Abstract
In a human menstrual cycle the endometrium undergoes remodeling, shedding and regeneration, all of which are driven by substantial gene expression changes in the underlying cellular hierarchy. Despite its importance in human fertility and regenerative biology, our understanding of this unique type of tissue homeostasis remains rudimentary. We characterized the transcriptomic transformation of human endometrium at single-cell resolution across the menstrual cycle, resolving cellular heterogeneity in multiple dimensions. We profiled the behavior of seven endometrial cell types, including a previously uncharacterized ciliated cell type, during four major phases of endometrial transformation, and found characteristic signatures for each cell type and phase. We discovered that the human window of implantation opens with an abrupt and discontinuous transcriptomic activation in the epithelia, accompanied with a widespread decidualization feature in the stromal fibroblasts. Our study provides a high-resolution molecular and cellular characterization of human endometrial transformation across the menstrual cycle, providing insights into this essential physiological process.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All raw and analyzed sequencing data associated with Figs. 1, 3–5 and 6a–c, all Extended Data figures and all supplementary figures in this study can be found at NCBI’s Gene Expression Omnibus (series accession code GSE111976) and Sequence Read Archive (accession code SRP135922). Data associated with Fig. 2, extracted from all representative fields of view, can be found in Supplementary Dataset 1. Source data are provided with this paper.
Code availability
Custom codes developed in this study for gene dispersion calculation, MI calculation and transcriptional regulator and secretory protein analyses are available at https://github.com/wanxinw/endometrium/. Specific codes are accessible, with no restrictions, upon request to W.W. (wanxinw@stanford.edu).
References
Martin, R. D. The evolution of human reproduction: a primatological perspective. Yearb. Phys. Anthropol. 50, 59–84 (2007).
Emera, D., Romero, R. & Wagner, G. The evolution of menstruation: a new model for genetic assimilation: explaining molecular origins of maternal responses to fetal invasiveness. BioEssays 34, 26–35 (2012).
Crichton, E. & Krutzsch, P. Reproductive Biology of Bats (Elsevier, 2000).
Bellofiore, N. et al. First evidence of a menstruating rodent: the spiny mouse (Acomys cahirinus). Am. J. Obstet. Gynecol 216, 40E.1–40E.11 (2017).
Noyes, R. W., Hertig, A. T. & Rock, J. Dating the endometrial biopsy. Fertil. Steril. 1, 3–25 (1950).
Croxatto, H. B. et al. Studies on the duration of egg transport by the human oviduct. II. Ovum location at various intervals following luteinizing hormone peak. Am. J. Obstet. Gynecol. 132, 629–634 (1978).
Wilcox, A. J., Baird, D. D. & Weinberg, C. R. Time of implantation of the conceptus and loss of pregnancy. N. Engl. J. Med. 340, 1796–1799 (1999).
Riesewijk, A. et al. Gene expression profiling of human endometrial receptivity on days LH+2 versus LH+7 by microarray technology. Mol. Hum. Reprod. 9, 253–264 (2003).
Ruiz-Alonso, M., Blesa, D. & Simón, C. The genomics of the human endometrium. Biochim. Biophys. Acta Mol. Basis Dis. 1822, 1931–1942 (2012).
Díaz-Gimeno, P. et al. A genomic diagnostic tool for human endometrial receptivity based on the transcriptomic signature. Fertil. Steril. 95, 50–60 (2011).
Van Der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 620, 267–284 (2008).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331–D338 (2017).
Zhou, F. & Roy, S. SnapShot: motile cilia. Cell 162, 224–224 (2015).
Schwab, K. E. & Gargett, C. E. Co-expression of two perivascular cell markers isolates mesenchymal stem-like cells from human endometrium. Hum. Reprod. 22, 2903–2911 (2007).
Masuda, H., Anwar, S. S., Bühring, H. J., Rao, J. R. & Gargett, C. E. A novel marker of human endometrial mesenchymal stem-like cells. Cell Transplant. https://doi.org/10.3727/096368911X637362 (2012).
Guo, Y., Manatunga, A. K., Chen, S. & Marcus, M. Modeling menstrual cycle length using a mixture distribution. Biostatistics 7, 100–114 (2006).
