versus placebo in patients with resected, BRAF(V600)-mutant, stage III melanoma (COMBI-AD): exploratory biomarker analyses from a randomised, phase 3 trial. Lancet Oncol. 21, 358–372 (2020).
Gide, T. N. et al. Distinct immune cell populations define response to anti-PD-1 monotherapy and anti-PD-1/anti-CTLA-4 combined therapy. Cancer Cell 35, 238–255 (2019).
Vilain, R. E. et al. Dynamic changes in PD-L1 expression and immune infiltrates early during treatment predict response to PD-1 blockade in melanoma. Clin. Cancer Res. 23, 5024–5033 (2017).
Massi, D. et al. The density and spatial tissue distribution of CD8(+) and CD163(+) immune cells predict response and outcome in melanoma patients receiving MAPK inhibitors. J. Immunother. Cancer 7, 308 (2019).
Lee, J. H. et al. Circulating tumour DNA predicts response to anti-PD1 antibodies in metastatic melanoma. Ann. Oncol. 28, 1130–1136 (2017).
Santiago-Walker, A. et al. Correlation of BRAF mutation status in circulating-free DNA and tumor and association with clinical outcome across four BRAFi and MEKi clinical trials. Clin. Cancer Res. 22, 567–574 (2016).
Riester, M. et al. PureCN: copy number calling and SNV classification using targeted short read sequencing. Source Code Biol. Med. 11, 13 (2016).
Oh, S. et al. Reliable analysis of clinical tumor-only whole-exome sequencing data. JCO Clin. Cancer Inf. 4, 321–335 (2020).
Atkinson, V. et al. Optimizing combination dabrafenib and trametinib therapy in BRAF mutation-positive advanced melanoma patients: guidelines from Australian melanoma medical oncologists. Asia Pac. J. Clin. Oncol. 12, 5–12 (2016).
Menzies, A. M. et al. Characteristics of pyrexia in BRAFV600E/K metastatic melanoma patients treated with combined dabrafenib and trametinib in a phase I/II clinical trial. Ann. Oncol. 26, 415–421 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Schroder, J. et al. Socrates: identification of genomic rearrangements in tumour genomes by re-aligning soft clipped reads. Bioinformatics 30, 1064–1072 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Daud, A. I. et al. Programmed death-ligand 1 expression and response to the anti-programmed death 1 antibody pembrolizumab in melanoma. J. Clin. Oncol. 34, 4102–4109 (2016).
Acknowledgements
We thank the patients and their families for their participation. We also thank the study site staff, additional investigators, R. Leary, C. Unitt and S. Mahan (Next Generation Diagnostics) for their contributions. We thank K. Gibbs for biomarker sample management, as well as A. Savchenko, J. Choi, C. Wong, B. Fu, G. Gorgun and R. Ramesh for support with biomarker analyses. We thank Navigate Biopharma for DNA and RNA extraction, as well as HistoGeneX and Bioagilytix for biomarker testing. We also thank M. Voi (Novartis Pharmaceuticals) for guidance and critical review. We thank A. Lytle and A. Ghiretti (ArticulateScience LLC) for providing medical writing support, which was funded by Novartis Pharmaceuticals Corporation in accordance with Good Publication Practice guidelines (http://www.ismpp.org/gpp3). COMBI-i (NCT02967692) is sponsored by Novartis Pharmaceuticals.
Author information
Authors and Affiliations
Contributions
C.R., D.S., H.A.T., J.C.B., E.G. and G.V.L. conceived or designed the work. R.D., C.L., V.A., M.M., P.D.N., A.A., E.R., N.Y., C.R., D.S., H.A.T., P.A.A., A.R., N.P., C.D.C., K.T.F., D.G., A.M., J.C.B., E.G. and G.V.L. acquired, analyzed or interpreted the data. R.D., C.L., A.A., E.R., N.Y., C.R., D.S., H.A.T., A.R., K.T.F., J.C.B., E.G. and G.V.L. drafted or substantively revised the work.