Tkačik, G. & Walczak, A. M. Information transmission in genetic regulatory networks: a review. J. Phys. Condens. Matter 23, 153102 (2011).
Talbi, S. M. et al. Molecular phenotyping of human endometrium distinguishes menstrual cycle phases and underlying biological processes in normo-ovulatory women. Endocrinology 147, 1097–1121 (2006).
Park, Y., Nnamani, M. C., Maziarz, J. & Wagner, G. P. Cis-regulatory evolution of forkhead box O1 (FOXO1), a terminal selector gene for decidual stromal cell identity. Mol. Biol. Evol. 33, 3161–3169 (2016).
Okada, H. et al. Regulation of decidualization and angiogenesis in the human endometrium: mini review. J. Obstet. Gynaecol. Res. 40, 1180–1187 (2014).
Ramathal, C. Y., Bagchi, I. C., Taylor, R. N. & Bagchi, M. K. Endometrial decidualization: of mice and men. Semin. Reprod. Med. 28, 17–26 (2010).
Hood, B. L. et al. Proteomics of the human endometrial glandular epithelium and stroma from the proliferative and secretory phases of the menstrual cycle. Biol. Reprod. 92, 106 (2015).
Lessey, B. A., Metzger, D. A., Haney, A. F. & McCarty, K. S. Immunohistochemical analysis of estrogen and progesterone receptors in endometriosis: comparison with normal endometrium during the menstrual cycle and the effect of medical therapy. Fertil. Steril. 51, 409–415 (1989).
Lessey, B. A. et al. Immunohistochemical analysis of human uterine estrogen and progesterone receptors throughout the menstrual cycle. J. Clin. Endocrinol. Metab. 67, 334–340 (1988).
Zhou, J., Dsupin, B. A., Giudice, L. C. & Bondy, C. A. Insulin-like growth factor system gene expression in human endometrium during the menstrual cycle. J. Clin. Endocrinol. Metab. 79,1723–1734 (1994).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Suryawanshi, H. et al. A single-cell survey of the human first-trimester placenta and decidua. Sci. Adv. 4, eaau4788 (2018).
Tulac, S. et al. Identification, characterization, and regulation of the canonical Wnt signaling pathway in human endometrium. J. Clin. Endocrinol. Metab. 88, 3860–3866 (2003).
Fan, X. et al. Dynamic regulation of Wnt7a expression in the primate endometrium: implications for postmenstrual regeneration and secretory transformation. Endocrinology 153, 1063–1069 (2012).
Yin, Y. & Ma, L. Development of the mammalian female reproductive tract. J. Biochem. 137, 677–683 (2005).
Tempest, N., Baker, A. M., Wright, N. A. & Hapangama, D. K. Does human endometrial LGR5 gene expression suggest the existence of another hormonally regulated epithelial stem cell niche? Hum. Reprod. 33, 1052–1062 (2018).
Lessey, B. A. et al. Luminal and glandular endometrial epithelium express integrins differentially throughout the menstrual cycle: implications for implantation, contraception, and infertility. Am. J. Reprod. Immunol. 35, 195–204 (1996).
Kelleher, A. M. et al. Integrative analysis of the forkhead box A2 (FOXA2) cistrome for the human endometrium. FASEB J. 33, 8543–8554 (2019).
Jeong, J.-W. et al. Foxa2 is essential for mouse endometrial gland development and fertility. Biol. Reprod. 83, 396–403 (2010).
Filant, J. & Spencer, T. E. Endometrial glands are essential for blastocyst implantation and decidualization in the mouse uterus. Biol. Reprod. 88, https://doi.org/10.1095/biolreprod.113.107631 (2013).
Hanna, J. et al. Decidual NK cells regulate key developmental processes at the human fetal–maternal interface. Nat. Med. 12, 1065–1074 (2006).
Krjutskov, K. et al. Single-cell transcriptome analysis of endometrial tissue. Hum. Reprod. 31, 844–853 (2016).
Wu, B. et al. Cell atlas of human uterus. Preprint at bioRxiv https://doi.org/10.1101/267849 (2018).
Lucas, E. S. et al. Recurrent pregnancy loss is associated with a pro-senescent decidual response during the peri-implantation window. Commun. Biol. 3, 37 (2020).
Benda, C. Klenusches Handbuch der Han und Sexualorgane (Hansebooks, 1894).
Turco, M. Y. et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat. Cell Biol. 19, 568–577 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Mann, H. B. & Whitney, D. R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Stat. 18, 50–60 (1947).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Hastie, T. & Stuetzle, W. Principal curves. J. Am. Stat. Assoc. 84, 502–516 (1989).
Petropoulos, S., Edsga, D., Reinius, B. & Linnarsson, S. Single-cell RNA-Seq reveals lineage and X chromosome dynamics in human preimplantation resource single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Kim, T. H. et al. Single-cell transcript profiles reveal multilineage priming in early progenitors derived from Lgr5+ intestinal stem cells. Cell Rep. 16, P2053–P2060 (2016).
Marco, E. et al. Bifurcation analysis of single-cell gene expression data reveals epigenetic landscape. Proc. Natl Acad. Sci. USA 111, E5643–E5650 (2014).
Ji, Z. & Ji, H. TSCAN: pseudo-time reconstruction and evaluation in single-cell RNA-seq analysis. Nucleic Acids Res. 44, e117 (2016).
Lachmann, A., Giorgi, F. M., Lopez, G. & Califano, A. ARACNe-AP: gene network reverse engineering through adaptive partitioning inference of mutual information. Bioinformatics 32, 2233–2235 (2016).
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res. 25, 1860–1872 (2015).
Whitfield, M. L. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, P1888–P1902 (2019).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, P329–P337 (2019).
Acknowledgements
We thank H. Ding for valuable discussions and advice; N. Neff and J. Okamoto for sequencing expertise; F. B. Yu, S. Darmanis and F. Zanini for technical expertise and discussions; S. Crasta and S. Kolluru for technical assistance; and the Stanford Cell Science Imaging Facility for assistance with imaging. This study was jointly supported by the March of Dimes, Chan Zuckerberg Biohub and MINECO/FEDER (no. SAF-2015-67164-R, to C.S.) (Spanish Government). W.W. was supported by the Stanford Bio-X Graduate Bowes Fellowship and Chan Zuckerberg Biohub. F.V. was supported by the Miguel Servet Program Type II of ISCIII (no. CPII18/00020) and the FIS project (no. PI18/00957). P.A. was supported by IVI-RMA Valencia. I.M. was supported by the Igenomix Foundation. M.M. and A.I. were supported by Chan Zuckerberg Biohub. W.P. was supported by the March of Dimes.
Author information
Authors and Affiliations
Contributions
W.W., W.P., F.V., C.S. and S.R.Q. conceived and designed the study. W.W., F.V., I.M. and A.I. performed experiments. W.W. performed single-cell experiments, RNAscope experiments and imaging. A.I. performed single-cell experiments. F.V. optimized the tissue dissociation protocol. I.M. performed tissue dissociation and immunofluorescence experiments. P.A. collected endometrial biopsies. W.W. and S.R.Q. analyzed single-cell RNA-seq data. M.M. and W.W. analyzed RNAscope data. W.W., F.V., C.S. and S.R.Q. interpreted the results. W.W., F.V., C.S. and S.R.Q. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A patent disclosure has been filed for the study under the inventors S.R.Q., C.S., W.W. and F.V. C.S. is Founder and Head of the Scientific Advisory Board of Igenomix SL. P.A., I.M., M.M., A.I. and W.P. declare no competing interests.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Data summary.