Corresponding author
Ethics declarations
Competing interests
R.D. reports intermittent, project-focused consulting and/or advisory relationships with Novartis, Merck Sharp & Dohme (MSD), Bristol Myers Squibb (BMS), Roche, Amgen, Takeda, Pierre Fabre, Sun Pharma, Sanofi, Catalym, Second Genome, Regeneron and Alligator outside the submitted work. C.L. reports research funding from Roche and BMS, speakers bureau for Roche, BMS, MSD, Amgen, Novartis and Pierre Fabre, and consulting and advisory roles for BMS, MSD, Roche, Novartis, Merck Serono, Sanofi and Pierre Fabre. V.A. reports advisory roles for BMS, Merck Serono, MSD, Novartis, Roche, Nektar and Pierre Fabre, and received speaker’s fees from BMS, Merck, MSD and Novartis, and travel support from BMS, MSD and Onco-Sec. M.M. reports consulting or advisory roles for MSD, Roche, BMS and Pierre Fabre and research funding from Roche, Novartis and BMS. P.D.N. reports advisory roles for BMS, Immunocore, Merck, MSD, Novartis and Pfizer, speaker’s bureaux for BMS and Novartis and steering committee membership for Immunocore, Merck and Novartis. A.A. reports personal fees and other from BMS, MSD, Roche, Novartis, Merck, Sanofi, Amgen and Pierre Fabre, outside the submitted work. E.R. reports consulting or advisory roles for Amgen, Bayer, BMS, MSD, Merck, Novartis, Pierre Fabre, Roche and Sanofi; reception of honoraria from Amgen, Bayer, BMS, MSD, Merck, Novartis, Pierre Fabre, Roche and Sanofi; speaker’s bureaux for Amgen, BMS, MSD, Merck, Novartis, Pierre Fabre and Sanofi; research funding from Amgen, BMS, MSD, Novartis, Pierre Fabre and Roche; travel support from Amgen, BMS, MSD, Merck, Novartis, Pierre Fabre, Roche and Sanofi; and President of the Austrian Cancer Aid/Styria. N.Y. reports consulting and advisory roles for Novartis, Ono, BMS and MSD, honoraria from Novartis, Ono, BMS and MSD and institutional research support from Novartis, Ono, BMS, MSD and Takara-Bio. C.R. reports consulting or advisory roles for BMS, Roche, Amgen, Novartis, Pierre Fabre, MSD, Sanofi, Biothera, CureVac and Merck. D.S. reports research funding from Novartis and BMS. H.A.T. reports consulting or advisory roles for Novartis, BMS, Roche-Genentech, Merck and Array BioPharma and research funding from BMS, Novartis, Merck, GlaxoSmithKline, Genentech/Roche and Celgene. P.A.A. reports consulting or advisory roles for BMS, Roche-Genentech, MSD, Array, Novartis, Merck Serono, Pierre Fabre, Incyte, NewLink Genetics, Genmab, Medimmune, AstraZeneca, Syndax, Sun Pharma, Sanofi, Idera, Ultimovacs, Sandoz, Immunocore, 4SC, Alkermes, Italfarmaco, Nektar and Boehringer Ingelheim, research funding from BMS, Roche-Genentech and Array and travel support from MSD. A.R. reports serving as a consultant/independent contractor for, and being the recipient of honoraria from, Amgen, Chugai, Merck, Novartis and Sanofi; advisory roles and receipt of honoraria from Arcus, Bioncotech, Compugen, CytomX, ImaginAb, Isoplexis, Merus, Rgenix, Lutris, PACT Pharma and Tango Therapeutics; self-managed stock shareholder in Arcus, Compugen, CytomX and Merus; and research support from Agilent and BMS. K.T.F. reports advisory roles for Clovis Oncology, Strata Oncology, Vivid Biosciences, Checkmate Pharmaceuticals, X4 Pharmaceuticals, PIC Therapeutics, Sanofi, Amgen, Asana, Adaptimmune, Fount, Aeglea, Stattuck Labs, Tolero, Apricity, Oncoceutics, Fog Pharma, Neon, Tvardi, xCures, Monopteros and Vibliome; consulting roles for Lilly, Novartis, Genentech, BMS, Merck, Takeda, Verastem, Boston Biomedical, Pierre Fabre and Debiopharm; research funding from Novartis and Sanofi; and stock shareholder in Clovis Oncology, Strata Oncology, Vivid Biosciences, Checkmate Pharmaceuticals, X4 Pharmaceuticals, PIC Therapeutics, Fount, Shattuck Labs, Apricity, Oncoceutics, Fog Pharma, Tvardi, xCures, Monopteros and Vibliome. N.P. reports employment at Novartis Healthcare Pvt. Ltd. C.D.C. reports employment with, and stock in, Novartis Pharmaceuticals. D.G. reports employment with Novartis Institutes for BioMedical Research. A.M. reports employment by Novartis and stock or ownership in Novartis and BMS. J.C.B. reports employment by, and stock ownership in, Novartis, and is a coinventor on a patent application related to reported biomarker subgroups of interest. E.G. reports employment by, and stock ownership in, Novartis. G.V.L. reports consultant advisory roles for Aduro Biotech, Inc., Pierre Fabre Medicament, BMS, Amgen, MSD, Novartis Pharma, Array BioPharma, Syneos and Sandoz Biopharmaceuticals. All authors received support for third-party medical writing and editorial assistance provided by ArticulateScience LLC and funded by Novartis Pharmaceuticals Corporation.