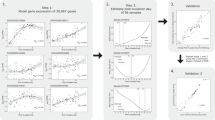
a, Number of single cells analyzed across the human menstrual cycle in the C1 dataset. b, C1: Relationship between the day of menstrual cycle (day) and endometrial phase/subphase assigned by single cell transcriptome data as in Fig. 4 for each woman (dot) in the C1 dataset. Arrows annotate discrepancy between day and phase/subphase where 1) orders were reversed for a pair of women between their orders in phase/subphase assignment and their cycle day (red) and 2) women on the same day of cycle were assigned into different phases or subphases (black). 10x: The day of menstrual cycle for each woman (dot) in the 10x dataset. Dots with * and # to the right are anchors shared between the two datasets (Methods). c, For each woman (dot: top, middle panel; bar: bottom panel) in the C1 dataset: (top) relationship between her pseudotime order (x-axis) and her cycle day, (middle) total number of single cells analyzed, (bottom) distribution of the six cell types identified by the C1 dataset. From left to right, women were ordered based on the median pseudotime of all her stromal fibroblasts and unciliated epithelia. Pseudotime and phase were as assigned in Fig. 3a. Related to Fig. 1, 3, 4.
Extended Data Fig. 2 Summary of cell types and states in the 10x validation dataset.
a, Dimension reduction (UMAP on top PCs) on 71032 single cells from 10 healthy human endometria. Integration was done across donors using Seurat IntegrateData() to decouple temporal signal. b, Distribution of cycling cells. c, Relative abundance of each cell type/subtype identified. Each black dot is a woman. Dots in pink are medians. d, Total number of cells analyzed per cell type/subtype. e, Expression of top discriminatory genes identified by the C1 dataset (Fig. 1b) and for smooth muscle cells. f, Expression of markers identifying cells with mesenchymal stem cell features in the human endometrium. g, Combinatorial expression of PDGFRB and MCAM in smooth muscle cells. In brackets: abundance of each of the four cell groups normalized by the total smooth muscle cell count. (Expression values were jittered (amount=0.1) for visualization.) h, Percent cells expressing SUSD2 in the four cell groups identified in (g). Related to Fig. 1, 5.
Extended Data Fig. 3 Constructing single cell resolution trajectories of the human menstrual cycle using mutual information (MI) based approach.
a, Dimension reduction (tSNE) using whole transcriptome data, that is all genes detected, for unciliated epithelia (epi) and stromal fibroblasts (str), respectively, led to identification of four major phases (insets). b, MI between expressions of each gene and time (red) or permutated time (black). Genes were ranked by their MI with time. c, tSNE using ‘time-associated genes’ for epi and str, respectively led to the identification of the same four major phases as in (a) and trajectories of endometrial transformation. In a, c main panels, cells were colored by the day of menstrual cycle (day). Related to Fig. 3.
Extended Data Fig. 4 Global temporal transcriptome dynamics across the human menstrual cycle.
a, MI between expressions of each pseudotime-associated gene (FDR<1E-05) and pseudotime (red) or permutated pseudotime (black) for unciliated epithelia (epi) and stromal fibroblasts (str). b, Dynamics of pseudotime- associated genes across the menstrual cycle. Related to Fig. 4.
Extended Data Fig. 5 Dynamics of WOI-associated genes in the 10x validation dataset.
a-b, Dimension reduction (UMAP on top PCs) on unciliated epithelia (a) and stromal fibroblasts (b) in the 10x dataset. Highlighted are anchors between the C1 and 10x datasets (Methods). Phases were assigned based on relative order between each cell group and the anchors. c, Relationship between the day of menstrual cycle and phase assignment. In blue are anchors. d-e, Dynamics of WOI-associated genes in Fig. 4 stratified by epithelial subpopulations (d) and stromal fibroblasts (e). MTIX, FGF7, and LMCD1 peaked at non-WOI phases in the C1 dataset and were included as references. White dots: medians; Boundaries of black bars: 25% and 75% quantiles. Highlighted in blue are cell groups where the anchors grouped with. Related to Fig. 4.
Extended Data Fig. 6 Dynamics of genes encoding transcriptional factors (TF) across the menstrual cycle.
a-b, All time-associated TFs for unciliated epithelia (epi, a) and stromal fibroblasts (str, b) (TFs bracketed by red bars were zoomed in c, d). c-d, TFs that are associated with the entrance/exit of WOI (bottom panel) or phase-defining (top) in epi (c) and str (d). e, Expression of TFs that are nuclear hormone receptors for steroid hormones estrogen (ESR1), progesterone (PGR), glucocorticoid (NR3C1), and androgen (AR). For heatmap, TFs were ordered first by the pseudotime of the major peak and then by the pseudotime of the peak’s inflection point. Full list of pseudotime-associated TFs can be found in Supplementary Table 2. Related to Fig. 4.