Additional information
Peer review information Javier Carmona was the primary editor on this article, and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Study designs for (a) part 1 and (b) part 2 of COMBI-i.
ALT, alanine aminotransferase; AST, aspartate aminotransferase; BID, twice daily; CD, cluster of differentiation; CNS, central nervous system; DCR, disease control rate; DLT, dose-limiting toxicity; DOR, duration of response; ECOG PS, Eastern Cooperative Oncology Group performance status; FFPE, formalin-fixed paraffin-embedded; ORR, overall response rate; OS, overall survival; PD, progressive disease; PD-L1, programmed death ligand 1; PFS, progression-free survival; PK, pharmacokinetics; Q4W, every 4 weeks; Q8W, every 8 weeks; QD, once daily; RECIST, Response Evaluation Criteria in Solid Tumors; RP3R, recommended phase 3 regimen; S, screening; ULN, upper limit of normal.a BRAF V600 mutation was assessed based on local testing (followed by central confirmation using the bioMérieux THxID-BRAF assay). b With systemic therapy including checkpoint inhibitors, targeted therapy, chemotherapy, biologic therapy, tumor vaccine therapy, or investigational treatment for unresectable or metastatic melanoma; prior intralesional, adjuvant, or neoadjuvant therapy was allowed if completed ≥ 6 months prior to start of study treatment, and prior radiation therapy was allowed if completed ≥ 4 weeks prior to start of study treatment. c DL-1b: DLT observation period starts on cycle 2, day 1 (C2D1 [day 29]). Patients who did not tolerate dabrafenib and/or trametinib and discontinued during the first 4 weeks were to be replaced due to insufficient exposure.
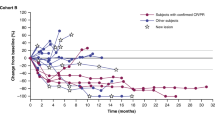
Extended Data Fig. 2 Best percent change from baseline in sum of diameters by local investigator review (N = 36).
A total of 33 patients experienced a reduction in the size of the target lesion. One patient with SD had a best percent change of 0% in the target lesion. Best percent change in the target lesion was not available for 1 patient with PD. Best percent change in target lesion could not be calculated for 1 additional patient as best overall response was unknown. CR, complete response; PD, progressive disease; PR, partial response; SD, stable disease.
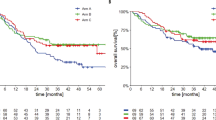
Extended Data Fig. 3 Time-to-event analyses, including (a) DOR, (b) PFS, and (c) OS, in patients enrolled in parts 1 and 2 of COMBI-i (N = 36).
12- and 24-month DOR rates were 80% (95% CI, 59%-91%) and 53% (95% CI, 29%-73%); 12- and 24-month PFS rates were 67% (95% CI, 49%-80%) and 41% (95% CI, 23%-59%); 12- and 24-month OS rates were 86% (95% CI, 70%-94%) and 74% (95% CI, 56%-86%). DOR, duration of response; PFS, progression-free survival; NE, not estimable; OS, overall survival.
Extended Data Fig. 4 Markers of response to immunotherapy were not associated with CR.