Extended Data Fig. 7 Dynamics of genes encoding secretory proteins (sec genes) across the menstrual cycle.
a-b, All time-associated sec genes for unciliated epithelia (epi, a) and stromal fibroblasts (str, b) (sec genes bracketed by blue bars were zoomed in c, d). c-d, Sec genes that are associated with the entrance/exit of WOI (bottom) in epi (c) and str (d). e-f, Expression of sec genes encoding canonical protein markers for decidualization during pregnancy in epi (e) and str (f). For heatmap, sec genes were ordered following the same strategy as in Extended Data Fig. 6. Full list of pseudotime-associated sec genes can be found in Supplementary Table 3. Related to Fig. 4.
Extended Data Fig. 8 Expression of genes that define human early-pregnant endometrial stromal fibroblast subtypes.
a, Abundance of stromal fibroblasts (str) and unciliated epithelia (epi) expressing IGFBP1, PRL, or both in non-pregnant human endometria across the menstrual cycle. Phases were assigned as in Extended Data Fig. 5. b, Distribution of stromal fibroblasts co-expressing IGFBP1 and PRL on the reduced dimension of all late-secretory stromal fibroblasts. c, Dispersion level of all genes detected in late-secretory stromal fibroblasts. Highlighted are genes reported to differentially express in the two early-pregnant subtypes that do not co-express IGFBP1 and PRL. d, Some of the subtype genes that are highly dispersed in (c), for example ACTA2, IGFBP1, showed a subtle expression gradient (arrow) on dimension-reduced late-secretory stromal fibroblasts. In c, d, red and blue indicate genes reported to define the two early-pregnant subtypes, respectively.
Extended Data Fig. 9 Dynamics of cell cycling activity across the menstrual cycle and transcriptomic signature of proliferative phases.
a-b, Endometrial G1/S and G2/M signatures for unciliated epithelia (epi, a) and stromal fibroblasts (str, b) identified in the C1 dataset. c-d, Distribution (left) and abundance (right) of cycling cells across major phases of the menstrual cycle. e-f, Top discriminatory genes for the two proliferative phases identified in epi (e) and str (f). Related to Fig. 4.
Extended Data Fig. 10 Deviating subpopulations in unciliated epithelia and their transcriptomic signatures across the menstrual cycle.
a. Dimension reduction (UMAP on top PCs) on all unciliated epithelia in the 10x validation dataset and distribution of cycling cells. Integration was done across donors using Seurat IntegrateData() to decouple temporal signal. b, Expression of markers differentially expressed between the two subpopulations that were identified in the C1 dataset. In black are genes in Fig. 5b. In orange are genes in Fig. 5c, d that were previously shown to be differentially expressed by glandular or luminal epithelia of human endometria in situ. c, Dynamics of differentially expressed genes between the two subpopulations in proliferative phases identified in the C1 dataset. Related to Fig. 5.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4.
Supplementary Data 1
Supplementary Dataset 1. Data were extracted from all representative fields of view captured for Fig. 2a–d and were used to generate Fig. 2e.
Source data
Source Data Fig. 3.
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, W., Vilella, F., Alama, P. et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle. Nat Med 26, 1644–1653 (2020). https://doi.org/10.1038/s41591-020-1040-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-1040-z
This article is cited by
-
Luteal phase support in assisted reproductive technology
Nature Reviews Endocrinology (2024)
-
Effect of aging on the human myometrium at single-cell resolution
Nature Communications (2024)
-
Applications of nature-inspired metaheuristic algorithms for tackling optimization problems across disciplines
Scientific Reports (2024)
-
Differential gene expression in decidualized human endometrial stromal cells induced by different stimuli
Scientific Reports (2024)
-
Characterizing the Extracellular Matrix Transcriptome of Endometriosis
Reproductive Sciences (2024)