Baseline T-cell–inflamed GES levels and TMB in samples from patients with and without a CR. For T-cell–inflamed GES: n = 27 independent tumor biopsy specimens (CR, n = 14; no CR, n = 13). For TMB: n = 24 independent tumor biopsy specimens (CR, n = 12; no CR, n = 12). Box plots show median, first and third quartiles (boxes), and range up to 1.5 times IQR from the bounds of the box [whiskers]. Points beyond 1.5 times IQR from the bounds of the box are plotted individually. For T-cell–inflamed GES: CR, 4.90 (4.31-5.27) [3.48-6.09]; no CR, 4.87 (3.99-5.30) [2.88-6.65]. For TMB: CR, 7.196 (5.957-9.205) [3.365-13.479]; no CR, 7.209 (6.639-9.140) [3.539-10.533]. Descriptive P values are based on a two-sided Wilcoxon rank sum test (T-cell–inflamed GES: W = 89, effect size -0.04 [95% CI, -0.75-0.83]; TMB: W = 79, effect size 0.05 [95% CI, -2.40-2.85]); no adjustments were made for multiple comparisons. CPM, counts per million; CR, complete response; GES, gene expression signature; IQR, interquartile range; TMB, tumor mutational burden.
Extended Data Fig. 5 Correlative analysis of GES levels and tumor shrinkage following treatment with spartalizumab in combination with dabrafenib and trametinib.
Association between a, T-cell–inflamed GES levels and b, PI3K pathway gene expression and best overall tumor reduction, based on Spearman correlation coefficient. n = 27 independent tumor biopsy specimens. CPM, counts per million; GES, gene expression signature.
Extended Data Fig. 6 Evidence of immune activation during treatment with spartalizumab in combination with dabrafenib and trametinib.
a, Analysis of intratumoral density of CD8+ cells for available paired tumor biopsy specimens (n = 9) at baseline and on treatment using exploratory H-score analysis. Scale bars = 100 μm. b, Modulation of plasma IFN-γ following treatment with spartalizumab plus dabrafenib plus trametinib. Of the 27 independent plasma specimens analyzed, 25 showed elevated IFN-γ levels on treatment, while the other 2 showed a slight decrease. Box plot shows median, first and third quartiles (boxes), and range up to 1.5 times IQR from the bounds of the box [whiskers]. Points beyond 1.5 times IQR from the bounds of the box are plotted individually. Baseline, 1.78 (1.06-2.37) [1.06-4.19]. On treatment, 6.08 (3.65-7.88) [1.69-10.51]. IFN, interferon; IHC, immunohistochemistry; IQR, interquartile range.
Supplementary information
Supplementary Information
Supplementary Tables 1–8 and Figs. 1–3.
Supplementary Tables
Supplementary Tables 9–11. Full listings of all gene sets for which P < 0.05 in unbiased analyses (Table 9: PFS ≤12 months versus all others; Table 10: CR versus all others; Table 11: on-treatment modulations).
Rights and permissions
About this article
Cite this article
Dummer, R., Lebbé, C., Atkinson, V. et al. Combined PD-1, BRAF and MEK inhibition in advanced BRAF-mutant melanoma: safety run-in and biomarker cohorts of COMBI-i. Nat Med 26, 1557–1563 (2020). https://doi.org/10.1038/s41591-020-1082-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-1082-2
This article is cited by
-
Binimetinib in combination with nivolumab or nivolumab and ipilimumab in patients with previously treated microsatellite-stable metastatic colorectal cancer with RAS mutations in an open-label phase 1b/2 study
BMC Cancer (2024)
-
Combined therapy of dabrafenib and an anti-HER2 antibody–drug conjugate for advanced BRAF-mutant melanoma
Cellular & Molecular Biology Letters (2024)
-
New clinical trial design in precision medicine: discovery, development and direction
Signal Transduction and Targeted Therapy (2024)
-
Epigenetic reprogramming of Runx3 reinforces CD8 + T-cell function and improves the clinical response to immunotherapy
Molecular Cancer (2023)
-
A cytosolic mutp53(E285K) variant confers chemoresistance of malignant melanoma
Cell Death & Disease (2023)